Molecular Cloning and Expression Analysis of NteIF2α A In ... · regulation of global protein...
Transcript of Molecular Cloning and Expression Analysis of NteIF2α A In ... · regulation of global protein...

Molecular Cloning and Expression Analysis of NteIF2α In Response to Stress in Nicotiana tabacum Zhaoqi, Li sibin, Li ning, Guo hao, Long yue, Cui hong, Yang yongxia, Jia hongfang, Zhang hongying, Zhang songtao*,
Henan agricultural university, College of tobacco science, Tobacco Cultivation Key Laboratory of China Tobacco, Zhengzhou 450002, China
* Corresponding Author: [email protected]
Abstract
eIF2α (α subunite eukaryotic translation initiation factor 2) plays an important role in the regulation of
protein synthesis. The phosphorylation of eIF2α could lead to down-regulate global protein synthesis. To
investigate the function of eIF2α and explore its roles in plant stress response, the ORF (Opening Read
Frame) of NteIF2α was cloned from Nicotiana tabacum L cv. K326 by RT-PCR and the mRNA expression
level in different tissues and stress conditions were analyzed by qRT-PCR method. Our results showed that
there were 2 copies of NteIF2α in N. tabacum K326, NteIF2α-1(GenBank accession number: KR184725)
and NteIF2α-2(GenBank accession number: KR184726). The nucleotide sequences shows 98.3%
similarity, while the protein sequences show 99.7% similarity to each other. Also, NteIF2α contains the
conserved domain of eIF2α which includes kinase interaction site, phosphorylation site and eIF2B
interaction site. Further, phylogenetic analysis revealed that NteIF2α-1 and NteIF2α-2 might be derived
from its progenitor Nicotiana sylvestris and Nicotiana tomentosiformis, respectively. Twenty-four
phosphorylation sites were predicted in the protein sequence. Among them, the Ser56 which was the
phosphorylated site of protein kinases were highly conserved from animals to plants. In addition, the
expression pattern was detected in different organs of N. tabacum K326. The results showed NteIF2α-1
and NteIF2α-2 were expressed in all organs examined and the expression profiles in N. tabacum displayed
differently, because the expression of NteIF2α-2 was higher than that of NteIF2α-1, which indicates the
function of the eIF2α family members were different in the organs. Finally, the activation of NteIF2α by
SA(salicylic acid) and MeJA(methyl jasmonate) suggests that NteIF2α participates in the plant defense
response to insect. Our results provides an important basis for further studying the function of eIF2α and
its roles in plant stress response.
Key Words: Nicotiana tabacum; eIF2α; gene cloning; gene expression
Introduction Eukaryotic translation initiation factor (eukaryotic translation initiation factor 2, eIF2) plays an important role during
the protein synthesis process. In the initiation of protein translation, eIF2 coupled with GTP (guanosine triphosphate)
and initial methionine-tRNA (Met-tRNAimet) to form eIF2·GTP·Met-tRNAimet Triplet complex, and then the
complex interacts with eIF1, eIF1A and eIF3 to form the 43s pre-initiation complexes. EIF2 consists of three subunits
heterologous trimer- α、β、γ. Serine56 of α-subunit could be phosphorylated by eIF2α kinase, such as GCN2 (general
controlnon derepressible-2), HRI (heme-regulated inhibitor), PERK (interferon inducible protein) and PKR (PKR-like
endoplasmic reciculum-resident protein kinase). Thus, leading to the suspension of translation initiation and down-
regulation of global protein synthesis.
