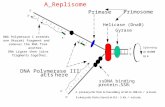
The quarternary structure of DNA Polymerase ε from S. cerevisiae#
Transcript of The quarternary structure of DNA Polymerase ε from S. cerevisiae#

1
The quarternary structure of DNA Polymerase ε from S. cerevisiae# Olga Chilkova1, Bengt-Harald Jonsson2 and Erik Johansson1* 1 Department of Medical Biochemistry and Biophysics Umeå University SE-901 87 UMEÅ Sweden 2 IFM/ Molecular Biotechnology Linköpings University SE-581 83 LINKÖPING Sweden Running Title: Quarternary structure of DNA polymerase ε Inquiries to: Erik Johansson TEL: +46-(0)90-7866638 FAX: +46-(0)90-7869795 E-mail: [email protected]
# This work was supported by the Swedish Cancer Society, Swedish Research Council, the Magnus Bergwalls stiftelse, and Kungliga vetenskapsakademiens stipendiefond. *Correspondence to this author.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

2
SUMMARY
DNA polymerase ε (Pol ε) from S. cerevisiae consists of four subunits (Pol2, Dpb2,
Dpb3, and Dpb4) and is essential for chromosomal DNA replication. Biochemical
characterizations of Pol ε have been cumbersome due to protease sensitivity and the
limited amounts of Pol ε in cells. We have developed a protocol for over-expression
and purification of Pol ε from S. cerevisiae. The native four-subunit complex was
purified to homogeneity by conventional chromatography. Pol ε was characterized
biochemically by sedimentation velocity experiments and gel filtration experiments.
The stoichiometry of the four subunits was estimated to be 1:1:1:1 from colloidal
Coomassie-stained gels. Based on the sedimentation coefficient (11.9 S) and the
Stokes radius (74.5 Å), a molecular mass for Pol ε of 371 kDa was calculated, in good
agreement with the calculated molecular mass of 379 kDa for a heterotetramer.
Furthermore, analytical equilibrium ultracentrifugation experiments support the
proposed heterotetrameric structure of Pol ε. Thus, both DNA polymerase δ and Pol ε
are purified as monomeric complexes, in agreement with accumulating evidence that
Pol δ and Pol ε are located on opposite strands of the eukaryotic replication fork.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

3
INTRODUCTION
Pioneering studies in the SV40 DNA replication system by several research groups
resulted in the reconstitution of the SV40 replication fork with mammalian proteins in
vitro (1). This system showed that DNA polymerase α is required for priming the
DNA since it is associated with primase activity. Only DNA polymerase δ (Pol δ) was
required to replicate both leading and lagging strand, leaving another essential
eukaryotic DNA polymerase, DNA polymerase ε (Pol ε), without a specific function.
However, there are several lines of evidence for the presence of Pol ε at the
eukaryotic replication fork. Evidence for a physical interaction came from cross-
linking experiments which showed that Pol ε is located at or near the replication fork
(2). Genetic experiments in S. cerevisiae have shown that Pol ε participates at the
replication fork. Cells grown in the presence of 6-hydroxylaminopurine showed a
strand specific mutation rate on the chromosome, depending on whether they carried
an exonuclease deficient Pol ε or Pol δ (3). It has also been shown that loss of the
exonuclease site in either Pol δ or Pol ε alters the mutation spectra of the sup-4 gene
when replicated on a plasmid (4). The altered mutational spectrum was again strand
specific. In addition, the overall mutation rate is increased about ten-fold in yeast
strains that carry a pol2 mutant that is lacking proofreading capability, suggesting that
Pol ε contributes to the bulk synthesis of newly replicated DNA (5). Evidence for a
direct role during the progression of the fork has emerged from in vitro replication
assays in Xenopus extracts in a cell-free system (6).
A very surprising result was obtained when the catalytic domain of Pol ε was deleted
in yeast (7). The catalytic domain was not essential. The essential property of the
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

