A physical map of the bioAB region in the λbio transducing phage
Click here to load reader
Transcript of A physical map of the bioAB region in the λbio transducing phage

104
Appendix
A physical map of the bioAB region in the X bio transducing phage
(E. coli bio operon; regulatory region: heteroduplex mapping; electron
tions)
microscopy; IS1 insertion; dele-
Graiyna Konopa , *a Waclaw Szybalski **a, Jeroo Kotval *b and Allan Campbell b
0 McArdle Laboratory for Cancer Research, University of Wisconsin, Madison, WI 53706, und ’ Deportment oj Biologicul
Sciences, Stanford University, Stanford, CA 94305 (U.S.A.)
(Received April 2nd, 1982)
(Accepted May 5th. 1982)
SUMMARY
The center of the p’,opC, promoter-operator region for the bioABFCD operon is located 1.7 1 kb clockwise
from the attX site on the Escherichia coli genome, as determined by the position of the ~13 1 (ISI ) insertion.
The order of several bio endpoints to the right of p131 is Xbio267, 122, 169, 74, 1, and 69. The endpoints of
the two bio deletions, A61 in bioA and A3h in bioB, were also determined.
In the preceding communication the physical
structure of the entire bioABFCD operon and
neighboring regions was outlined, and the position
of the controlling p’,o& region indicated on the
basis of the restriction mapping and sequencing
data of Otsuka and Abelson (1978) and Barker et
al. (198 1). Here we present our electron micro-
scopic heteroduplex data which locate the ~131
(ISI ) insertion (Ketner and Campbell, 1975) in
relation to attB and specify the position of several
nearby bio endpoints and deletions in relation to
* Present addresses: (G.K.): Department of Biochemistry, Uni-
versity of Gdadsk, “1. KIadki 24, 80-822 Gdarisk (Poland);
(J.K.): Department of Microbiology and Immunology, Albany
Medical College, Albany, NY 12208 (U.S.A.).
** To whom reprint requests should be directed.
Abbreviations: bp, base pairs; IS, insertion sequence; kb, kilo-
base pairs; o, operator; p, promoter; A, deletion; %X unit. 485
bp; Xarr2, XarrLarrR or XattBOP’atrPOB’ (Fiandt et al., 1976).
0378-I 119/82/0000-0000/$02.75 ‘Q 1982 Elsevier Biomedical Press
this ISI marker (see Figs. 1 and 2, and Table I).
Deletion A61 removes part of the bioA gene and
fuses it to the gal region. This deletion was crossed
into the phage XgalSbio69cI857 (Ketner and
Campbell, 1974; Barker and Campbell, 1980) re-
sulting in Xgaf8(A61)bio69cI857. Phage Xbio3h-1
of Kayajanian (1968) carries within the bio DNA
the A3h deletion extending from bioB to uvrB
(Szybalski and Szybalski, 1982). Other bio end-
points shown in Fig. 1 are within gene bioB with
the exception of hbiol, which contains the entire
gene bioB, and hbio69, which also contains most
of bioC. Phage hbio267, 122, 169, and 74 were
isolated by J.K., and some of them were used for
testing the effect of rightward function on leftward
bioA transcription (Kotval et al., 1982).
The present data complement those of Szybal-
ski and Szybalski (1982) and suggest that the
center of the p,op, region is located about 1.71 kb
from the attR site, i.e., 0.23 kb closer to the att site
than the data of Otsuka and Abelson (1978) would

