Site-specific T-DNA integration in Arabidopsis thaliana mediated by the...
Transcript of Site-specific T-DNA integration in Arabidopsis thaliana mediated by the...

Acc
epte
d A
rtic
le
This article has been accepted for publication and undergone full peer review but has not been through the copyediting, typesetting, pagination and proofreading process which may lead to differences between this version and the Version of Record. Please cite this article as an 'Accepted Article', doi: 10.1111/tpj.12202 This article is protected by copyright. All rights reserved.
Received Date : 06-Jun-2012
Revised Date : 04-Apr-2013
Accepted Date : 08-Apr-2013
Article type : Technical Advance
Site-specific T-DNA integration in Arabidopsis thaliana mediated by the combined
action of CRE recombinase and φC31 integrase
Annelies De Paepe, Sylvie De Buck, Jonah Nolf, Els Van Lerberge and Ann Depicker*
Department of Plant Systems Biology, VIB, Technologiepark 927, B-9052 Gent, Belgium and
Department of Plant Biotechnology and Bioinformatics, Ghent University, Technologiepark
927, B-9052 Gent, Belgium
* Correspondence (Tel +32(0)3313940; fax +32(0)3313940; email [email protected]
ugent.be)
E-mail adresses authors:
Annelies De Paepe: [email protected]
Sylvie De Buck: [email protected]
Jonah Nolf: [email protected]
Els Van Lerberge: [email protected]
Ann Depicker: [email protected]
Running title: Site-specific T-DNA integration in Arabidopsis
Keywords: T-DNA, site-specific integration, φC31 integrase, CRE recombinase, gene
stacking, Arabidopsis.
SUMMARY
Random T-DNA integration into the plant host genome can be problematic for a variety of
reasons, including potentially variable transgene expression as a result of different

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
integration positions and multiple T-DNA copies, the risk of mutating the host genome and
the difficulty of stacking well defined traits. Therefore, recombination systems have been
proposed to integrate the T-DNA at a pre-selected site in the host genome. Here, we
demonstrate the capacity of the φC31 integrase (INT) for efficient targeted T-DNA integration.
Moreover, we show that the iterative site-specific integration system (ISSI), which combines
the activities of the CRE recombinase and φC31 integrase, allows to target genes to a pre-
selected site with concomitant removal of the resident selectable marker. To begin, plants
expressing both the CRE and INT recombinase and containing the target attP-site were
constructed. These plants were supertransformed with a T-DNA vector harboring the loxP
site, the attB-sites, a selectable marker and an expression cassette encoding a reporter
protein. Three out of 35 obtained transformants (=9%) showed transgenerational site-
specific integration (SSI) of this T-DNA and removal of the resident selectable marker, as
demonstrated by PCR, Southern blot, and segregation analysis.
In conclusion, our results show the applicability of the ISSI system for precise and
targeted Agrobacterium-mediated integration, allowing the serial integration of transgenic
DNA sequences in plants.
INTRODUCTION
The widely used Agrobacterium plant transformation method allows efficient genetic
modification of many plants species, but still has some drawbacks. Due to a lack of control
on the inserted copy number and the site of integration, the transgenics often harbor multiple
copies of the transgene at one or more random genomic locations (Jones et al., 1987;
Hobbs et al., 1993; Cheng et al., 1997; De Neve et al., 1997; De Buck et al., 1999; De Buck
et al., 2009). Variations in copy number and complex transgene integration events have
been correlated with transgene expression variability and instability caused by post-
transcriptional and transcriptional transgene silencing (Matzke et al., 2000; Muskens et al.,
2000; Wang and Waterhouse, 2000; De Buck et al., 2001). Also, random integration of the
transgenes into the host plant genome can be problematic for a variety of reasons, including

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
potential variable transgene expression resulting from position effects and the risk of
mutating the host genome during transgene integration (Matzke and Matzke, 1998). As a
consequence, often hundreds of transformants must be screened to find an elite event
conferring good stable transgene expression and uncompromised agronomic performance.
Therefore, more precise technologies are highly desirable for the production of
transgenic crop plants with predictable transgene integration sites, and also for addressing
issues in regulatory approval and social acceptance. Furthermore, the challenge for future
crop improvement depends on the ability to stack multiple transgenic traits at a
predetermined integration site in a single-copy fashion. Recycling of marker genes in
successive transformations is therefore also essential as not enough efficient and approved
selectable markers are available.
Site-specific recombination methods have been developed to remove selectable
markers to produce marker-free transgenic plants and to integrate the transgene into an
appropriate genomic target (Wang et al., 2011 and references therein; Tuteja et al., 2012),
as well as to reduce the number of transgene copies (Srivastava and Ow, 2001; De Buck et
al., 2007; De Paepe et al., 2009). The Cre/loxP, R/Rs and FLP/FRT recombinases from
bacteriophage P1 of Escherichia coli (Dale and Ow, 1990; Odell et al., 1990; Russell et al.,
1992), from the SR1 plasmid of Zygosaccharomyces rouxii (Onouchi et al., 1991; Onouchi et
al., 1995) and from the plasmid of Saccharomyces cerevisiae (Lyznik et al., 1993; Kilby et
al., 1995), respectively, have been the focus of multiple studies in plants and other
organisms. These site-specific recombination systems are working efficiently and are simple
to use; however they work bidirectionally, meaning that these recombinases catalyze freely
reversible reactions and are therefore less ideal for gene stacking. Unwanted events such as
deletion of previously introduced DNA cannot be prevented because fully functional
recombination sites are needed for subsequent gene targeting of other DNA fragments. In
contrast, the integrase from the Streptomyces phage φC31 (Thorpe and Smith, 1998), a
member of the serine family of recombinases, catalyzes the recombination reaction of the

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
non-identical attB and attP sites of φC31 in only one direction to generate attL and attR sites.
This feature gives the φC31 system a distinct advantage as a tool for site-specific DNA
integration (Smith et al., 2010). In plants, the φC31 integrase was applied for stable plastid
transformation (Lutz et al., 2004), marker gene excision in tobacco (Ow, 2002;
Kittiwongwattana et al., 2007), removal of transgene fragments in Arabidopsis (Gils et al.,
2008; Thomson et al., 2010), chromosomal recombination in Arabidopsis (Ow, 2002),
intrachromosomal inversion and transgene excision in wheat (Rubtsova et al., 2008; Kempe
et al., 2010), and φC31 integrase-mediated excision in barley (Kapusi et al., 2012). More
recently, the use of the Bxb1 integrase, a recombinase from phage Bxb1 analogous to φC31,
was described for site-specific integration with 10% efficiency in tobacco (Ow, 2011; Yau et
al., 2011) and for site-specific excision in Arabidopsis (Thomson et al., 2012).
The combined use of multiple site-specific recombination systems to direct DNA
integration and concomitant marker gene removal to produce marker-free targeted site-
specific integration events has been proposed and demonstrated in previous studies
(Srivastava and Ow, 2001; Ow, 2002; Akbudak and Srivastava, 2011; Wang et al., 2011;
Nandy and Srivastava, 2012). Furthermore, the combination of a non-reversible with a
reversible recombination system permits efficient sequential gene stacking of many
transgenes.
Brown and co-workers established a method in chicken DT40 cells that enables the
iterative integration of transgenic DNA sequences by combining the site-specific
recombinase activities of the unidirectional φC31 integrase and the bidirectional CRE
recombinase (Dafhnis-Calas et al., 2005). This method, termed iterative site-specific
integration (ISSI), allows to stack genes at a pre-selected site with concomitant removal of
the resident selectable marker. Also another member of the serine recombinase family, the
φBT1 integrase, in combination with the CRE recombinase, was shown to promote targeted
iterative integration of transgenic DNA in vertebrate chromosomes (Xu et al., 2008). Using
this method, four successive cycles of gene stacking were demonstrated in vertebrate cells