4
enzyme was instead located at the C-terminus of the largest subunit(7;8). Cells that
were expressing only the C-terminus were viable, although they had growth problems.
However, later it was shown that cells were not viable when carrying a point mutation
in the conserved catalytic site of Pol ε (9). Although Pol ε is normally present at the
fork, in the absence of the catalytic domain, another DNA polymerase, presumably
Pol δ, can substitute its function.
Pol ε consists of four subunits, Pol2p (256 kDa), Dpb2p (79 kDa), Dpb3p (23 kDa),
and Dpb4p (22 kDa) (10). The largest subunit, Pol2p, contains the catalytic site and
an essential putative zinc-finger domain at the C-terminus (9). The other three
subunits have an unknown function. There has been very limited biochemical
characterization of the 4-subunit enzyme, since it was first purified in 1990 (10). This
is in part due to the limited amount of enzyme in the cells, the tedious purification
protocol, and the sensitivity of the enzyme to proteolytic cleavage (10;11).
Several models of the structure of the eukaryotic replication fork have been proposed
(1;12-14). A common theme has been the coupled synthesis of the leading and
lagging strands in which the replicative DNA polymerase should form a dimer as has
been observed in E. coli (15;16). Earlier studies suggested the existence of a dimeric
Pol δ with two catalytic assemblies, and a model where a dimeric Pol δ would
replicate both leading and lagging strand was proposed (13;17). However, it has now
been demonstrated that Pol δ actually is a monomer in vitro, and that the previous
estimates of the molecular weight of Pol δ were misleading due to the elongated shape
of the enzyme (18). The initial analysis of the 4-subunit Pol ε included an estimate of
the molecular weight and the subunit stoichiometry (10). The molecular weight was
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

5
estimated to be 295 kDa from the hydrodynamic properties of the enzyme, and the
molar ratio of the four subunits was estimated at Pol2/Dpb2/Dpb3/Dpb4 =1:1:3:4.
This molar ratio predicted a molecular mass of 492 kDa, which was substantially
different from that determined experimentally. The authors concluded that the enzyme
was purified as a monomer with an elongated shape. Pol ε has also been proposed to
form a dimer based on yeast two-hybrid assays and gel filtration experiments using
Pol ε overproduced in a baculovirus system (9;19). In light of these contradictory
results we wished to clarify whether Pol ε exist as a monomer or a dimer when
overproduced in yeast. Therefore, we developed an over-expression system that
allowed us to purify native Pol ε from yeast, i.e. without tags. Our studies of the
assembly state of this enzyme show it to be a monomeric assembly of one each of the
four subunits.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

6
EXPERIMENTAL PROCEDURES
Strains - A protease deficient yeast strain was used for the over-expression of Pol ε.
PY116 (ura 3-52, trp 1∆, his 3-11, 15, pep 4-4, prb 1-1122 (prc1-407) (CANsens), leu
2-3, 112, nuc 1: LEU 2) was transformed with plasmids pJL1 and pJL6 (see below).
This strain for over-expression of Pol ε was called YEJ1. PJL1 contained the POL2
gene and pJL6 contained the DPB2, DPB3 and DPB4 genes under the control of the
GAL1-10 promoter. The plasmids carried a 2µ origin and either TRP1 or URA3 as
selective markers.
Plasmids – All plasmids were based on the pRS424GAL and pRS426GAL from the
pRS424-426GAL series of plasmids (a kind gift from P. Burgers). The POL2 gene
was amplified from the plasmid YCplac33 POL2 (20) with the following
oligonucleotides; POL2-1: 5´-
TTCTAGAGCGGCCGCATGATGTTTGGCAAGAAAAAAAACAACGG -3´;
POL2-2: 5´-
AACACGAGCTCTCATATGGTCAAATCAGCAATACAACTCAATAA -3´. The
resulting DNA fragment was digested with NotI and SacI, purified and ligated into
prs424GAL opened in NotI and SacI. The 500 nucleotides most approximate to NotI
and SacI were sequenced to make sure that no mutations were inserted by KlenTAQ
LA during the amplification of the insert. After this, the over-expression plasmid was
digested with PshAI and NgoMIV to remove most of the POL2 gene. The YCplac33
POL2 plasmid was also digested with PshAI and NgoMIV and the 6151 nt fragment
containing the POL2 gene was subcloned into the opened prs424GAL-POL2 plasmid.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