Phosphorylation of eIF2α by eIF2α kinase to down-regulate protein synthesis are widespread in eukaryotic cells. In
mammalian cells, the phosphorylation of serine at site 51 of eIF2a is mediated by four protein kinases, each of which
respond to different stress stimuli. For example, in the absence of heme, HRI is activated to regulate the synthesis of
erythrocyte globulin. GCN2 kinases are activated by uncharged tRNAs in the absence of amino acids. PERK is activated
when the endoplasmic reticulum protein misfolded. All these kinases contain a typical protein kinase domain that could
activate the phosphorylation of eIF2a. In Arabidopsis thaliana, the serine52 of AteIF2a could be phosphorylated when
Arabidopsis thaliana was treated with herbicides that may affect amino acid synthesis, and the phosphorylation is
accomplished by AtGCN2.
eIF2α contains about 300 amino acid residues with high sequence similarity from different species. About 180 amino
acids at the N-terminus of the eIF2α are necessary for the phosphorylation of serine at site 51. The N-terminal sequence
of the yeast and human eIF2α shows the similarity of 56%, while the similarity of the first 100 amino acid residues
reach 75%. Moreover, it was showed that sequences prior to Serine51 and the conserved regions KGYID83 were
important for the binding of eIF2B. The conserved KGYID83 region is also involved in the interaction of eIF2α with
kinases.
In Arabidopsis, various stresses (including amino acid deficiency, purine deficiency, UV, cold stress and injury
treatment) could activate phosphorylation of eIF2α, leading to decrease of protein synthesis. Besides, plant homorney
include methyljasmonate (MeJA) and salicylic acid (SA) could also phosphorylate eIF2α, suggesting that
phosphorylation of eIF2α might also be involved in plant defense response. However, little is known about the reseach
of eIF2α from tabacco and its roles in plant defense response.
Materials and methods Plant materials
Nicotiana tabacum K326 plants were grown in soil in a growth chamber with a temperature/light setting of 28 ℃/16 h.
Preparation of cDNA
Total RNA was isolated from leaf material using RNeasy Plant Mini Kit (Qiagen), then the cDNA was prepared by
using the random primers and obtained RNA as template.
Molecular Clone of NteIF2α
Using the prepared cDNA as a template, primers were shown as following: NteIF2α-ORF-F: 5'-
CGCGCGGCAGCCATATGGCGACCAACTCCCCAAACCTC-3’ and NteIF2α-ORF-R: 5'-
CCATGCATGGTCGACTTACTCTCTAATTCCAGCTCCTG-3’. PCR was performed with 50 μl PCR reaction
mixture that included 1 μL template, 5 μL each of the upstream and downstream primers, 1 μL of DNA polymerase, 5
μL of dNTP, and 33 μL of ddH2O. PCR program: pre-denaturation at 95 ℃ for 5 min; denaturation at 95 ℃ for 1 min,
annealing at 58 ℃ for 45 s and extension at 72 ℃ for 1 min for 30 cycles,Then extended at 72 ℃ for 1 min, and
stored at 4 ℃. PCR products were purified and ligated into pMD19-T, then transformed into E. coli. Positive clones
were sequenced.
Bioinformatics Analysis
Clustal W 1.83 was used for Multiple alignment analysis and phylogenetic tree construction. Protein hydrophobicity
Analysis and transmembrane prediction were performed using ProtScale (http: // web.
Expasy.org/protscale/ ,TMpred(http://embnet.vitAl-it.Ch/software/TMPRED_form.Html) and Phosphorylation site
prediction was analyzed by the online tools (http://www.cbs.dtu.dk/services/Net-Phos /).The secondary structure is
predicted using online tools SOPMA (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.Pl? Page = npsa_sopma.html).
Amino acid sequences were submitted to the Swiss Bioinformatics Institute website(Http://swissmodel.expasy.org/)
for homology modeling. Three-dimensional structure was displayed by PyMol software.