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
and five serial integration cycles were described in mammalian cells (Dafhnis-Calas et al.,
2005; Xu et al., 2008).
We investigated the potential of the ISSI strategy as a tool for iterative integration of
transgenic DNA after Agrobacterium-mediated floral dip transformation in Arabidopsis.
Therefore, we developed two T-DNA vectors, i.e. the T-DNA target vector and the T-DNA
targeting vector. The former contains an attP target site for subsequent integration of
targeting T-DNAs and a loxP site for removal of the selectable marker by the CRE
recombinase upon integration of the targeting T-DNA.; the latter contains two directly
oriented loxP sites inside the T-DNA borders needed for circularization; subsequent site-
specific integration is mediated by φC31 between one of its two attB sequences and the
resident attP sequence.
Here, we describe the first cycle of the ISSI strategy in Arabidopsis consisting of site-
specific T-DNA integration (SSI) at the attP target site and excision of the resident marker
construct in a single step. We also demonstrate that site-specific T-DNA integration occurred
in the progeny of 9% of the obtained T-DNA transformants.
RESULTS
Experimental design to demonstrate the feasibility of the ISSI strategy
In the presented ISSI strategy, an Arabidopsis thaliana ecotype C24 transformant was
selected, constitutively expressing the CRE and INT recombinases, and is referred to as the
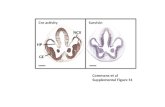
parental transgenic line, Cre-Hsc1 (Figure 1a). This target line contains two T-DNAs, i.e. the
Hsc1 T-DNA and the Cre T-DNA. The Hsc1 T-DNA contains a loxP site (Figure 1a, open
triangle) at the left and an attP (PP’ box) site at the right side of a selectable hygromycin
(Hyg) resistance marker gene. The loxP site allows removal of the selectable marker by the
CRE recombinase upon integration of the targeting T-DNA, while the PP’ site serves as the
target for site-directed integration. Besides this, the Hsc1 T-DNA contains the Pcas driven
φC31 integrase (INT). The Cre-Hsc1 line also harbors the unlinked Cre T-DNA with the

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
biolaphos resistance marker (BAR) and the 35S promoter driven CRE gene expression
cassette.
To demonstrate the targeted stacking of a gene of interest (GOI), a dedicated
targeting vector, pK2L2B (Figure 1b), was constructed. This targeting T-DNA vector contains
two directly oriented loxP sites just between the T-DNA borders to obtain a double-stranded
circular intermediate from the linear T-DNA by CRE-mediated recombination (Figure 1c1). In
between those loxP sites, this targeting vector contains the neomycin phosphotransferase II
(NPTII) selectable marker and a 35S promoter driven enhanced green fluorescent protein/β-
glucuronidase (eGFP/GUS) fusion gene as GOI to stack in the PP’ site in the target line. The
eGFP/GUS gene is flanked by two copies of an attB sequence, BB’I and BB’II (Figure 1b).
The INT-mediated recombination may occur between the attP site and either one of the two
attB sites present in the targeting vector, leading to two different intermediate recombination
products. The INT-produced recombined intermediate I (Figure 1c2) is obtained by INT
recombination between the PP’ and BB’I site (Figure 1c1). CRE-mediated recombination
between the in tandem oriented loxP sites surrounding the resident HPT selectable marker
finally results in a targeted insertion of the T-DNA and concomitant removal of the selectable
marker (Figure 1d). Thus, this SSI transformant can be used for a next round of INT-
mediated site-specific recombination, as it contains a single attB target site. The other
possible INT-produced recombined intermediate II (Figure 1c3), obtained by INT
recombination between PP’ and BB’II, however contains the eGFP/GUS gene together with
the HPT selectable marker between the two loxP sequences, and thus CRE recombination
deletes the transiently inserted GOI, resulting in recombined outcome II as shown in Figure
1e. Half of the INT-mediated recombination products is thus not useful for ISSI and can be
distinguished from recombined outcome I by PCR or Southern blot analysis (see further).
We have illustrated the ISSI strategy in Figure 1 as though the site-specific
integration reaction occurred first by INT after CRE-mediated circularization of the targeting
vector, but ISSI can also be obtained by integration of the targeting T-DNA circle by CRE
into the genomic loxP site of the Hsc1 T-DNA followed by the INT recombination for the

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
excision reaction (Figure S1). However, the order in which the recombinases act does not
make a difference in the final and stabilized outcome.
Construction of the φC31 and CRE recombinase-expressing target line Cre-
Hsc1 with attP target site
Homozygous CRE-expressing Arabidopsis plants (Marjanac et al., 2008) were transformed
with the Hsc1 T-DNA vector (De Paepe et al., 2009) (Figure 1a) by floral dip to obtain a
single-copy Cre-Hsc1 target line suitable for ISSI recombination reactions. The obtained T1
Cre-Hsc1 transformants grew normally and did not show any clear phenotypic alterations,
and single-copy lines were selected by Southern blot analysis (De Paepe et al., 2009). Leaf
extracts of the T1 transformants containing a single Hsc1 T-DNA copy were tested for the
expression of the φC31 integrase by immunoblot analysis with an anti-φC31 integrase
antibody. We selected one single-copy target Cre-Hsc1 transformant with a good φC31
integrase expression. This T1 plant was selfed and seven T2 Hyg-resistant offspring plants
(Table 1- Recipient plants 4, 7, 13, 15, 16, 19 and 23), homozygous (Ho) or hemizygous (He)
for the Hsc1 T-DNA were used for floral dip transformation with the pK2L2B targeting vector.
Creation of SSI events and selection of transformants
The pK2L2B targeting construct as shown in Figure 1b was used for transformation into the
seven Cre-Hsc1 target plants to evaluate how many targeted integration events versus
random insertions would be recovered. In fact, only the former are expected, as in case of
random integration, resolvement of the internal T-DNA segment with the kanamycin (Km)
selectable marker between the loxP sites would leave behind only the LB-loxP-RB footprint
at the integration site. The seeds obtained after floral dip of the T2 Cre-Hsc1 plants with the
pK2L2B targeting construct are indicated as the R1 seed stocks. Selection of R1 seedlings
on Km yielded a total of 35 transformants from about 14,000 seeds, corresponding to an
average transformation frequency of 0.25% (Table 1). In parallel, transformation of the same
vector in CRE-expressing plants without a target attP recombination site generated five Km-
resistant plants out of 4,000 seeds (=0.13%) (Table 1). This shows that pK2L2B

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
transformants can be obtained in which the T-DNA is randomly inserted and not excised by
the CRE recombinase, for instance due to deleted or inaccessible loxP sites. Thus, among
the 35 R1 transformants, a fraction of those is expected to result from random T-DNA
integration concomitant with maintaining the Km selectable marker. Also, C24 wild-type
plants were transformed with the same targeting vector for estimating the random integration
efficiency of the targeting vector. Here, the transformation efficiency was 0.45%.
Molecular analysis of the obtained R1 recombinants
Leaf DNA of the 35 R1 Km-resistant plants was PCR analyzed with primer sets a/b and a/c
(Figure 1d,e). Amplification of a fragment of 478 bp (primers a/b) or 1022 bp (primers a/c)
indicates a targeted recombined outcome I or II, respectively. The diagnostic fragment of
478 bp was found in three and that of 1022 bp in six of the 35 R1 transformants (Table 1).
These nine recombination products were derived from four recipient Cre-Hsc1 plants (4, 7,
13 and 16) while no positive PCR recombination products were seen in the R1 transformants
of the other three Cre-Hsc1 recipient plants, nor in the Km-resistant Cre-1 and C24
transformants. The nine obtained PCR fragments were sequenced and the results confirmed
precise site-specific recombination for recombined outcomes I and II in the leaf DNA of R1
transformants (Table 1). Subsequently, these nine R1 Km-resistant transformants with a
scored SSI event in the leaves were selfed for analysis of the R2 seed stocks.
Segregation analysis to investigate germ line transmission of the SSI event
To determine whether the recombined outcome products were heritably transmitted to the
progeny and whether the resident selectable HPT marker was deleted, we examined the
segregation of the R2 progenies of the nine R1 SSI PCR positive transformants on Km- and
Hyg-containing medium. Surprisingly, several of those seed stocks showed a very low
number of Km-resistant seedlings, as compared to the expected 75% frequency (Table 2).
Also unexpected was that R2 progeny plants of most transformants are still 100% Hyg
resistant. This result indicates that the SSI event demonstrated in the leaves of the R1
transformants was not transmitted through the germ cells and also that the targeting Km-