7
The resulting plasmid was named pJL1. DPB2, DPB3, and DPB4 were amplified
from W303 genomic DNA with the following oligonucleotides; DPB2-1: 5´-
TTCTAGGATCCATGTGTGAAATGTTTGGCTCTGGGA -3´; DPB2-2: 5´-
TTCTAGAGCGGCCGCAAACGGTAGGCCAAGTAAACTGCCC -3´; DPB3-1: 5´-
TTCTAGAGCGGCCGCTCAACCGTGTTGCAAAAAAAATGTCC -3´; DPB3-2:
5´- AACACGAGCTCGTAATTGTGGCACAGGCAAGCTG -3´; DPB4-1: 5´-
TTCTAGAGAATTCATGCCACCAAAAGGTTGGAGAAAAG -3´; DPB4-2: 5´-
AACACGTCGACATATTCAGTTGCTCTAATCGGAGC -3´. The amplified DPB2
gene was digested with BamHI and NotI, purified and ligated into the BamHI-NotI
sites of prs424GAL. The resulting plasmid was called pJL2. The amplified DPB3
gene was digested with NotI and SacI, purified and ligated into the NotI-SacI sites of
prs426GAL. This plasmid was called pJL3. The amplified DPB4 gene was digested
with EcoRI and SalI, purified and ligated into the EcoRI-SalI sites of prs426GAL.
This plasmid was called pJL4. Next, DPB2, DPB3 and DPB4 were sequenced to
confirm that no mutations had been inserted by the polymerase during the
amplification. This was followed by the digestion of pJL3 and pJL4 with BamHI and
SacI. The fragment from pJL3 that contained DPB3 was cloned into the BamHI-SacI
sites of the pJL4 plasmid. This plasmid was digested with SacI and blunt-ended with
Klenow. pJL2 was digested with SapI and HincII, over-hangs were filled in by
Klenow and DPB2 was ligated into the opened blunt-ended plasmid containing DPB3
and DPB4. The resulting plasmid, prs426GAL with DPB2, DPB3, and DPB4, was
called pJL6.
Cell growth - For Pol ε over-expression the YEJ1 strain was grown as previously
described in (21). Briefly, the YEJ1 strain was grown in 100 ml of Glycerol-lactate
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

8
media overnight. After 24 hours, the culture was aliquoted into six 6 liter flasks, each
containing 1.2 liter glycerol lactate media, and left vigorously shaking at 30 °C. After
another 24 hours 1.2 l of YPGL media was added to every flask. YPGL media
consists of 1% yeast extract, 2% bactopeptone, 3% glycerol, 2% lactic acid and 0.1%
glucose, pH 5.5-6. After 3-4 hours the protein expression was induced by adding solid
galactose to a final concentration of 2%. After another 6 hours the cells were
harvested by centrifugation for 5 min at 4 ˚C. The cells were resuspended in H2O and
frozen in liquid nitrogen.
Enzyme purification - All purification steps were carried out at 4 ˚C. The following
buffers were used. Buffer A: 150 mM Tris-Ac pH 7.8, 50 mM NaAc, 2 mM EDTA, 1
mM EGTA, 10 mM NaHSO3, 1 mM dithiothreitol, 5 µM pepstatin A, 5 µM
leupeptin, 0.3 mM p-phenylmethylsulfonyl fluoride, 5 mM benzamidine. Buffer B: 25
mM Hepes-NaOH pH 7.6, 10% glycerol, 1 mM EDTA, 0.5 mM EGTA, 0.005%
Nonidet P-40, 1 mM DTT, 5 µM pepstatin A, 5 µM leupeptin, 5 mM NaHSO3. The
concentration of sodium acetate is indicated as suffix, for example, buffer B100 =
buffer B with 100 mM NaAc. A beadbeater was used to open 60-70 g of cells in
Buffer A. Beating continued for 5 min with intervals of 1 min cooling every 30
seconds. The volume of the lysate from the beadbeater was measured and saturated
ammonium sulfate was added to a final concentration of 175 mM, followed by the
addition of 40 µl 10 % Polymin P per ml of extract. The lysate was incubated on ice
with occasional stirring for 15 min, followed by centrifugation at 18,000 rpm in a
Beckman JA25.5 rotor for 1 hour. The volume of the cleared lysate was measured and
0.28 g of solid ammonium sulfate was added per ml of extract. The ammonium sulfate
was dissolved for 45 min on a magnetic stirrer, followed by centrifugation at 18,000
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