Real-time quantitative RT-PCR
RNA was extracted from the middle leaves, roots, stems and flowers of N.tabacum K326. qRT-PCR was performed
with 20 μl PCR reaction mixture that contained 2 μl of cDNA, 10 μl of 2 × LightCycler® 480 SYBR Green I Master
(Roche, Swiss), 0.4 μl of primers (Table 1), and 7.2 μl of nuclease-free water. The mixure was incubated at 95℃ for
10 min, followed by 40 cycles of 95℃ for 10 s, 60℃ for 30 s ,72℃ for 30 s. Each sample was run in triplicate for
analysis. The expression levels of mRNAs were normalized to tobacco ribosome gene L25(L18908) and were
calculated using the 2-ΔΔCt method (Livak and Schmittgen, 2001).
Stress treatment
The seedlings of N.tabaccum K326 was grown on MS medium, and were sprayed with 2 mmol·L-1 salicylic acid (SA)
or 100 μmol·L-1 methyl jasmonate (MeJA) after grown for 1 month. 200 mg samples were taken at 0,3,6,12,24,48 and
72 h after SA treatment and 0, 3, 6, 24, 48 and 72 h for MeJA treatment.
A
B
D
Results and discussion
1.Molecular clone of NteIF2a from N.tabacum
The K326 cDNA was used as template to amplify the PCR fragment. The PCR product was ligated into the
pMD19-T vector and tranformed into E.coli. The resulted recombinant plasmid verified and The fragment was
about 1000 bp(Fig. 1). The results showed that there were 2 copies of NteIF2α in N. tabacum K326, Named
NteIF2α-1(GenBank accession number: KR184725) and NteIF2α-2(GenBank accession number: KR184726),
respectively.
Fig.1 Clony PCR identification of the NteIF2α
from N. tabacum K326
2.Homologous alignment and phylogenetic analysis of NteIF2α
The obtained eIF2α nucleotide was translated into amino acid sequence and used for alignment analysis (Figure 2).
The results showed that NteIF2α-1 and NteIF2α-2 was showed 99.7% similarity in amino acid sequence, and
NteIF2α contains the conserved domain of eIF2α which includes kinase interaction site, phosphorylation site and
eIF2B interaction site.
In addition, these plant-derived eIF2α sequences showed high similarity over 76.1%, while the NteIF2α showed
low similarity with human (45.5%) and yeast (45.1%). The results showed that the NteIF2α sequence was the
coding sequence of Nicotiana tabacum eIF2α and the typical domain was identical to that of human and yeast
eIF2α, which suggest the conserved regulation mechanism of protein synthesis among animal and plant.
The phylogenetic tree was constructed according to the nucleotide sequence of eIF2α(Fig 3). The
phylogenetic analysis revealed that NteIF2α-1 and NteIF2α-2 might be derived from its progenitor
Nicotiana sylvestris and Nicotiana tomentosiformis, respectively.
3. Analysis of protein hydrophobicity and phosphorylation site prediction of NteIF2α
Hydrophobicity analysis showed that the maximum hydrophobicity of NteIF2α protein was 2.011 (268 amino acids)
and the maximum hydrophilicity was -3.20 (323 amino acids). Phosphorylation site prediction showed that there
are 14 potential serine (Ser) phosphorylation sites, 3 threonine (Thr) phosphorylation sites and 7 tyrosine (Tyr)
phosphorylation sites in the amino acid sequence of eIF2α protein. The 56th serine phosphorylation site score was
as high as 0.917, suggesting that the site is a potential phosphorylation site, which is involved in the regulation of
Table 1 Primer sequences of NteIF2α-1 and NteIF2α-2 used for qRT-PCR
Fig.2 Alignment of the amino acid
sequences of eIF2α (The same amino
acid residues are highlighted in black)
NteIF2a-1 and NteIF2a -2: the eIF2a sequence
of tobacco N. tabacum K326; N. sylvestris:
eIF2α sequence of tobacco Nicotiana sylvestris
(gi: 698443790); N.tomentosiformis: eIF2α
sequence of Nicotiana tomentosiformis (gi:
697118772); S. lycopersicum: eIF2α sequence
of tomato Solanum lycopersicum (gi:
460378959); Arabidopsis: eIF2α sequence of
Arabidopsis thaliana (gi: 18405334);Zea mays:
eIF2α sequence of maize (gi: 226505440);
Homo sapiens: human eIF2α sequence (gi:
4758256);S. cerevisiae: eIF2α sequence of
yeast Saccharomyces cerevisiae (gi: 903889);
The sites indicated by the arrows are the
phosphorylation site of the protein kinase (S56)
and the site of action of the kinase
(V38Y86I87D88), and the triangular symbol
represents the eIF2B interaction site
(E54K84G85).