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
resistant T-DNA was lost from many of the germline precursor cells in several of the R1
transformants. This is probably due to resolvement of the in tandem oriented loxP-sites in
the targeting T-DNA after random integration.
We further reasoned that the SSI event could take place in the germline progenitors
in kanamycin resistant R2 plants. Therefore, some of the Km-resistant R2 plants from each
transformant were selfed and the obtained R3 seed stocks were again analyzed for the
segregation of the targeting Km-resistant and the target Hyg-resistant constructs. As an SSI
T-DNA allele should be correlated with the loss of the Hyg resistance marker at the
integration locus, all R3 SSI progeny plants derived from a hemizygous target plant should
be 100% sensitive to Hyg and 100% or 75% resistant to Km. On the other hand, SSI R3
progeny plants derived from a homozygous Hsc1 locus plant should segregate (3:1) on Hyg
and Km, or should be 100% sensitive to Hyg and 100% resistant to Km.
We found that all R3 progeny plants of the 13-3, 16-11 and 16-12 R2 transformants had
lost Hyg resistance while being 100% Km resistant, or showed Hyg and Km resistance in the
predicted segregation ratio (3:1) (Table 2). In the other tested R3 seed stocks (4-1, 4-2, 7-1,
16-3 and 16-4) the segregation pattern on Hyg and Km was indicative for random integration
of the non-excised T-DNA targeting construct. Indeed, these R3 seed stocks derived from
different R2 transformants were fully Hyg resistant, or showed unlinked segregation of the
Km and Hyg selectable markers.
In conclusion, site-specific integration of a targeting cassette in the attP site with
concomitant removal of the residual selectable marker and stable transmission of the SSI
allele from the R2 to the R3 generation did occur in three of the nine analyzed transformants
(13-3, 16-11 and 16-12).
Molecular confirmation of germ line transmitted SSI events: PCR and Southern
blot analysis
Genomic DNA was isolated for PCR and Southern blot analysis from the Km-resistant
homozygous R3 or R4 (see Table 3) SSI offspring plants of the 16-11, 16-12 and 13-3

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
transformants and from the progeny plants of the 16-4 transformant without an SSI
recombinant product. Amplification with primers a and b showed that all R1 16-11 derived
progenies (Table 3 (5-8)) harbor the recombination event I (Figure 1d), while in none of
these transformants the original attP-PCR fragment could be amplified. Interestingly, from
some of the R1 transformants, the two different recombination outcomes could be recovered.
The a/b primer set amplified the PCR fragment indicative for recombined outcome I in two
out of four analyzed 16-12 progenies and one out of three 13-3 progenies (Table 3
(9,10,15)), while the primer set a/c amplified the fragment diagnostic for recombined
outcome II in the other progenies (Table 3 (11-14)). Primer set a/d amplified the predicted
PCR product for the target attP site in all 16-4 progenies without an SSI event (Table 3 (1-4)).
The seven obtained outcome I PCR fragments with the a/b primer set were also sequenced
and the sequencing results confirmed precise recombinant attL products in the leaf DNA of
these transformants.
In parallel, we performed Southern blot analysis on the same DNA samples. DNA was
digested with either NcoI or SphI (Figure 2a-e). The blots with NcoI-digested DNA were
hybridized with the INT probe and the blots with SphI-digested DNA with the GUS, HPT,
NPTII and INT probe (Figure 1 and 2). The INT probe (probe 2) is predicted to detect a 2.6-
kb NcoI hybridizing fragment in the attP-containing target construct (Figure 1a), while it
should reveal either a 1.8-kb or a 2.8-kb NcoI fragment for recombined outcome I (Figure 1d)
or II (Figure 1e), respectively. The GUS probe should detect a 3.7-kb SphI hybridizing
fragment in plants with outcome I specific for the targeting construct (Figure 1d), while no
signal should be present in plants with recombined outcome II, and one or more T-DNA–
plant junction fragments bigger than 4.1 kb should be detected if random integration of the
targeting construct has occurred (Figure 1b). When using the HPT probe after SphI DNA
digestion, no hybridizing fragments should be detected in the recombinants, while one T-
DNA–plant junction fragment bigger than 2.2 kb should appear in the case of random
integration of the targeting construct (Figure 1a). Furthermore, the NPTII probe should detect
the same hybridizing T-DNA–plant junction fragment bigger than 1.8 kb in recombinants with

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
outcome I and II (Figure 1d,e), while this T-DNA–plant junction fragment should be of a
different size in transformants with random integration of the targeting construct (Figure 1b).
Finally, the same T-DNA–plant junction fragment bigger than 3.3 kb should appear in all
SphI-digested DNA samples when using the INT probe (Figure 1a,d,e).
In agreement with the PCR analysis, Southern blot results confirmed the targeted
integration in 16-11, 16-12 and 13-3 homozygous offspring (Figure 2). Indeed, the expected
hybridization fragments for SSI outcome I and II were detected on the NcoI/INT (Figure 2a)
and SphI/GUS (Figure 2b) blots, indicating that in these lines SSI had occurred at the attP
target site as predicted from the PCR analysis. In the offspring of R1 transformants 16-12
and 13-3, both recombined outcomes I (Fig. 2a lanes 9, 10, and 15) and II (Fig. 2a lanes 11,
12, 13 and 14) were found. As predicted, in offspring with recombined outcome II, a 2.8-kb
NcoI fragment was detected with the INT probe and no 3.7-kb hybridization fragment with
the GUS probe, while the lines with recombined outcome I showed the expected 1.8-kb NcoI
band with the INT probe (Figure 2a) and the 3.7-kb SphI fragment with the GUS probe
(Figure 2b). On the contrary, in the offspring of the non-SSI R1 16-4 transformation event,
the internal attP-containing 2.6-kb NcoI hybridizing fragment was observed with the INT
probe (Figure 2a). In addition, in the R1 16-4 offspring, a new 4.8-kb SphI hybridizing
fragment was seen with the GUS probe, corresponding to a T-DNA–plant junction fragment
(Figure 2b).
An additional weak INT probe hybridizing NcoI fragment of about 5.5 kb was detected in
most of the SSI outcome I recombinants. Similarly, an additional GUS hybridizing SphI
fragment of about 8 kb was seen in all but one of the SSI outcome I recombinants
specifically (Figure 2b). In the R1 13-3 SSI offspring for both outcome I and II recombinants,
multiple extra hybridization bands were observed with the GUS probe probably
corresponding to the integration of additional copies of the targeting vector at the target
locus (Fig. 2b lanes 13,14,15).
Further, the absence of a HPT hybridizing fragment in the offspring of the R1 16-11 and
16-12 transformants and one R1 13-3 progeny plant as expected for recombinant outcome I

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
was confirmed (Fig. 2c, lanes 5-12,15), while in R1 13-3 offspring plants with outcome II, a
HPT hybridizing fragment of about 5 kb was observed (Figure 3c, lanes 13-14). As the
segregation analysis of these 13-3 offspring plants on Hyg-containing medium demonstrated
100% sensitivity towards Hyg (Table 3), we attribute this to a truncated HPT hybridizing
fragment, most likely linked to the extra GUS hybridizing fragments, still present in this
complex SSI locus.
Also, as expected for both recombinant outcome I and II, the same NPTII hybridizing
plant–T-DNA left border fragment was observed. Indeed, an NPT probe hybridizing SphI
fragment of about 6.5 kb was observed in all SSI recombinants (Figure 2, lanes 5-14) except
in the complex R1 13-3 recombinant outcome I (Figure 2, lane 15). The R1 16-4 progeny
plants without an SSI event contained a different NPT hybridizing T-DNA–plant junction
fragment of 4.7 kb (Fig. 2d, lanes 1-4). Also, additional NPTII hybridizing fragments were
seen in the 16-12 recombinants with outcome II and in all the 13-3 recombinant progenies
with both outcomes. This can only be explained by rearrangements during the SSI event at
the left T-DNA end or by a complex targeting T-DNA integration pattern.
Finally, the expected T-DNA–plant junction fragment bigger than 3.3 kb was present in
all SphI-digested DNA samples after hybridization with the INT probe, confirming the single-
copy integration pattern of the target locus and no rearrangements at the right T-DNA plant
junction in transformants with or without an SSI event.
From this molecular data analysis, we can conclude that among the identified SSI
integration events, at least one, ie the progeny plant from R16-12 (Figure 2, lane 9) shows
the exactly predicted desired outcome, namely integration in the PP’ target site and removal
of the resident selectable marker with only one hybridizing fragment with all the probes and
digests tested. The other progenies with recombined outcome I (16-11, 16-12 and 13-3) do
show the correct recombinant PCR fragments, do not have the resident Hyg resistance
marker, show the expected GUS- and INT hybridizing restriction fragments, but do contain in
addition other fragments most likely due to aberrancies in the integration and resolvement
process.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Determination of the presence of LB-loxP-RB footprints
Next, we determined whether the recombinants with a targeted T-DNA integration event
contained footprints of random integration. Indeed, if random T-DNA integration of the
targeting construct occurs and the sequence between the loxP sites is subsequently deleted
by CRE-mediated site-specific recombination, an LB-loxP-RB footprint will remain. To this
end, the primer combination e/f (Figure 1b) was used and was predicted to yield a PCR
fragment of 591 bp in case of CRE-mediated excision of the DNA between the loxP sites in
the correctly processed T-DNA from the targeting vector. For most of the homozygous
offspring lines of R1 16-11, 16-12 and 13-3 transformants amplification of this specific
fragment (Table 3) was obtained, indicating that LB-loxP-RB footprints were present. Only in
one of the R1 16-12 progenies this footprint segregated from the targeted SSI locus.
Furthermore, also the progenies from R1 transformants with random integration events of
the targeting construct (such as those from 16-4) contained the LB-loxP-RB footprint,
indicating that one or more targeting T-DNA insertions did occur from which the internal T-
DNA was deleted.
DISCUSSION
For the insertion of new DNA at a known locus, both recombinase-mediated and
homologous recombination-dependent site-specific DNA targeting have been demonstrated
to work. For homologous recombination (HR), targeting frequencies of 10-3-10-6 have been
reported in plants, being still quite inefficient (Britt and May, 2003; Hanin and Paszkowski,
2003; Iida and Terada, 2004). Studies have shown that double stranded breaks (DSBs)
induced by site-specific edonucleases can stimulate the HR repair machinery and thus gene
targeting (Puchta, 2005). Also, in planta gene targeting was achieved in Arabidopsis with a
frequency up to 8.3 x 10-3 by crossing an endonuclease expressing locus with a target and a
transgenic donor locus both containing the corresponding restriction sites in homologous
sequences, resulting in recombination between the activated target site and both ends of the