9
rpm in a Beckman JA25.5 rotor for 1 hour. The ammonium sulfate precipitate was
resuspended in buffer B50 and frozen. The next day, the frozen extract was thawed and
dialyzed against 1000 ml of buffer B100 for 3 hours, and thereafter centrifuged for 30
min. The cleared lysate was loaded onto a 20 ml phosphocellulose column
equilibrated in B200. The column was washed with B200 and eluted with B750. Fractions
with protein were loaded onto a MonoQ column eluted with a 20 ml linear gradient
from B800 to B1200. After dialysis against 500 ml of buffer B100, peak fractions from
MonoQ were loaded onto a MonoS column equilibrated in B100 and eluted with a 20
ml linear gradient from B100 to B500. Finally, the MonoS peak fraction was passed
over a Superose6 gel filtration column as described below. The protein concentration
in various fractions throughout the purification was determined by the method of
Bradford (22) with BSA as a standard.
Gel filtration analysis - A Superose 6 gel filtration column was equilibrated at 4 °C in
Buffer C. Buffer C: 25 mM Hepes pH 7.6, 10% glycerol, 1 mM EDTA, 0.005%
Nonidet P-40, 400 mM NaAc, 5 mM dithiothreitol, 5 mM NaHSO3, 2 µM leupeptin, 2
µM pepstatin A. The column was calibrated by injection of proteins with a known
Stokes radius. The Stokes radii of the standard proteins were plotted against their
elution volumes. The standard proteins used were: carbonic anhydrase (Ve = 18.07
ml), bovine serum albumin (Ve = 16.68 ml), catalase (Ve = 15.92 ml), ferritin (Ve =
14.82 ml), E. coli β-galactosidase (Ve = 14.45 ml), and thyroglobulin (Ve = 12.65 ml).
0.2-20 µg Pol ε from the MonoS peak fraction was passed over the equilibrated
superose6 column. Fractions were analyzed by SDS-PAGE and the polymerase
activity was measured. The specific activity of purified Pol ε was determined to be
25-40000 U/mg protein, as defined by Hamatake et.al. (10).
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

10
Glycerol Gradient Centrifugation — 0.2-20 µg of Pol ε from the MonoS peak fraction
together with 0.5 µg of bovine liver catalase (Sigma) as an internal control, in a total
volume of 100 µl, were layered on top of a 4 ml 15–30% linear gradient of glycerol in
buffer C. The samples were spun for 16 h at 40,000 rpm in a Beckman SW60 rotor at
1°C. Fractions were collected from the bottom of the tube, and aliquots were analyzed
by SDS-PAGE, DNA polymerase assays or Western-blot analysis with Dpb4p
antibodies. The rabbit Dpb4p polyclonal antibodies were raised against Dpb4p that
was over-expressed and purified from E.coli. Aliquots of each fraction were also
removed for spectrophotometric assay of catalase activity (23). Whole cell extracts
from wild-type yeast were prepared as previously described, and cleared lysate was
loaded onto the 15-30% glycerol gradient in buffer C400. The density of glycerol
gradient fractions was measured refractometerically. Standard proteins with known
sedimentation coefficients were used to calibrate the gradients: carbonic anhydrase
(2.8 S), bovine serum albumin (4.3 S), aldolase (8.9 S), catalase (11.3 S) and ferritin
(17.6 S).
Polymerase assays - For DNA polymerase assays an oligo(dT)16 primed poly(dA)300
template was used (24). A typical reaction (50 µl) contained 20 mM Tris-HCl pH 7.5,
4% glycerol, 0.1 mg/ml bovine serum albumin, 5 mM dithiothreitol, 8 mM MgAc2, 80
µM each dATP, dCTP, and dGTP, 20 µM [3H]dTTP (400-600 cpm/pmol), 0.01 U
poly(dA)⋅oligo(dT), 1 mM spermidine and enzyme. Reactions were assembled on ice
and incubated for 30 sec to 5 min at 30 °C. The reactions were stopped by the
addition of 150 µl stop solution (50 mM sodium pyrophosphate, 25 mM EDTA and
50 µg/ml salmon sperm DNA). The insoluble material was precipitated by adding 1.5
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