105
0 1 2 3 4 kb -- I
attlR V
1 I I II II
bio 267 I
I --
1 I
----A3h---_-_-_-_-_---
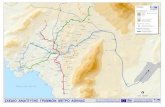
Fig. 1. Physical and genetic map of the regulatory and neighboring regions of the bio operon, showing the endpoints of the bio DNA in
several Xbro phages. The top scale shows the length in kb of bio DNA starting from the aft R site. The open rectangles represent the bio
proteins, positioned as described in the preceding communication of Szybalski and Szybalski (1982). pa is the rightward and j?A is the
leftward bio promoter, and they flank the o operator in the ,?a@*. Symbol ~131 specifies the site of the 1% insertion in Xbiolbio
p98bio ~131, positioned according to the electron microscopic analysis of the heteroduplex Xbiolbio p98bio p131/Xbio+att’imm434
(see Fig. 2). The positions of 5’-ends of the pA and pa-promoted bio transcripts (open arrows) in relation to the ~13 1 insertion site are
taken from Barker et al. (1981). The heavy lines represent bio DNA and the thinner lines (left of artR) correspond to X DNA. The
right endpoint of the A61 deletion in Xgul8(A6l)bio69 and the left endpoint of the A3h deletion in Xbio3h-1 (see also Fig. 2 in
Szybalski and Szybalski, 1982) are positioned in respect to ~131 and all other bio’s. These positions and that of the bio endpoints are
based on the data in column 4(A) of Table I (pl31-bio measurements) and are drawn to scale. For the structure of harr2ir7nn434
(b519b515arrRbiot uorB+pg/-atrLXimm434) see Fiandt et al. (1976) and Szybalski and Szybalski (1982).
seem to indicate. The sequence data of Barker et
al. (1981) and Otsuka and Abelson (1978) specify
the distances between the ~131 insertion site and
the start points of the pA and p,-promoted tran-
scripts, but they do not extend to the att site or all
termini of the bio genes.
ACKNOWLEDGEMENTS
This report was supported by grants 5-POl-CA-
23076 and 5-P30-CA-07175 from the National
Cancer Institute (to W.S.) and by grant AI08573
from the NIAID (to A.C.).

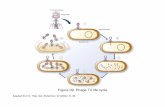
Fig. 2. Electron photomicrograph of the heteroduplcx, Xhiolhw p98bio pl31/b5 196.5 15bio’ (rtr2mnt434. The single-stranded loops A
and B correspond to the b519 and h515 deletions, respectively. loop C to the ~131 (ISf) insertion. and loop D to the combined
&I’- -uurB’ -pgf region of katt’ (see legend to Fig. 1). The I~~nh~mology E is between the immh and wrww434 regions.

TA
BL
E
I
Phys
ical
en
dpoi
nts
of
the
bio
DN
A
inse
rtio
ns
and
dele
tions
h&i0
st
rain
B
(1)
Phys
ical
di
stan
ces
b515
to
bio
end
poin
t b
(%A
*SD
.)
(n)
(2)
pl3
1 to
bi
o en
dpoi
nt
c
(%X
2S.
D.)
(n
)
(3)
Phys
ical
di
stan
ces
Cal
cula
ted
leng
th
of
bio
DN
A
d (k
b)
(A)
(B)
(4)
(5)
h en
dpoi
nt
to
Cal
cula
ted
’ im
m 4
34 e
h
endp
oint
(%;X
*S
D.)
(n
) (%
A)
(6)
(7)
Xbi
o267
nin5
Xbi
ol22
imm
434n
in5
Xbi
o169
imm
434
Xbi
o74i
mm
434
Xbi
olbi
op98
p131
Xga
/( A
61)b
io69
hbio
3h-I
(A3h
)
7.77
~0.5
7 (2
3)
0.26
-cO
.06
(12)
1.
83
1.90
6.
3020
.50
(27)
67
.20
8.27
r0.7
1.
(25)
0.
83*0
.09
(17)
2.
11
2.14
__
_ __
8.58
-to.
78
(44)
1.
07-c
0.08
(27
) 2.
23
2.29
.-
- __
8.52
20.5
3 (4
4)
1.20
20.1
0 (2
2)
2.29
2.
26
__^
_-
9.77
r0.6
6 (4
5) g
2.
3920
.19
(44)
2.
87
2.87
5.
1620
.29
(46)
68
.34
___
1.48
~0.2
1 (1
9)
h 3.
50 i
__
7.14
-tO
.57
(25)
66
.36
-__
0.50
~0.1
0 (2
0) h
-_
__
__
_ __
a Se
e Fi
g.
1 (a
nd
Tab
le
III
of
Szyb
alsk
i an
d Sz
ybal
ski,
1982
; pr
eced
ing
pape
r).
b M
easu
red
in h
eter
odup
lexe
s of
th
e ty
pe
hbio
/Xb5
19b5
15bi
o’ar
t’im
m43
4 as
the
di
stan
ce
from
th
e b5
15 d
elet
ion
loop
to
th
e “a
rt2
loop
”,
the
nonh
omol
ogy
mar
king
th
e bi
o-A
junc
tion
in
the
Xbi
o (s
ee S
zyba
lski
an
d Sz
ybal
ski,
1982
).
S.D
., st
anda
rd
devi
atio
n;
n,
num
ber
of
mol
ecul
es
mea
sure
d (i
n pa
rent
hese
s).
’ M
easu
red
in
hete
rodu
piex
es
of
the
type
X
bio/
Xbi
oibi
o ~
98~
131
as
the
dist
ance
fr
om
the
~131
(I
SI)
in
sert
ion
loop
to
th
e bi
o en
dpoi
nt.
d(A
) C
alcu
late
d in
X
h un
its
as
the
sum
of
th
e di
stan
ce
from
at
f to
~13
1 (3
.52%
&h,
as
mea
sure
d in
th
e hb
rolb
io
p98p
131/
harr
2 he
tero
dupl
ex)
and
the
dist
ance
fr
om
~131
to
th
e bi
o
endp
oint
, an
d co
nver
ted
to
kb.
(B)
Cal
cula
ted
as
the
dist
ance
fr
om
b51S
to
th
e bi
o en
dpoi
nt
min
us
the
bS15
-nrt
R
dist
ance
(i
n SX
),
and
conv
erte
d to
kb
. T
he
b515
-attR
di
stan
ce
is
3.86
kO.4
0 %
X (
Fian
dt
et a
l.,
1976
).
’ D
ista
nce
from
th
e “a
ft*
loop
” to
th
e im
m43
4/im
mX
no
nhom
olog
y in
het
erod
uple
xes
hbio
/XbS
19b5
15bi
o+at
r’im
m43
4.
fCal
cuIa
ted
as
the
diff
eren
ce
betw
een
the
left
im
m43
4 en
dpoi
nt,
take
n as
73
S%h
(Szy
bals
ki
and
Szyb
alsk
i, 19
79)
and
the
h en
dpoi
nt-t
o-im
m43
4 di
stan
ce.
s The
mea
sure
d di
stan
ce
from
ar
rR
to ~
131
(IS
I)
is 3
.522
0.47
8X
(see
fo
otno
te
d).
The
IS
f le
ngth
is
not
in
clud
ed
in
this
b5
1S-L
:o
endp
oint
di
stan
ce.
h D
ista
nce
from
~1
31
left
war
d to
th
e ri
ght
endp
oint
of
de
letio
n A
61
(lin
e 6)
or
ri
ghtw
ard
to
the
left
en
dpoi
nt
of
dele
tion
A3h
(l
ine
7) (
see
Fig.
1)
i Dis
tanc
e fr
om
dele
tion
A61
to
th
e bi
o en
dpoi
nt
in
the
Xga
18(A
61)b
io69
/Xar
r’im
m43
4 he
tero
dupl
ex.