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
released donor construct (Fauser et al., 2012). Recently, sequence-specific nucleases
introducing double-stranded breaks such as zinc-finger nucleases (Shukla et al., 2009;
Townsend et al., 2009; Tzfira et al., 2012), TALEN nucleases (Christian et al., 2010; Tzfira et
al., 2012; Zhang et al., 2013) and mega-nucleases (Yang et al., 2009; Gao et al., 2010;
Tzfira et al., 2012) were shown to all target DNA integration to those double-stranded breaks
with reproducible frequencies. On the other hand, site-specific recombination systems have
been described to allow gene targeting (Wang et al., 2011 and references therein).
Compared to homology dependent gene replacement, which involves end-joining ligation,
site-specific recombinases are precise, but need pre-existing recombination sites in the DNA.
Here, we demonstrate that the combined activities of the unidirectional φC31 integrase
and the reversible CRE recombinase can integrate a T-DNA at a predetermined position in
the Arabidopsis genome upon Agrobacterium-mediated floral dip transformation with
concomitant removal of the resident selectable marker. Indeed, one cycle of ISSI of an
Agrobacterium-delivered T-DNA is shown to have occurred by PCR, Southern blot, and
segregation analyses in the offspring of three out of 35 selected T-DNA transformants.
Fragments of the expected sizes were detected on Southern blots and the selectable marker
from the target locus was lost in the homozygous SSI recombinants. In addition, the
identified integration events with outcome I are Km-resistant, and contain the added GOI
(GFP/GUS) in the target locus.
From the results, it is clear that random insertion of the targeting T-DNA did happen in
most, if not all, of the transformed female gametocytes, for which an R1 Km resistant
seedling could be selected. First of all, most of the obtained R1 transformants lost the Km
selectable marker during the first generation. This is probably the result of CRE mediated
excision of random integrated T-DNA copies, generating a circular extra-chromosomal
product which then gets degraded or lost upon cell division. Indeed, the nine R1
transformants with a PCR proven SSI event in the leaves did not transmit the Km resistance
as predicted, and only few R2 seedlings were still Km resistant. Also, more than half of the
other 26 R1 Km-selected transformants without SSI-specific PCR fragments in the leaves

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
showed loss of the Km resistance in the R2 generation. Furthermore, LB-loxP-RB footprints
were found in most of the transformants and also in the offspring of the three SSI site-
specific recombinants. This was also observed by Vergunst et al. (1998). These footprints
should anyway be removed from the final recombinant, which can be easily done by
segregation or through back-crosses, because the RB-loxP-LB footprints could possibly
interact with other loxP sites in the genome (SSI locus or others).
Altogether, the results suggest that the CRE recombinase activity was too low in the
transformed female gametocyte to efficiently obtain targeted integration. Indeed, it was
already observed before that the CRE activity was lower in germinal cells compared to
somatic cells when expressed from a 35S driven expression cassette (Marjanac et al.,
2008). Thus, to succeed optimally in the proposed ISSI set-up and to drive targeted
integration immediately after T-DNA transfer, one needs to have both the CRE recombinase
and the INT integrase simultaneously working well in the transformed female gametocyte. In
particular the CRE recombinase needs to be very active to generate the circular T-DNA
product and to prevent random insertion. Therefore, adequate female gametocyte specific
promoters could be used as described by Van Ex et al. (2009).
Further, analysis of the R2 and R3/R4 progenies of selected R1 transformants with an
SSI event in the leaves led to the following insights. The randomly integrated targeting T-
DNA construct in R2 plants could serve as donor locus for release of CRE-mediated excised
circles which were then either inserted stably in the attP target locus or got lost. Probably
site-specific integration did occur independently in many different somatic cells. This
explains why several of the R1 transformants with an SSI-specific PCR fragment amplified
from R1 leaf material, did not transmit this SSI event to the R2 and R3 generation.
Srivastava and Ow (2003) described the rare instance of the maintenance of the CRE-
mediated deletion product as an extra-chromosomal circular molecule in mature somatic
tissue. In our experiments the CRE gene was expressed by the constitutive 35S promotor,
which is active at all stages of development. Somatic accumulation of CRE-mediated
excision circles could also have occurred in our experiments yielding donor molecules for

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
SSI into the target locus of somatic cells. Also Arumugam et al. (2007) found that CRE-
mediated excisions occurred with very high efficiency in quiescent, differentiated somatic
tissues, but were less abundant in meristematic tissues.
Only if the SSI occurs in the primordial cells of the germ line it is stable and transferred
to the next generation. This can occur several times during development. Indeed, several R2
seedlings selected to still contain the Km resistant targeting construct, acquired independent
SSI events during development of the primordial germ cells, as both SSI recombined
outcome I and II events were found in the R3 generation. We postulate that the excised
targeting construct recombined in different attP sites in different germline primordial cells,
resulting in offspring with two different SSI alleles.
Finally, in some of the selected R1 transformants, the randomly integrated targeting T-
DNA remained stably present in the genome, and CRE-mediated loss of the Km resistant
marker did not occur. This can be explained by either the absence of one of the loxP sites
due to truncation of the T-DNA ends, or to mutated or inaccessible loxP sites. Previous
reports also documented that CRE-mediated excision can be suppressed in a significant
amount of transformants because of deleted, mutated or non-accessible loxP sequences
due to chromatin modifications in complex loci (De Buck et al., 2007; Marjanac et al., 2008;
De Paepe et al., 2009).
For certain applications, one would eventually like to obtain CRE- and INT-free plants
containing the stacked insert locus. The most straightforward approach is to flank the CRE
and INT expression cassettes with directly oriented site-specific recombination sites other
than loxP sites (f.e. FRT or RS sites), so that subsequent introduction of the corresponding
recombinase by co-transformation, crossing or re-transformation would excise the CRE and
INT gene expression cassettes. Moreover, the selectable marker could also be excised in
this manner to obtain a final marker-free T-DNA integration event. Also, the incorporation of
a promoter-trap target construct, where the loxP site is placed between the promoter and the
coding region of the selectable marker in the target construct, and the use of a targeting
vector with a loxP site at the promoterless 5’ end of the coding region of a second selectable