11
ml of 10% trichloroacetic acid, followed by a 30 min incubation on ice. The insoluble
material was filtered on GF/C filters, washed 4x2 ml with 1 M HCl in 0.05 M sodium
pyrophosphate, rinsed with ethanol, dried, and counted by liquid scintillation. One
unit of enzyme activity incorporates 1 nmol of total nucleotide / hour (10).
Sedimentation equilibrium experiments - The analytical ultracentrifugation
experiments were carried out at 10 °C in a Beckman XL-A/XL-I analytical
ultracentrifuge, equipped with an An50Ti rotor using six sector centerpieces.
Sedimentation equilibrium experiments on Pol ε were analyzed in Buffer D (as Buffer
C but with 5 mM 2-mercaptoethanol instead of dithiothreitol) at three different initial
protein concentrations, 2.1 µM, 0.28 µM and 0.14 µM. Absorbance at 280 nm was
measured as a function of radial position for three rotor speeds, ranging from 4000 to
7500 rpm. Equilibrium at each rotor speed was reached after 20 hours. At that time,
five scans were averaged for each sample at each rotor speed. The sedimentation
properties of Pol ε were analyzed by using the self association model in the Beckman
software package.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

12
RESULTS
We chose to over-express Pol ε in S. cerevisiae for several reasons. Earlier attempts to
over-express Pol ε in a baculovirus-system experienced several problems (19;25). Not
only was the largest subunit of 256 kDa expressed at low levels, but in addition Pol2p
was cleaved by a protease and the proteolytic fragment copurified with the
holoenzyme. Finally the purified protein was tagged and this could affect the
properties of the enzyme. We chose to over-express Pol ε in S. cerevisiae without any
tags added, similar to earlier over-expression studies of Pol δ from S. cerevisiae (18).
The POL2 gene was subcloned into one plasmid and DPB2, DPB3, and DPB4 were
subcloned into the second plasmid. All genes were cloned under the control of the
galactose-inducible GAL1-10 promoter (21). This protocol proved successful and
allowed us to purify 400 µg of Pol ε from as little as 70 g of cells. The complex was
purified by successive phosphocellulose, MonoQ, and MonoS chromatography, and
the enzyme was finally purified to homogeneity by gel filtration column (Fig 1) or by
glycerol gradient centrifugation.
Our over-expressed Pol ε was purified as a four-subunit complex with all subunits co-
migrating through a glycerol gradient and co-eluting from a Superose6 column (Fig
2B and Fig 3B). The peak fractions from the gel filtration column and the glycerol
gradient were analyzed by 10% SDS-PAGE, the stained gels were scanned and the
intensity of the protein bands was quantified. The molar ratio of the four subunits was
estimated to be 1.00:1.12:1.07:1.06 (Pol2p:Dpb2p:Dpb3p:Dpb4p) (Fig 4). In contrast,
we found that the apparent molar ratio of the four subunits was approximately 1:1:3:4,
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

13
when the SDS-PAGE was silver-stained (data not shown). Earlier estimates of the
subunit stoichiometry have been based solely on silver-staining, and it appears that
the amounts of Dpb3p and Dbp4p may have been overestimated due to the silver-
staining (10;19).
The sedimentation coefficient for Pol ε was determined to be 11.9 S (Fig 2). Using an
antibody to Dpb4p we measured the sedimentation coefficient of Pol ε, when a crude
extract from a wild-type strain was separated over the same glycerol gradient. The
sedimentation of Pol ε in a crude extract was similar to that of the purified Pol ε (data
not shown).
Pol ε was analyzed on a Superose6 gel filtration column (Fig 3). The enzyme was
injected over the column in the concentration range between 5 nM and 0.5 µM (as
monomer) and the elution volume was 13.8 ml over the entire concentration range.
The Stokes radius was determined to be 74.5 Å from the standard curve. The
measured Stokes radius and the sedimentation coefficient allowed us to estimate the
Mr, according to Siegel and Monty (26). Our calculated Mr of 371 kDa suggests that
Pol ε is purified as a monomer of a Pol2:Dpb2:Dpb3:Dpb4 complex with the molar
ratio of 1:1:1:1 in the concentration range studied (Table I). The frictional coefficient
f/fo of Pol ε is 1.56, suggesting a moderately elongated shape of the enzyme.
Sedimentation equilibrium experiment. To obtain a more precise molecular mass
independent of the shape of the enzyme, we carried out sedimentation equilibrium
ultracentrifugation. Three different starting concentrations of Pol ε ranging from 0.14
µM to 2.1 µM were analyzed at four different speeds until equilibrium was reached.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

