Protein Interactions Visualized By Atomic Force Microscopy
Transcript of Protein Interactions Visualized By Atomic Force Microscopy

science/technology ϋ?ν.
Protein Interactions Visualized By Atomic Force Microscopy
Aresearch group has extended the applications of atomic force mi-Icroscopy (AFM) by using the
technique to obtain dramatic "movies" of individual protein molecules interacting with each other in real time. The work-by graduate student Mario B. Viani, post-doc lia I. Pietrasanta, physics professors Helen G. and Paul Κ Hansma, and coworkers at the University of California, Santa Barbara (UCSB)— has potential implications for the direct visual analysis of protein function and dynamics [Nat Struct Biol, 7, 644 (2000)].
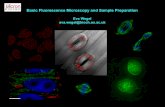
The researchers used AFM to image binding interactions between the bacterial chaperonin protein GroEL and its co-chaperonin, GroES. The barrel-shaped GroEL assists protein folding by accepting unfolded proteins into either of two internal chambers. GroES then caps the chamber, creating a protected environment in which the proteins have an opportunity to fold to their native states.
In the study, the UCSB group's AFM images show GroES binding to and dissociating from individual GroEL molecules on a solid support. The images made it possible to analyze dynamics of the interactions, such as the lifetime of individual GroEIXiroES binding events. AFM images acquired in the study are of such high resolution that the central chambers of the GroEL proteins are visible.
AFM has been used previously to observe motions of individual proteins. However, these studies looked just at protein motion, rather than at protein interactions. In addition, the signal-to-noise ratios of the images obtained in
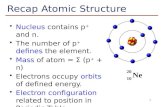
ί Scanning electron micrograph of small AFM cantilevers (right) and a conventional commercial cantilever (left). Small cantilevers, such as the one used in the UCSB study (arrow), permit faster measurements and higher signatto-noise ratios. Reprinted with permission from Nature Structural Biology, ©2000
the current study are much higher than those of images from the earlier studies. 'The GroEI^GroES work is a significant step because finally we can image quickly and gently enough to see protein-protein interactions that we can understand," Paul Hansma says.
Julio M. Fernandez, professor and chair of the department of physiology and biophysics at Mayo Clinic, Rochester, Minn., whose research interests include AFM of proteins, believes the study is an important one, calling it "a clear breakthrough that deserves significant recognition."
The ability to visualize GroEI^GroES interactions in real time was made possible by the UCSB group's recent development of AFM cantilevers much small-
r'^nj*^^^
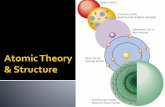
AFM Image shows association and dissociation of GroEL-GroES complexes at the single-molecule level. Horizontal "tubes" each represent individual chaperonin molecules that have been repeatedly scanned by AFM. During the time range (horizontal axis) of the scans, height changes are observed along each tube, with brighter areas indicating taller features. Trace at bottom shows height variations along one of the tubes (indicated by arrow at top). Increases in height occur when GroES caps GroEL and decreases correspond to dissociations of GroES from the complex. These complexation and dissociation processes are depicted with electron microscopy images (by crystallography professor Helen Saibil of Birkbeck College, London, and coworkers) of barrel-shaped uncapped GroEL and beehivelike GroES-GroEL complexes.
er than those found in conventional AFM instruments \J. Appl Phys., 86, 2258 (1999)]. In AFM, images are obtained by measuring forces between an AFM tip and a sample. These forces cause deflections of a cantilever on which the tip is mounted, and the deflections are detected by a laser beam to generate an image of the surface topology. The small cantilevers used by the UCSB group in the current study "have higher resonant frequencies and better signal-to-noise ratios than larger [conventional AFM] cantilevers with the same spring constant," Viani says.
Can a technique that visualizes protein interactions provide information thafs truly useful? 'That depends on who you talk to," Viani says. "Some researchers in the chaperonin field think this work is useless because the system is artificial"—that is, because the GroEL molecules are adsorbed to a surface instead of a being in a more natural biological en-
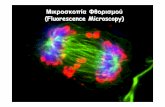
AFM image of GroEL adsorbed on mica. Central channels of the individual GroEL molecules are visible. Reprinted with permission from Nature Structural Biology, ©2000
vironment. "Others in the field like our [technique] because it will let them look at the properties of individual molecules, even if they are in a model system."
Associate professor of cellular and molecular pharmacology Jonathan S. Weiss-man of the University of California, San Francisco, whose research interests include chaperonins, says: "Ifs a really neat experiment. There is no big punch line yet, but single-molecule experiments like this one have a great potential for revealing behavior that is obscured by the bulk properties. I am sure this approach will be useful for a variety of different systems, even if it does end up being mostly confirmatory for GroEL."
Stu Borman
4 0 AUGUST 14,2000 C&EN