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
marker (Dafhnis-Calas et al., 2005; Wang et al., 2011 and references therein) may aid in
obtaining only precise SSI lines because then transformants with random T-DNA events will
not survive on selective growth medium. Further, optimization of the promoters of both
recombinases could greatly enhance the targeting frequency at the moment of the
transformation/integration event itself. In addition, additional single-copy target lines could be
compared in a follow-up study to select the best performing target line and to obtain a clear
view of the effciency of the proposed SSI method.
The ISSI described here opens new ways for the development of transgenics with
single-copy and stable multigene inserts at one genetic locus. The SSI integrated
recombined product (recombined outcome I) can now be the target for stacking another
sequence by site-specific recombination (Figure S2) and the ISSI cycle can be repeated for
an arbitrary number of times with only two selectable markers and two sets of recombination
sites, namely the loxP and the attB/attP sites. This is in contrast to the stacking possibilities
of the recombinase-mediated DNA cassette exchange (RMCE) method (Li et al., 2009)
which is dependent on the availability of compatible recognition sites. The possibility to stack
multiple transgenes at a single locus should be very useful in the near future. A merit of
assembling transgene arrays is that it will avoid the difficulties arising from segregation of
independent transgenic loci. In addition, stacking of multiple transgenes at a predicted site of
integration ensures single copy insertions and allows control of the position in the genome.
EXPERIMENTAL PROCEDURES
Plasmid construction: pHsc1, pCre and pK2L2B vectors
To produce the targeting construct pK2L2B, compatible oligonucleotides (5’-
GGCCCGTTAACATAACTTCGTATAGCAT-3’ and 5’-ACATTATACGAAGTTATCCATATTA-
3’) containing the loxP sequence were first ligated and inserted into the PspOMI/BstXI site of
pXD610 (De Loose et al., 1995). Then, compatible oligonucleotides (5’-
AATTCAGATCTATAACTTCGTATA-3’ and 5’-ATGTATGCTATACGAAGTTATG-3’)

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
containing the second loxP sequence in the same orientation were ligated and inserted into
the EcoRI/BamHI site. Next, compatible oligonucleotides (5’-
CGTTAACCGGTGCGGGTGCCAGGGCGTGCCC-3’ and 5’-
TTGGGCTCCCCGGGCGCGTACTCCACTCTAGAGAGCT-3’) containing the attB sequence
were ligated and inserted into the SacI/HpaI site of the p35S-Fs7 vector, which contains the
P35S::eGFPGUS expression cassette, followed by insertion of compatible oligonucleotides
(5’-GGCCCTCTAGACGGTGCGGGTGCCAGGGCGTGCCC-3’ and 5’-
TTGGGCTCCCCGGGCGCGTACTCCACCATATG-3’) containing the second attB site into
the PspOMI site. Finally, the XbaI-cut attB-GUS/GFP-attB fragment was ligated to the XbaI-
cut loxP-containing fragment to yield the pK2L2B vector. Plasmid pHsc1, which harbors the
φC31 integrase coding cassette (De Paepe et al., 2009), and the CRE T-DNA were
described previously (De Buck et al., 2000; Marjanac et al., 2008; De Paepe et al., 2009).
Agrobacterium strain, plant transformation and selection of transformants
The pHsc1 and pK2L2B plasmids were transferred to the Agrobacterium tumefaciens strain
C58C1RifR(pMP90) by electroporation. Homozygous single-copy Cre1 (in C24 background)
(Marjanac et al., 2008) plants were transformed with the pHsc1 vector by the floral-dip
method (Clough and Bent, 1998). Seeds of the dipped plants were harvested and sown on
germination medium supplemented with Hyg (20 mg/L) for selection of primary
transformants (T1). A single-copy transformant, Cre-Hsc1, was chosen as target for
supertransformation. T2 progeny plants of the Cre-Hsc1 plant (homozygous or hemizygous
for the Hsc1 locus), the homozygous Cre1 plant (Marjanac et al., 2008) and C24 wild-type
plants were transformed with the pK2L2B T-DNA vector. Selection of the R1 seedling
transformants was performed on Km (50 mg/L). After growing and selfing these plants, R2
seed stocks were obtained and the zygosity of the Km locus and the presence of Hyg
resistance was determined in the R2 and R3 progeny by sowing at least 50 seeds on either
Km- or Hyg-containing medium (20 mg/L).
Plant DNA preparation and Southern blot analysis

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
DNA of Arabidopsis leaf material was prepared from 200 mg frozen plant tissue using the
DNeasy Plant Mini Kit as described by the manufacturer (Qiagen, http://www.qiagen.com/).
Digested Arabidopsis DNA (1 µg) was loaded into each lane of a 1% agarose gel. For the
Hsc1 T-DNA, the HPT (Figure 1a, probe 1) and the INT (Figure 1a, probe 2) probes were
used. The HPT and INT probes were prepared via PCR amplification from the plasmids
pH610 (De Buck et al., 1998) and pGEMINT (De Paepe et al., 2009), respectively, with the
following primers: for the HPT probe with hpt1 (5’- ACCTGCCTGAAACCGAACTGC-3’) and
hpt2 (5’-GCGTCTGCTGCTCCATACAA-3’) and for the INT probe with anpae32 (5’-
ACGCCTGAAGCTCATACCAC-3’) and anpae33 (5’- GGAAACGTCATGGACCTGAT-3’). For
the K2L2B T-DNA, the NPTII (Figure 1b, probe 3) and the GUS (Figure 1b, probe 4) probes
were used. The NPTII probe was PCR amplified from the pK2L610 plasmid (De Buck et al.,
1998) with the primers napod8 (5’- GCTTTTCTGGATTCATCGACTGTG-3’) and napod13
(5’- CGCTATAAACCTATTCAGCACA-3’), and the GUS probe was amplified from the
pGEMgus plasmid (De Buck et al., 1999) with the primers GUS3F (5’-
TGGCAGTGAAGGGCCAACAG-3’) and GUS5R (5’- TACCTGTTCACCGACGACGG-3’).
The probe DNA was labeled and detected using the PCR DIG probe synthesis mix (Roche,
Mannheim, Germany), Anti-digoxigenin (DIG)-AP Fab fragments and the CDP-star detection
module (Roche) (De Paepe et al., 2012).
PCR analysis
For PCR analysis, 100 ng of leaf DNA was used. Primers were: a, 5’-
CGCCTGGTTCTTCTTTTTCA-3’; b, 5’-TAACCATCTGTGGGTCAGCA-3’; c, 5’-
TCAGTGACAACGTCGAGCACA-3’; d, 5’-CGGAATTGACGGATCTGGTA-3’; e, 5’-
CAGGCCAAATTCGCTCTTAG-3’; f, 5’- CCTGGCCTAACATCACACCT-3’. The PCR
reaction conditions were: initial denaturation at 94°C for 5 min, followed by 30 cycles of
denaturation at 94°C for 45 sec, annealing at 56°C for 45 sec, and elongation at 72°C for 60
sec, with a final elongation step at 72°C for 5 min. 10 µL of the PCR samples was loaded on
1% agarose gels and visualized by addition of EtBr.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
ACKNOWLEDGEMENTS
The authors thank William Brown (Institute of Genetics, University of Nottingham, UK) for
providing the plasmid with the φC31= integrase gene, our colleagues of the floral dip and
sequencing groups, Marta Walczak for help with experimental work, and Annick Bleys for
help in preparing the manuscript. This work was supported by grants from the 6th
Framework Program of the European Union ‘GENINTEG’ (LSHG-CT2003-503303), the
European Union BIOTECH program (QLRT-2000-00078), with additional co-financing from
the Flemish Community and the Research Foundation Flanders (G.021106).
CONFLICTS OF INTEREST
No conflicts of interest are to be declared.
SUPPORTING INFORMATION
Additional Supporting Information may be found in the online version of this article:
Figure S1. Schematic outline of the alternative ISSI process when using the pK2L2B vector
for site-specific recombination into the Cre-Hsc1 target line.
Figure S2. ISSI T-DNA stacking strategy.
REFERENCES
Akbudak, M.A. and Srivastava, V. (2011) Improved FLP recombinase, FLPe, efficiently
removes marker gene from transgene locus developed by Cre-lox mediated site-specific
gene integration in rice. Mol. Biotechnol., 49, 82-89.
Arumugam, N., Gupta, V., Jagannath, A., Mukhopadhyay, A., Pradhan, A.K., Burma,
P.K. and Pental, D. (2007) A passage through in vitro culture leads to efficient production