14
The molecular weight of Pol ε was determined to be 366 kDa under these conditions.
A comparison with the predicted distribution of a monomeric versus a dimeric
assembly clearly showed that Pol ε was present as a monomer in the solution under all
conditions investigated (Fig 5).
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

15
DISCUSSION
We have presented a method to over-express and purify Pol ε to homogeneity from S.
cerevisiae. The yield and purity of the enzyme will allow us to perform a careful
characterization of the enzyme. We have initiated this study by determining the
molecular weight and subunit stoichiometry of the enzyme. These are essential
parameters for further studies of the enzyme. From these studies and recent studies of
Pol δ we can conclude that, at least under the conditions studied, both enzymes are
monomeric multi-subunit assemblies with each subunit occurring once.
An attractive model of the replication fork describes how there should be coupling of
leading and lagging strand synthesis. This is a model that has emerged from studies in
E. coli (15;16). It is very likely that a similar mechanism is present at the eukaryotic
replication fork. Initial experiments with Pol δ suggested that Pol δ was a dimeric
enzyme (17;27). A model was proposed in which Pol δ formed a dimeric unit with
two catalytic domains. In this model Pol δ would replicate both leading and lagging
strand in a coordinated manner (13). This model has recently been questioned and it
has been shown that Pol δ is a monomer with one catalytic site (18;28). Meanwhile,
Pol ε was also suggested to be dimeric (19). The evidence for a dimeric enzyme
originated from yeast two-hybrid assays and gel filtration experiments (9;19). Yeast
two-hybrid assays, particularly with yeast genes, can be misleading because false
positives can result from a bridging interaction through a third yeast protein, as for
instance was observed with the third subunit of Pol δ. Pol32 appeared to interact with
itself in a two-hybrid analysis, but this interaction was actually mediated by PCNA
(18;29). A second line of evidence for a dimeric Pol ε was based upon gel filtration
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

16
experiments (19). The calculated molecular weight of the protein was based upon the
assumption that Pol ε was globular. We have measured both the sedimentation
coefficient and the Stokes radius of Pol ε and from those data calculated the frictional
coefficient to be 1.56 (Table I), indicating that the complex is slightly elongated. As a
consequence, the molecular weight will be overestimated when determined by gel-
filtration only (19). The calculated molecular weight of 371 kDa from both gel
filtration and density centrifugation measurements agrees well with the theoretical
molecular weight of a four subunit complex with the subunit stoichiometry of 1:1:1:1
(Table I). Finally, the sedimentation equilibrium experiments confirmed the results of
our other studies where Pol ε was found to exist as a monomer in solution (Fig 5 and
Table I). Thus, our results clearly support a model of Pol ε as a monomer of a
(Pol2:Dpb2:Dpb3:Dpb4) heterotetrameric complex.
The role of Pol ε and Pol δ at the eukaryotic replication fork is still not known with
certainty. Genetic experiments in S. cerevisiae have shown that there is an asymmetry
present at the replication fork (30). The current hypothesis that the coupled synthesis
of the leading and lagging strand involves both Pol ε and Pol δ agrees with our
observations that both Pol δ and Pol ε are monomeric assemblies. Pol δ has been
suggested to replicate the lagging strand, based on a genetic interaction with the flap
endonuclease (RAD27) (31). Pol ε has been suggested to replicate the leading strand,
based on experiments in cell free Xenopus extracts (6). However, the fact that both
Pol δ and Pol ε are purified as monomers in solution does not exclude the possibility
that they dimerize in the presence of a template. The yeast two-hybrid result
indicating that Dpb2p may interact with itself may suggest that Pol ε could be present
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

17
as a dimer in vivo (19). More experiments are required in order to determine which
strands Pol ε and Pol δ do replicate.
We are now in a position to study the various activities of Pol ε using a well-defined
source of the enzyme. These studies may help us understand the role of Pol ε during
DNA replication and DNA repair.
ACKNOWLEDGEMENTS
We thank P. Burgers for comments on the manuscript and providing the prs421-
424GAL plasmids and pY116 strain.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

