Project 3 SS_Yeh, George_poster FINAL
-
Upload
george-yeh -
Category
Documents
-
view
84 -
download
2
Transcript of Project 3 SS_Yeh, George_poster FINAL

GGTTTT
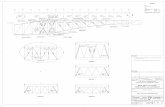
Selective Regulation of IL12b Transcription by NF-κB cRel George Yeh1, Abraham Chang1, Trevor Siggers2, Kevin Wiliams3, Shomyseh Sajanbi4, and Stephen Smale1,4
1Molecular Biology Inst., UCLA, 2Dept. of Biology, Boston U. , 3Dept.of Pathology, UCLA, 4Dept.of Microbiology and Immunology, UCSF, and 4Dept.of Microbiology, Immunology, and Molecular Genetics, UCLA
Abstract Introduction
Summary
Combinatorial gene regulation principles and experimental results support the view that typical transcription factors directly regulate hundreds of genes in a given cell type. Surprisingly, using genome-wide RNA-seq and ChIP-seq assays, we discovered that, in macrophages stimulated with bacterial products, the cRel member of the NF-κB family of transcription factors makes a major contribution to the regulation of only one gene: the IL12b gene, which encodes the p40 subunit of IL12 and IL23. Only a small number of genes were significantly mis-regulated in cRel -/- macrophages and the magnitude of IL12b mis-regulation was much greater than observed at any other gene. Furthermore, ChIP-seq experiments revealed that, although RelA and c-Rel bind similarly to over 8000 sites throughout the genome, the IL12b promoter was unique in its ability to support much stronger binding to cRel than to RelA. Biochemical studies revealed that this preferential and highly selective binding of cRel to the IL12b promoter is due to differences in the intrinsic binding properties of cRel and RelA. These results suggest that key immunoregulatory genes are regulated by highly unique mechanisms, raising the possibility that these unique mechanisms might be attractive therapeutic targets for the treatment of diseases associated with the immune system.
Contact: George Yeh [email protected]
Stephen T. Smale [email protected]
IL12b Transcription is Dependent on 46 Residues of cRel Located in the N-terminus of Rel Homology Domain (RHD)
IL12b is a Subunit of IL-12 and IL-23 Cytokines
Cooperative Binding to the IL12b is Unique to cRel Homodimer
Therapeutic Potentials of Targeting IL12 (Il12b)
Goals
1. Elucidate the MECHANISM that explains IL12b’s strong dependence on cRel transcription factor. a. Generate chimeric proteins of cRel and RelA to identify key regions for IL12b
transcription. b. Utilize electrophoretic mobility shift assay (EMSA) to prove cRel binds
stronger than RelA at IL12b promoter. c. Perform protein binding microarray (PBM) to recognize novel DNA motifs
that may prefer cRel binding over RelA.
2. Explore the degree of SELECTIVITY in which cRel is able to exquisitely regulate a specific set of genes. a. Use chromatin RNA-seq to scrutinize cRel’s selectivity on gene transcription
at genome-wide scale. b. Perform cRel and RelA ChIP-seq to peruse their binding disparity at genome-
wide scale.
NFκB protein cRel is exclusively dedicated to the transcription of IL12b due to its unique binding properties. Namely, it strongly and cooperatively binds to IL12b promoter non-consensus κB sites, to which RelA does poorly. Strikingly, this unique cRel property is only observed in IL12b promoter even on a genome-wide scale, as evident by whole-genome analysis of RNA-seq and ChIP-seq experiments. This exquisite selectivity may be an attractive target to build therapeutic strategies on for diseases that are related to aberrant IL12 or IL23 expression.
Through the generation of chimeric proteins and their expressions in cRel -/- macrophages, we identified a region of cRel, which consists of 46 residues of DNA binding domain that is critical for IL12 expression. This is our first clue that the IL12b dependency on cRel has something to do with DNA binding.
cRel is Required for IL12b Gene Transcription in LPS-Stimulated Macrophages
• It is well-established that the control of IL12 can be therapeutically affective. For example, STELARA®, a human monoclonal antibody, blocks IL12b and is an effective treatment for psoriasis.
• In contrast, in case of infectious diseases like leprosy, we would like to boost IL12 expression as it limits the reproduction of M. leprae.
In 2000, we first discovered that cRel is important for IL12b expression. When stimulating macrophages with LPS, where as RelA and p50 knockouts only had a moderate effect on IL12b expression, the absence of cRel almost completely abolished IL12b expression. This is the same in both peritoneal as well as fetal liver macrophages.