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
of marker-free transgenic plants in Brassica juncea using the Cre-loxP system.
Transgenic Res., 16, 703-712.
Britt, A.B. and May, G.D. (2003) Re-engineering plant gene targeting. Trends Plant Sci., 8,
90-95.
Cheng, M., Fry, J.E., Pang, S., Zhou, H., Hironaka, C.M., Duncan, D.R., Conner, T.W.
and Wan, Y. (1997) Genetic transformation of wheat mediated by Agrobacterium
tumefaciens. Plant Physiol., 115, 971-980.
Christian, M., Cermak, T., Doyle, E.L., Schmidt, C., Zhang, F., Hummel, A., Bogdanove,
A.J. and Voytas, D.F. (2010) Targeting DNA double-strand breaks with TAL effector
nucleases. Genetics, 186, 757-761.
Clough, S.J. and Bent, A.F. (1998) Floral dip: a simplified method for Agrobacterium-
mediated transformation of Arabidopsis thaliana. Plant J., 16, 735-743.
Dafhnis-Calas, F., Xu, Z., Haines, S., Malla, S.K., Smith, M.C.M. and Brown, W.R.A.
(2005) Iterative in vivo assembly of large and complex transgenes by combining the
activities of φC31 integrase and Cre recombinase. Nucleic Acids Res., 33, e189.
Dale, E.C. and Ow, D.W. (1990) Intra- and intermolecular site-specific recombination in
plant cells mediated by bacteriophage P1 recombinase. Gene, 91, 79-85.
De Buck, S., Jacobs, A., Van Montagu, M. and Depicker, A. (1998) Agrobacterium
tumefaciens transformation and cotransformation frequencies of Arabidopsis thaliana root
explants and tobacco protoplasts. Mol. Plant-Microbe Interact., 11, 449-457.
De Buck, S., Jacobs, A., Van Montagu, M. and Depicker, A. (1999) The DNA sequences
of T-DNA junctions suggest that complex T-DNA loci are formed by a recombination
process resembling T-DNA integration. Plant J., 20, 295-304.
De Buck, S., De Wilde, C., Van Montagu, M. and Depicker, A. (2000) Determination of the
T-DNA transfer and the T-DNA integration frequencies upon cocultivation of Arabidopsis
thaliana root explants. Mol. Plant-Microbe Interact., 13, 658-665.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
De Buck, S., Van Montagu, M. and Depicker, A. (2001) Transgene silencing of invertedly
repeated transgenes is released upon deletion of one of the transgenes involved. Plant
Mol. Biol., 46, 433-445.
De Buck, S., Peck, I., De Wilde, C., Marjanac, G., Nolf, J., De Paepe, A. and Depicker, A.
(2007) Generation of single-copy T-DNA transformants in Arabidopsis by the CRE/loxP
recombination-mediated resolution system. Plant Physiol., 145, 1171-1182.
De Buck, S., Podevin, N., Nolf, J., Jacobs, A. and Depicker, A. (2009) The T-DNA
integration pattern in Arabidopsis transformants is highly determined by the transformed
target cell. Plant J., 60, 134-145.
De Loose, M., Danthinne, X., Van Bockstaele, E., Van Montagu, M. and Depicker, A.
(1995) Different 5’ leader sequences modulate β-glucuronidase accumulation levels in
transgenic Nicotiana tabacum plants. Euphytica, 85, 209-216.
De Neve, M., De Buck, S., Jacobs, A., Van Montagu, M. and Depicker, A. (1997) T-DNA
integration patterns in co-transformed plant cells suggest that T-DNA repeats originate
from co-integration of separate T-DNAs. Plant J., 11, 15-29.
De Paepe, A., De Buck, S., Hoorelbeke, K., Nolf, J., Peck, I. and Depicker, A. (2009)
High frequency of single-copy T-DNA transformants produced by floral dip in CRE-
expressing Arabidopsis plants. Plant J., 59, 517-527.
De Paepe, A., De Buck, S., Nolf, J. and Depicker, A. (2012) High frequency of single-copy
T-DNA transformants produced after floral dip in CRE-expressing Arabidopsis plants. In
Transgenic Plants: Methods and Protocols (Dunwell, J.M. and Wetten, A.C. eds.) New
York: Humana Press, pp. 317-333.
Fauser, F., Roth, N., Pacher, M., Ilg, G., Sánchez-Fernández, R., Biesgen, C. and
Puchta, H. (2012) In planta gene targeting. Proc. Natl. Acad. Sci. USA, 109, 7535-7540.
Gao, H., Smith, J., Yang, M., Jones, S., Djukanovic, V., Nicholson, M.G., West, A.,
Bidney, D., Falco, S.C., Jantz, D. and Lyznik, L.A. (2010) Heritable targeted
mutagenesis in maize using a designed endonuclease. Plant J., 61, 176-187.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Gils, M., Marillonnet, S., Werner, S., Grützner, R., Giritch, A., Engler, C.,
Schachschneider, R., Klimyuk, V. and Gleba, Y. (2008) A novel hybrid seed system for
plants. Plant Biotechnol. J., 6, 226-235.
Hanin, M. and Paszkowski, J. (2003) Plant genome modification by homologous
recombination. Curr. Opin. Plant Biol., 6, 157-162.
Hobbs, S.L.A., Warkentin, T.D. and DeLong, C.M.O. (1993) Transgene copy number can
be positively or negatively associated with transgene expression. Plant Mol. Biol., 21, 17-
26.
Iida, S. and Terada, R. (2004) A tale of two integrations, transgene and T-DNA: gene
targeting by homologous recombination in rice. Curr. Opin. Biotechnol., 15, 132-138.
Jones, J.D.G., Gilbert, D.E., Grady, K.L. and Jorgensen, R.A. (1987) T-DNA structure
and gene expression in petunia plants transformed by Agrobacterium tumefaciens C58
derivates. Mol. Gen. Genet., 207, 478-485.
Kapusi, E., Kempe, K., Rubtsova, M., Kumlehn, J. and Gils, M. (2012) PhiC31 integrase-
mediated site-specific recombination in barley. PLoS One, 7, e45353.
Kempe, K., Rubtsova, M., Berger, C., Kumlehn, J., Schollmeier, C. and Gils, M. (2010)
Transgene excision from wheat chromosomes by phage phiC31 integrase. Plant Mol.
Biol., 72, 673-687.
Kilby, N.J., Davies, G.J., Snaith, M.R. and Murray, J.A.H. (1995) FLP recombinase in
transgenic plants: constitutive activity in stably transformed tobacco and generation of
marked cell clones in Arabidopsis. Plant J., 8, 637-652.
Kittiwongwattana, C., Lutz, K., Clark, M. and Maliga, P. (2007) Plastid marker gene
excision by the phiC31 phage site-specific recombinase. Plant Mol. Biol., 64, 137-143.
Li, Z., Xing, A., Moon, B.P., McCardell, R.P., Mills, K. and Falco, S.C. (2009) Site-specific
integration of transgenes in soybean via recombinase-mediated DNA cassette exchange.
Plant Physiol., 151, 1087-1095.
Lutz, K.A., Corneille, S., Azhagiri, A.K., Svab, Z. and Maliga, P. (2004) A novel approach
to plastid transformation utilizes the phiC31 phage integrase. Plant J., 37, 906-913.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Lyznik, L.A., Mitchell, J.C., Hirayama, L. and Hodges, T.K. (1993) Activity of yeast FLP
recombinase in maize and rice protoplasts. Nucleic Acids Res., 21, 969-975.
Marjanac, G., De Paepe, A., Peck, I., Jacobs, A., De Buck, S. and Depicker, A. (2008)
Evaluation of CRE-mediated excision approaches in Arabidopsis thaliana. Transgenic
Res., 17, 239-250.
Matzke, A.J.M. and Matzke, M.A. (1998) Position effects and epigenetic silencing of plant
transgenes. Curr. Opin. Plant Biol., 1, 142-148.
Matzke, M.A., Mette, M.F. and Matzke, A.J.M. (2000) Transgene silencing by the host
genome defense: implications for the evolution of epigenetic control mechanisms in plants
and vertebrates. Plant Mol. Biol., 43, 401-415.
Muskens, M.W.M., Vissers, A.P.A., Mol, J.N.M. and Kooter, J.M. (2000) Role of inverted
DNA repeats in transcriptional and post-transcriptional gene silencing. Plant Mol. Biol., 43,
243-260.
Nandy, S. and Srivastava, V. (2012) Marker-free site-specific gene integration in rice based
on the use of two recombination systems. Plant Biotechnol. J., 10, 904-912.
Odell, J., Caimi, P., Sauer, B. and Russell, S. (1990) Site-directed recombination in the
genome of transgenic tobacco. Mol. Gen. Genet., 223, 369-378.
Onouchi, H., Yokoi, K., Machida, C., Matsuzaki, H., Oshima, Y., Matsuoka, K.,
Nakamura, K. and Machida, Y. (1991) Operation of an efficient site-specific
recombination system of Zygosaccharomyces rouxii in tobacco cells. Nucleic Acids Res.,
19, 6373-6378.
Onouchi, H., Nishihama, R., Kudo, M., Machida, Y. and Machida, C. (1995) Visualization
of site-specific recombination catalyzed by a recombinase from Zygosaccharomyces
rouxii in Arabidopsis thaliana. Mol. Gen. Genet., 247, 653-660.
Ow, D.W. (2002) Recombinase-directed plant transformation for the post-genomic era. Plant
Mol. Biol., 48, 183-200.
Ow, D.W. (2011) Recombinase-mediated gene stacking as a transformation operating
system. J. Integr. Plant Biol., 53, 512-519.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Puchta, H. (2005) The repair of double-strand breaks in plants: mechanisms and
consequences for genome evolution. J. Exp. Bot., 56, 1-14.
Rubtsova, M., Kempe, K., Gils, A., Ismagul, A., Weyen, J. and Gils, M. (2008) Expression
of active Streptomyces phage phiC31 integrase in transgenic wheat plants. Plant Cell
Rep., 27, 1821-1831.
Russell, S.H., Hoopes, J.L. and Odell, J.T. (1992) Directed excision of a transgene from
the plant genome. Mol. Gen. Genet., 234, 49-59.
Shukla, V.K., Doyon, Y., Miller, J.C., DeKelver, R.C., Moehle, E.A., Worden, S.E.,
Mitchell, J.C., Arnold, N.L., Gopalan, S., Meng, X., Choi, V.M., Rock, J.M., Wu, Y.-Y.,
Katibah, G.E., Zhifang, G., McCaskill, D., Simpson, M.A., Blakeslee, B., Greenwalt,
S.A., Butler, H.J., Hinkley, S.J., Zhang, L., Rebar, E.J., Gregory, P.D. and Urnov, F.D.
(2009) Precise genome modification in the crop species Zea mays using zinc-finger
nucleases. Nature, 459, 437-441.
Smith, M.C.M., Brown, W.R.A., McEwan, A.R. and Rowley, P.A. (2010) Site-specific
recombination by φC31 integrase and other large serine recombinases. Biochem. Soc.
Trans., 38, 388-394.
Srivastava, V. and Ow, D.W. (2001) Single-copy primary transformants of maize obtained
through the co-introduction of a recombinase-expressing construct. Plant Mol. Biol., 46,
561-566.
Srivastava, V. and Ow, D.W. (2003) Rare instances of Cre-mediated deletion product
maintained in transgenic wheat. Plant Mol. Biol., 52, 661-668.
Thomson, J.G., Chan, R., Thilmony, R., Yau, Y.-Y. and Ow, D.W. (2010) PhiC31
recombination system demonstrates heritable germinal transmission of site-specific
excision from the Arabidopsis genome. BMC Biotechnol., 10, 17.
Thomson, J.G., Chan, R., Smith, J., Thilmony, R., Yau, Y.-Y., Wang, Y. and Ow, D.W.
(2012) The Bxb1 recombination system demonstrates heritable transmission of site-
specific excision in Arabidopsis. BMC Biotechnol., 12, 9.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Thorpe, H.M. and Smith, M.C.M. (1998) In vitro site-specific integration of bacteriophage
DNA catalyzed by a recombinase of the resolvase/invertase family. Proc. Natl. Acad. Sci.
USA, 95, 5505-5510.
Townsend, J.A., Wright, D.A., Winfrey, R.J., Fu, F., Maeder, M.L., Joung, J.K. and
Voytas, D.F. (2009) High-frequency modification of plant genes using engineered zinc-
finger nucleases. Nature, 459, 442-445.
Tuteja, N., Verma, S., Sahoo, R.K., Raveendar, S. and Reddy, I.N.B.L. (2012) Recent
advances in development of marker-free transgenic plants: Regulation and biosafety
concern. J. Biosci., 37, 167-197.
Tzfira, T., Weinthal, D., Marton, I., Zeevi, V., Zuker, A. and Vainstein, A. (2012) Genome
modifications in plant cells by custom-made restriction enzymes. Plant Biotechnol. J., 10,
373-389.
Van Ex, F., Verweire, D., Claeys, M., Depicker, A. and Angenon, G. (2009) Evaluation of
seven promoters to achieve germline directed Cre-lox recombination in Arabidopsis
thaliana. Plant Cell Rep., 28, 1509-1520.
Verdaguer, B., de Kochko, A., Fux, C.I., Beachy, R.N. and Fauquet, C. (1998) Functional
organization of the cassava vein mosaic virus (CsVMV) promoter. Plant Mol. Biol., 37,
1055-1067.
Vergunst, A.C., Jansen, L.E.T. and Hooykaas, P.J.J. (1998) Site-specific integration of
Agrobacterium T-DNA in Arabidopsis thaliana mediated by Cre recombinase. Nucleic
Acids Res., 26, 2729-2734.
Wang, M.-B. and Waterhouse, P.M. (2000) High-efficiency silencing of a β-glucuronidase
gene in rice is correlated with repetitive transgene structure but is independent of DNA
methylation. Plant Mol. Biol., 43, 67-82.
Wang, Y., Yau, Y.-Y., Perkins-Balding, D. and Thomson, J.G. (2011) Recombinase
technology: applications and possibilities. Plant Cell Rep., 30, 267-285.
Xu, Z., Lee, N.C.O., Dafhnis-Calas, F., Malla, S., Smith, M.C.M. and Brown, W.R.A. (2008)
Site-specific recombination in Schizosaccharomyces pombe and systematic assembly of