18
Reference List
1. Waga, S. and Stillman, B. (1998) Annu.Rev.Biochem. 67, 721-751
2. Aparicio, O. M., Weinstein, D. M., and Bell, S. P. (1997) Cell 91, 59-69
3. Shcherbakova, P. V. and Pavlov, Y. I. (1996) Genetics 142, 717-726
4. Karthikeyan, R., Vonarx, E. J., Straffon, A. F. L., Simon, M., Faye, G., and Kunz, B. A. (2000) Journal of Molecular Biology 299, 405-419
5. Morrison, A. and Sugino, A. (1994) Mol.Gen.Genet. 242, 289-296
6. Waga, S., Masuda, T., Takisawa, H., and Sugino, A. (2001) PNAS 98, 4978-4983
7. Kesti, T., Flick, K., Keranen, S., Syvaoja, J. E., and Wittenberg, C. (1999) Mol.Cell 3, 679-685
8. Navas, T. A., Zhou, Z., and Elledge, S. J. (1995) Cell 80, 29-39
9. Dua, R., Levy, D. L., and Campbell, J. L. (1999) J.Biol.Chem. 274, 22283-22288
10. Hamatake, R. K., Hasegawa, H., Clark, A. B., Bebenek, K., Kunkel, T. A., and Sugino, A. (1990) J.Biol.Chem. 265, 4072-4083
11. Liu, W. and Linn, S. (2000) Nucleic Acids Res. 28, 4180-4188
12. Falaschi, A. (2000) Trends Genet. 16, 88-92
13. Burgers, P. M. (1998) Chromosoma 107, 218-227
14. Kawasaki, Y. and Sugino, A. (2001) Mol.Cells 12, 277-285
15. Glover, B. P. and McHenry, C. S. (2001) Cell 105, 925-934
16. Yuzhakov, A., Turner, J., and O'Donnell, M. (1996) Cell 86, 877-886
17. Burgers, P. M. and Gerik, K. J. (1998) J.Biol.Chem. 273, 19756-19762
18. Johansson, E., Majka, J., and Burgers, P. M. (2001) J.Biol.Chem. 276, 43824-43828
19. Dua, R., Edwards, S., Levy, D. L., and Campbell, J. L. (2000) J.Biol.Chem. 275, 28816-28825
20. Araki, H., Ropp, P. A., Johnson, A. L., Johnston, L. H., Morrison, A., and Sugino, A. (1992) EMBO J. 11, 733-740
21. Burgers, P. M. (1999) Methods 18, 349-355
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

19
22. Bradford, M. M. (1976) Anal.Biochem. 72, 248-254
23. Chance, B. and Herbert, D. (1950) Biochem.J. 46, 402-414
24. Burgers, P. M. (1995) Methods Enzymol. 262, 49-62
25. Dua, R., Levy, D. L., Li, C. M., Snow, P. M., and Campbell, J. L. (2002) J.Biol.Chem. 277, 7889-7896
26. Siegel, L. M. and Monty, K. J. (1966) Biochim.Biophys.Acta 112, 346-362
27. Zuo, S., Bermudez, V., Zhang, G., Kelman, Z., and Hurwitz, J. (2000) J.Biol.Chem. 275, 5153-5162
28. Bermudez, V. P., MacNeill, S. A., Tappin, I., and Hurwitz, J. (2002) J.Biol.Chem. 277, 36853-36862
29. Gerik, K. J., Li, X., Pautz, A., and Burgers, P. M. (1998) J.Biol.Chem. 273, 19747-19755
30. Pavlov, Y. I., Newlon, C. S., and Kunkel, T. A. (2002) Mol.Cell 10, 207-213
31. Jin, Y. H., Obert, R., Burgers, P. M., Kunkel, T. A., Resnick, M. A., and Gordenin, D. A. (2001) Proc.Natl.Acad.Sci.U.S.A 98, 5122-5127
FOOTNOTES
1The abbreviations used are: Pol ε, DNA polymerase ε; Pol δ, DNA polymerase δ; PCNA, proliferating cell nuclear antigen.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