IL12a Il23a IL12b
IL12 IL23
Naïve CD4+ T Cell
Th1
Th17
Th2
IL-6 TGFβ
IL-23
• IL12b forms IL12 heterodimer with IL12a, and IL23 with IL23a. • Both cytokines are very important for immune responses. • For example, IL23 is important for stimulating T helper 17 cell. • IL12, on the other hand, is important for T helper 1 cell differentiation,
proliferation, and maintenance. • Particularly, Th1 response is important for controls of intracellular pathogens,
including microbacterial infections like Mycobacterium leprae.
Psoriasis - Treatment with IL12b antibody (STELARA®; FDA approved)
Leprosy - Lepromatous patients (severe)
- Low IL12 expression - Tuberculoid patients (controlled)
- High IL12 expression
Results
Peritoneal Fetal-liver derived Peritoneal Macrophages Fetal Liver Macrophages PBM Discovered Novel DNA Motif that Prefers cRel Homodimer
1) cRel TD
4) RelA (C46) 3) cRel (A46)
0 5 10 15 20
ng/ml of IL12 (p40/p70)
2) RelA
RHD(N)
Chimeric RelA (C46)
RelA amino acids
46 cRel residues
dsDNA
ELISA Assay in cRel -/-
cRel Binds to Non-consensus κB site with a Higher Affinity than RelA
IL12b (κB2) AAAATTCCCC
Consensus κB GGAAATTCCC
cRel 1.2±0.1 0.4±0.1
RelA 51.4±6.5 1.1±0.1
RelA(C46) 4.4±0.4 0.8±0.1
RelA/cRel 42.8
2.8
RelA/(C46) 11.7
1.4
Frac
tion
Boun
d
[NFκB] (nM)
RelA cRel
RelA (C46)
(Concentration of protein required to bind 50% DNA in EMSA)
Mouse IL12b -154 AGGGGGGGAGGGAGGAACTTCTTAAAATTCCCCCAGAATGTTTTGACACTA -104
Surface Plasmon Resonance
Offrate - t1/2 (s)
RelA/RelA Binding Affinity
cRel
/cRe
l Bin
ding
Aff
inity
DNA RelA/RelA cRel/cRel RelA(C46)
GGGGGGAGAT 4±1 4±1 8±5
GGGGGTTTTT 7±1 58±11 28±6
IL12b promoter sequence NF-kB 1 NF-kB 2 NF-kB 3 NF-kB 4 C/EBP Mouse GGAGGAACTTCTTAAAATTCCCCCAGAATGTTTTGACACTAGTTTTCAGTGTTGCAATTGAGACTAG *********** ************** ******* * * ******* ******** * ** Human AAAGGAACTTCTTGAAATTCCCCCAGAAGGTTTTGAGAGTTGTTTTCAATGTTGCAA------CAAG -125/-116 -115/-106 -104/-95 -92/-83 -80/-72
NF-kB1 NF-kB2 NF-kB3 EMSA probe - AGGAACTTCTTAAAATTCCCCCAGAATGTTTTGACA
RelA/RelA p50/p50 RelA/p50 cRel/p50 c-Rel/c-Rel
Chromatin RNA-seq Revealed Strikingly Selective Dependence of IL12b Transcription on cRel at Genome-wide Scale
IL12b
1 SD Mean
1 SD 2 SD
2 SD
WT vs cRel -/-
WT Log2 (Max RPKM)
cRel
Log
2 (M
ax R
PKM
)
ChIP-seq with RelA and cRel Exhibits Exquisite Selective cRel Binding at IL12b Promoter at Genome-wide Scale
1 SD 2 SD
1 SD 2 SD
IL12b Promoter
Mean
cRel : RelA Binding Ratio in All Peaks
8669 Called Peaks
cRel : RelA Binding Ratio in Promoters of Inducible Genes
IL12b Promoter
38 Promoter Peaks
Non-consensus κB site
Ratio
(Log
2)
Ratio
(Log
2)
Out of ~8000 peaks, the peak in IL12b promoter is ranked the 2nd showing bias towards cRel binding over RelA. The peak ranked 1st is about 22,000 bp away from the nearest TSS. No transcriptional aberration is observed for that gene in cRel -/- RNA-seq (Arghap35).
If we only look within the promoters of genes strongly induced by lipid A, IL12b is the ONLY gene that stands out as the one that strongly prefers cRel binding over RelA.













![SURP Final Paper [Final] DW](https://static.fdocument.org/doc/165x107/5881c6c61a28ab87638b46b3/surp-final-paper-final-dw.jpg)