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
a 400kb transgene array in mammalian cells using the integrase of Streptomyces phage
φBT1. Nucleic Acids Res., 36, e9.
Yang, M., Djukanovic, V., Stagg, J., Lenderts, B., Bidney, D., Falco, S.C. and Lyznik,
L.A. (2009) Targeted mutagenesis in the progeny of maize transgenic plants. Plant Mol.
Biol., 70, 669-679.
Yau, Y.-Y., Wang, Y., Thomson, J.G. and Ow, D.W. (2011) Method for Bxb1-mediated site-
specific integration in planta. In Plant Chromosome Engineering (Birchler, J.A. ed.) New
York: Springer, pp. 147-166.
Zhang, Y., Zhang, F., Li, X., Baller, J.A., Qi, Y., Starker, C.G., Bogdanove, A.J. and
Voytas, D.F. (2013) Transcription activator-like effector nucleases enable efficient plant
genome engineering. Plant Physiol., 161, 20-27.
Tables
Table 1: Transformation efficiency of the targeting T-DNA construct and PCR analysis of R1
transformants for site-specific recombination
Recipient T2
Cre-Hsc1
planta
# of sown
seeds/
plant line
# of
Kmr R1
transfor-
mants
Transformation/
recombination
efficiency (TE)
(%)b
Positive R1
PCR for
recombined
outcome Ic
Positive R1
PCR for
recombined
outcome IId
R1 plant with
positive SSI
PCR product
4 2000 7 0.35 0 2 4-1
4-2
7 2000 4 0.2 0 1 7-1
13 2000 3 0.15 0 2 13-2
13-3
15 2000 6 0.3 0 0
16 4000 12 0.3 3 1
16-3
16-4
16-11
16-12

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
19 1000 0 0 0 0
23 1000 3 0.3 0 0
Total 14000 35 0.25 3 6
Cre1 4000 5 0.13 0 0
C24 4000 18 0.45 0 0
Kmr, kanamycin resistant
a Seven T2 progeny plants, Ho or He for the Hsc1 locus (Hyg resistant), from a single T1
transformant being Ho for the CRE locus, were numbered and used for floral dipping with the
pK2L2B construct.
bTransformation/recombination efficiency: the # of transformants divided by #
seeds/plant*100.
c PCR reaction with primers a and b yields the 478-bp fragment if the recombined outcome I
is present in leaf DNA of the R1 plant (see Figure 1d).
d PCR reaction with primers a and c yields the 1022-bp fragment if the recombined outcome
II is present in leaf DNA of the R1 plant (see Figure 1e).