5.Gene expression analysis of NteIF2α in different organs and under stress treatment
The gene expression of NteIF2α-1 and NteIF2α-2 were analyzed by RT-PCR(Fig 6). The results showed NteIF2α-1 and NteIF2α-2 were
expressed in all organs examined and the expression profiles in N. tabacum displayed differently, because the expression of NteIF2α-2 was
higher than that of NteIF2α-1, which indicates the function of the eIF2α family members were different in the organs.
The activation of NteIF2α by SA(salicylic acid) and MeJA(methyl jasmonate) was shown in Fig 7. These suggests that NteIF2α
participates in the plant defense response to insect.
Fig.4 Protein hydrophobicity (left) and phosphorylation sites (right) analysis of
NteIF2α
Fig.3 The phylogenetic tree of the nucleotide
sequences of eIF2α
Fig.7 RT-qPCR analysis of NteIF2α gene expression under stress treatment(A: SA treatment;
B: MeJA treatment )
A B
Table 1 Primer sequences of NteIF2α-1 and NteIF2α-2 used for qRT-PCR
Primer name Sequence (5’-3’) Purpose
NteIF2α-1-F: CTCCGAAACCCTAATAACCTTCACT Amplification of
NteIF2α-1-R: CCTTCTATGTTGTTGTACTCGAGAAGC NteIF2α-1
NteIF2α-2-F: CTCCGAAACCCTAATAACTCCCAC Amplification of
NteIF2α-2-R: CCTTCTATGTTGTTGTACTCGAGAAGT NteIF2α-2
NteIF2α-F: CCTCGAATGCCGAATGTACGAAGCC Amplification of
NteIF2α-R: CCTTCTATGTTGTTGTACTCGAGAAG NteIF2α
L25-F: GCTTTCTTCGTCCCATCA Reference gene
L25-R: CCCCAAGTACCCTCGTAT
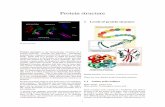
4. Protein tertiary structure prediction
The protein secondary structure of NteIF2α-1 contained α-helix(48.98%) , β-sheet (16.33%), β-turn (10.79%) and
random coil (23.91%). In addition, the N-terminal of the protein contains five antiparallel β-sheets, which are
similar to the eIF2α spatial structure in human and yeast (Fig 5).
Fig. 5 The tertiary structure prediction of
NteIF2α-1(blue)and NteIF2α-2(green)
Fig.6 Relative expression of NteIF2α-1 and NteIF2α-2
in different organs of N. tabacum L cv. K326
Roots Stems Leaves Flowers
Conclusion 1. We clone NteIF2α from Nicotiana tabacum L cv. K326. NteIF2α-1and
NteIF2α-2 might be derived from its progenitor Nicotiana sylvestris
and Nicotiana tomentosiformis, respectively.
2. The mRNA expression level of NteIF2α-1and NteIF2α-2 in different tissues
indicates the function of the eIF2α family members were different in the organs.
3. The activation of NteIF2α by SA and MeJA suggests that NteIF2α participates
in the plant defense response to insect. The results provides an important basis
for further studying the function of eIF2α and its roles in plant protein synthesis.
2016
_AP
PO
ST
03_Z
hang
Son
gtao
Con
gres
s201
6 -
Doc
umen
t not
pee
r-re
view
ed b
y C
OR
ES
TA