108
REFERENCES
Barker. D.F. and Campbell, A.M.: Use of hro-lac fusion strains
to study regulation of biotin biosynthesis in Escherrchio cob.
J. Bacterial. 143 (1980) 7899800.
Barker, D.F.. Kuhn, J. and Campbell, A.M.: Sequence and
properties of operator mutations in the hro operon of
Escherrchia co/i. Gene I3 (1981) 89- 102.
Fiandt, M.. Gottesman, M.E., Shulman. M.J.. Szybalski, E.H..
Szybalski. W. and Weisberg, R.A.: Physical mapping of
coliphage hart’. Virology 72 (1976) 6- 12.
Kayajanian, G.: Studies on the biotin-transducing, defective
variants of bacteriophage X. Virology 36 (1968) 30-41.
Ketner, G. and Campbell, A.: A deletion placing the galac-
tokinase gene of Eschmchia co/i under control of the biotin
promoter. Proc. Nat]. Acad. Sci. USA 7 I (1974) 2698~~2702.
Ketner. G. and Campbell, A.: Operator and promoter mu-
tations affecting divergent transcription in the hrr~ gene
cluster of Escherrrhrcr CO/I. J. Mol. Biol. I96 (1975) 13-27.
Kotval. J., Campbell, A.. Konopa. G. and Szybalski. W.:
Leftward transcription in the Esc,hrrrchra dt operon does
not require the products of the rightward transcript. Gene
17 (1982) 219-222.
Otsuka, A. and Abelson, J.: The regulatory region of the biottn
operon in Escherichru co/r. Nature 276 (1978) 689-694.
Szybalski. E.H. and Szybalski, W.: A physical map of the
Escherrchm w/i bra operon. Gene I8 ( 1982) 93- 103.
Communicated by M. Lieb.