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Table 2: Segregation analysis of the targeting construct containing the Kmr marker and the
target locus containing the Hygr marker in the nine selfed progenies of R1 transformants
positive for the SSI-specific PCR
tested R2 #Kmr/total #Hygr/total tested R3 #Kmr/total #Hygr/total
seed stock R2 R2 seed stock R3 R3
4-1 5/60 58/58 4-1-1 34/50 50/50
4-1-2 37/50 50/50
4-1-3 34/50 50/50
4-1-4 40/52 50/50
4-2 26/60 49/60 4-2-1 46/60 0/58
4-2-2 42/57 55/55
4-2-3 45/60 54/54
4-2-4 40/58 33/57
4-2-5 42/57 51/51
7-1 48/60 46/60 7-1-1 40/59 58/58
7-1-2 41/60 60/60
13-2 0/58 46/60
13-3 30/60 9/60 13-3-1 49/57 0/58
13-3-2 41/59 0/59
13-3-3 47/59 0/58
13-3-6 46/59 0/60
13-3-7 46/60 0/59
16-3 36/58 59/59 16-3-1 39/55 58/58
16-3-2 28/52 58/58
16-3-3 35/56 58/58
16-3-4 37/50 58/58
16-3-5 23/54 58/58

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
16-4 5/60 60/60 16-4-1 44/56 58/58
16-4-2 50/50 52/52
16-4-3 58/58 58/58
16-11 3/78 60/60 16-11-1 42/54 33/53
16-11-2 58/58 0/58
16-11-3 58/58 0/58
16-12 2/60 59/59 16-12-1 21/59* 38/58
16-12-2 7/60* 38/56
Kmr, kanamycin resistant; Hygr, hygromycin resistant
ISSI positive lines are underlined: if the site-specific integrants were derived from a
hemizygous target plant, the R2 plant should be all Hyg-sensitive and all derived R3 seeds
should be sensitive to Hyg while segregating (3:1) on Km or 100% resistant to Km. If the
site-specific recombinants were derived from a line homozygous for the Hsc1 locus, the R3
seeds should segregate on Hyg and Km or should yield progeny for 100% sensitive to Hyg
and 100% resistant to Km; progeny plants of the R3 or R4 seed stocks indicated in bold
were tested by PCR (see Table 3).
* We observed less than expected Kmr seedlings in the 16-12 R3 progeny seed stocks.
However in the next R4 generation, normal segregation patterns for a single locus insert
were obtained.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Table 3: PCR analysis on leaf DNA of SSI and non-SSI transformants homozygous for the
targeting Kmr construct.
(e) Plant Generation Positive PCR for
target locus
(attP site)a
Positive PCR for
recombined
outcome Ib
Positive PCR for
recombined
outcome IIc
Positive PCR
LB-RB footprintd
Non-SSI
1 16-4-3-4 R3 + - - -
2 16-4-2-5 R3 + - - +
3 16-4-2-4 R3 + - - +
4 16-4-2-3 R3 + - - +
SSI
5 16-11-2-1 R3 - + - +
6 16-11-2-3 R3 - + - +
7 16-11-3-8 R3 - + - +
8 16-11-1-2 R3 - + - +
9 16-12-1-2-9 R4 - + - -
10 16-12-1-2-8 R4 - + - +
11 16-12-2-3-7 R4 - - + +
12 16-12-2-3-10 R4 - - + +
13 13-3-7-1 R3 - - + +
14 13-3-6-1 R3 - - + +
15 13-3-1-2 R3 - + - +
a Primers a and d yield the 305-bp fragment present in the original target locus.
b Primers a and b yield the 478-bp fragment diagnostic for SSI recombined outcome I.
c Primers a and c yield the 1022-bp fragment diagnostic for SSI recombined outcome II.
d Primers e and f yield the 591-bp fragment.
e The numbers in the first column are corresponding to the lanes used for Southern blot
analysis (see Figure 2).

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
Figure legends
Figure 1 Schematic outline of the site-specific integration strategy when using the pK2L2B
vector for site-specific recombination into the Cre-Hsc1 target line. Sizes of the expected
fragments after digestion with SphI or NcoI when using the HPT (probe 1), the INT (probe 2),
the NPTII (probe 3) and the GUS (probe 4) probes are indicated. Primer pairs are shown as
a, b, c, d, e, f above the T-DNA construct schemes. (a) Structure of the T-DNAs in the CRE
and INT-expressing target line with an attP site (PP’) for targeted integration. The two T-
DNAs are inserted at different loci. The Cre locus is homozygous and is stably present. (b)
Structure of the targeting T-DNA vector with two copies of an attB site, the Km resistance
selectable marker, the P35S::eGFP/GUS expression cassette and two directly oriented loxP
sites next to the T-DNA borders. (c1) Structure of the CRE intermediate, obtained by CRE-
mediated circularization of the K2L2B T-DNA. (c2) and (c3) Structure of the INT
intermediates I (c2) and II (c3): INT activity will lead to site-specific recombination of the PP’
site in the target locus with attB I or attB II sites of the CRE intermediate. (d) Recombined
outcome I as a result of CRE activity on the INT intermediate I. (e) Recombined outcome II
as a result of CRE activity on the INT intermediate II. Abbreviations: LB, left border; RB, right
border; 3’ocs, 3’ end of the octopine synthase gene; NPTII, neomycin phosphotransferase II
gene; Pn, promoter of the nopaline synthase gene; T35S, 3’ terminator of CaMV; eGFPGUS,
eGFPGUS fusion gene of the enhanced green fluorescent protein and β-glucuronidase;
P35S, CaMV 35S promoter; 3’n, 3’ end of nopaline synthase gene; HPT, hygromycin
phosphotransferase gene; Pcas, cassava vein mosaic virus promoter (Verdaguer et al.,
1998); INT, integrase gene of bacteriophage φC31 (Dafhnis-Calas et al., 2005); 3’chs’, 3’
end of the chalcone synthase gene; CRE, CRE recombinase gene; BAR, phosphinothricin
transferase gene; PSSUARA, the small subunit promoter of Arabidopsis; TP, transit peptide;
3”g7, 3’ terminator of gene 7; SI, SphI; NI, NcoI. The open triangle represents the loxP site;
the open boxes with PP’ and BB’ symbolize the attP and attB recombination substrate sites
for the INT recombinase and the open boxes with BP’ and PB’ symbolize the attL and attR

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.
recombination sites. Note: the P35S::eGFP/GUS expression cassette can be replaced by
any desired DNA fragment.
Figure 2 Southern blot analysis on the progenies of transformants with site-specific
integration and a randomly inserted pK2L2B T-DNA integration. (a) DNA gel-blot of NcoI-
digested DNA hybridized with the INT probe. The target locus in the Cre-Hsc1 plant yields a
2.6-kb NcoI fragment and the expected NcoI fragment in the site-specific integrants is 1.8 kb
for recombined outcome I and 2.8 kb for recombined outcome II. (b) DNA gel-blot SphI
fragments detected using the GUS probe. The expected fragment in the site-specific
integrants is 3.7 kb for recombined outcome I. No hybridization bands are expected in
recombinants with outcome II. Random integration of the targeting construct will yield a right
T-DNA–plant junction of more than 4.1 kb. (c) DNA gel-blot of SphI-digested DNA hybridized
with the HPT probe. No hybridizing fragments should be detected in the recombinants, while
one T-DNA–plant junction fragment bigger than 2.2 kb is expected in the case of random
integration of the targeting construct. (d) DNA gel-blot SphI fragments detected using the
NPTII probe. The NPTII probe should detect the same hybridizing T-DNA–plant junction
fragment bigger than 1.8 kb in recombinants with outcome I and II, while this T-DNA–plant
junction fragment should be of a different size in transformants with random integration of
the targeting construct. (e) DNA gel-blot of SphI-digested DNA hybridized with the INT probe.
A T-DNA–plant junction fragment bigger than 3.3 kb is expected in all DNA samples. Lanes
1-4 contain DNA from different homozygous progenies of 16-4, lanes 5-8 that from 16-11,
lanes 9-12 that from 16-12 and lanes 13-15 that from 13-3. The numbers indicating the lanes
are also used in Table 3.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.

Acc
epte
d A
rtic
le
This article is protected by copyright. All rights reserved.