20
LEGENDS TO FIGURES
Fig 1. Over-expressed Pol ε was purified to homogeneity. 1 µg of Pol ε from the
Superose6 column was loaded on a 4-20% linear gradient SDS-polyacrylamide gel.
The proteins were visualized by colloidal Coomassie-staining.
Fig 2. Glycerol gradient centrifugation A. Gradient calibration was carried out with
ferritin (data point 1, 17.6 S), catalase (data point 2, 11.3 S), aldolase (data point 3,
8.9 S), bovine serum albumin (data point 4, 4.3 S), and carbonic anhydrase (data
point 5, 2.8 S). The sedimentation positions of the complexes are as indicated. The
estimated sedimentation coefficient of Pol ε was 11.88 S +0.84 S. The sedimentation
coefficient of Pol ε was measured over a concentration range from 5 nM to 0.5 µM
(as monomer), with no observed shift in S value. B. Fractions containing Pol ε were
analyzed on a 10% SDS-PAGE and visualized with colloidal Coomassie-staining.
Catalase was added as an internal standard before loading the sample and the catalase
activity was measured in a colorimetric assay. The peak fraction containing catalase
activity has been indicated with catalase. The contrast of the shown gel was enhanced
to visualize Dpb3 and Dpb4.
Fig 3. Gelfiltration. A. The calibration of the Superose6 column was carried out with
thyroglobulin (data point 1, 85 Å), β-galactosidase (data point 2, 69 Å), ferritin (data
point 3, 61 Å), catalase (data point 4, 52.2 Å) bovine serum albumin (data point 4,
35.5 Å), and carbonic anhydrase (data point 5, 23.9 Å). The elution volumes of the
complexes are as indicated. Pol ε eluted at 13.8 ml and the Stokes radius was
estimated to be 74.5 Å +2.4 Å. B. Fractions containing Pol ε were analyzed on a 10%
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

21
SDS-PAGE and visualized with colloidal Coomassie-staining. The contrast of the
shown gel was enhanced to visualize Dpb3 and Dpb4.
Fig 4. Subunit stoichiometry of Pol ε. Peak fractions from the Superose6 column were
separated on a 10% SDS-PAGE and stained with colloidal Coomassie. The gel was
digitized with a CanonFB1210U scanner and the lanes were scanned and quantified
using NIH Image software. The molar ratio of the subunits was
Pol2p:Dpb2p:Dpb3p:Dpb4p = 1:1.12:1.07:1.06.
Fig 5. Sedimentation equilibrium experiment. Pol ε with a starting concentration of
2.1 µM was centrifuged at three consecutive speeds, beginning with 4000 rpm. The
theoretical curves for the monomeric and dimeric species are calculated based upon
the polypeptide sequence.
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

22
Table I. Molecular weight of DNA polymerase ε. Mr values in kDa and Stokes radii in Å. Theoretical Mr values were calculated from the polypeptide sequences; Mr and f/fo from S and Å was obtained from (26); Mr from equilibrium sedimentation centrifugation was calculated with the Beckman software package. Indicated errors in the calculated Mr reflect measurement errors only. See text for details. Theoretical Mr for S value Stokes Mr from f/f0 Mr from monomer dimer radius S and Å equil. sed. Pol ε 379 758 11.88 +0.84 74.5 +2.4 371 +38 1.56 366 +7
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

Pol2p Dpb2p Dpb3p Dpb4p
Fig4R
elat
ive
Inte
nsity
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

Abs
orba
nce
(280
nm
)
Radius (mm)
Fig 5
6.3 6.4 6.5 6.6
1.0
1.5
2.5
0.5
2.0
0.5
2.0
3.0
6.3 6.4 6.5 6.6
1.0
1.5
2.5
3.5
3.5
6.3 6.4 6.5 6.6
0.5
1.0
1.5
2.0
2.5
3.0
Monomer
Dimer
7500 rpm
5500 rpm
4000 rpm
Monomer
Dimer
Monomer
Dimer
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from

Olga Chilkova, Bengt-Harald Jonsson and Erik JohanssonALIGN="BASELINE" ALT="epsilon "> from S. cerevisiae
The quarternary structure of DNA polymerase <IMG SRC="/math/epsilon.gif"
published online February 5, 2003J. Biol. Chem.
10.1074/jbc.M211818200Access the most updated version of this article at doi:
Alerts:
When a correction for this article is posted•
When this article is cited•
to choose from all of JBC's e-mail alertsClick here
by guest on February 16, 2018http://w
ww
.jbc.org/D
ownloaded from