Inhibition of γ-thrombin-induced human platelet aggregation by histone H1subtypes and H1.3...
Transcript of Inhibition of γ-thrombin-induced human platelet aggregation by histone H1subtypes and H1.3...

Platelets, August 2009; 20(5): 349–356
ORIGINAL ARTICLE
Inhibition of c-thrombin-induced human platelet aggregation by histoneH1subtypes and H1.3 fragments
GERALD SOSLAU1,2, PHILLIP J. PREST2, REINER CLASS3,4, MONIKA JOST3,
& LYNN MATHEWS2
1Department of Biochemistry & Molecular Biology, 2Office of Professional Studies in the Health Sciences,3Department Radiation Oncology, Drexel University College of Medicine, Philadelphia, PA, USA, and4Toxicology & Premedical Development, Pharmacelsus, GmbH Science Park 2, 66123 Saarbrucken, Germany
(Received 12 March 2009; revised 1 May 2009; accepted 13 May 2009)
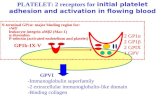
AbstractHuman platelets are differentially activated by varying concentrations of �-thrombin or by �- and g-thrombin viathree thrombin receptors, PAR-1, PAR-4 and GPIb ~�. It is likely that the development of a normal or abnormal hemostaticevent in humans is dictated, in part, by the selective activation of these receptors. The ability to differentially inhibitthese thrombin receptors could, therefore, have clinical significance. We have previously demonstrated that histoneH1 selectively inhibits the PAR-4 receptor. In the current study we investigated whether five subtypes of the H1 moleculeor fragments of the H1.3 subtype differentially inhibited the PAR-4 receptor. PAR-4 inhibition by all H1 subtypeswas saturated at 1 uM with no statistical difference observed with the five H1 subtypes tested. Of the five fragmentsgenerated from the H1.3 molecule only one had significant inhibitory activity against PAR-4. The C-terminal fragment,N.1, generated by the proteolysis of the parent molecule by Asp-N endoproteinase (Aeromonas proteolytica) at the singleaspartate residue, showed the same level of PAR-4 inhibition as the intact H1.3 at 1 uM concentrations. Removal of twoN-terminal amino acids (Asp-Val as determined by MALDI analysis) from the N.1 fragment further enhanced itsinhibitory activity. These studies may help to develop specific drugs to differentially inhibit the platelet thrombin receptors.
Keywords: Platelet aggregation, �–thrombin, histone 1
Introduction
Hemostasis and thrombosis are orchestrated by the
generation of �-thrombin, a serine protease, derived
from the inactive precursor, prothrombin (factor II).
Thrombin is required for both the intrinsic and
common coagulation cascades [1], generates the
proteolytic conversion of fibrinogen to fibrin
required for clot formation and also activates
platelets via specific thrombin receptors. Evidence
indicates that �-thrombin may be autolytically
cleaved, in vivo, to initially form �-thrombin which
in turn can be further hydrolyzed to yield g-thrombin
[2, 3]; the same process can be induced in vitro with
the addition of plasmin, factor Xa, or trypsin [2].
These additional forms of thrombin may be physi-
ologically significant by differentially activating three
human platelet thrombin receptors, PAR-1 [4],
PAR-4 [5], and GPIb [6, 7]. The mechanism of
thrombin activation of platelets via the protease
activated receptors, PAR-1 and PAR-4, has been
well established [8]. The mode of thrombin-induced
activation of platelets via GPIb has not been as well
resolved. While one group indicated that GPIb was
not involved in thrombin-induced platelet aggrega-
tion at all [9], the accumulating evidence demon-
strates that GPIb-thrombin interactions are directly
involved in platelet aggregation by a unique pathway
[6, 7, 10–12]. �-Thrombin activates platelets pri-
marily via PAR-1 and/or GPIb, while g-thrombin
and �-thrombin only function through PAR-4 [6, 13,
14]. The specificity of the platelet thrombin receptors
offers the potential to selectively inhibit responses for
both clinical and research purposes.
Correspondence: Gerald Soslau, Office of Professional Studies in the Health Sciences, Drexel University College of Medicine, 245 N. 15th Street, MS 344,
Philadelphia, PA 19102, USA. Tel: þ215-762-7831. Fax: þ215-762-7434. E-mail: [email protected]
ISSN 0953–7104 print/ISSN 1369–1635 online � 2009 Informa Healthcare Ltd.
DOI: 10.1080/09537100903047745
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

The structural interactions of histone H1 with
DNA are well documented. Eukaryotic DNA is
efficiently packaged into a relatively small nuclear
volume called chromatin [15]. Chromatin consists
of repeating nucleosomes: four histone proteins – two
each of H2A, H2B, H3, and H4 – form an octamer
around which 160–220 base pairs of genomic DNA
wrap [15]. The nucleosome further contains an
average of one lysine-rich linker histone, H1,
resulting in the ‘‘beads on a string’’ motif that can
be visualized by electron microscopy [16]. In the
absence of H1, chromatin unravels into unstable
linearized filaments, demonstrating the importance of
H1 in stabilizing the structure of the nucleosome [17].
The protective role of DNA structure by histone
H1 is but one of many biological roles ascribed to
this protein. Histone H1 has been shown to affect
several different cellular processes: directly causing
cytolysis [18]; reducing aerobic and anaerobic
metabolism [19]; altering cell morphology and con-
fluent density [20]; limiting DNA and RNA produc-
tion [21], and; one subtype, H1.2, has been shown to
trigger apoptosis [22]. Histone H1 also affects the
progress and outcome of the inflammatory response
by activating macrophages and neutrophils while also
regulating several leukocyte functions [23]. In addi-
tion, histone H1 appears to possess anti-neoplastic
properties, including suppressing tumor activity
in experimental mammary carcinoma [24] and anti-
tumor effects against leukemic cells [25]. The
origin of H1’s cytotoxicity may be related to its
similarity with proteins secreted by activated macro-
phages [26]. Furthermore, H1 directly activates
macrophages to have nonoxidative antimicrobial
potential; these activated macropages may recognize
and destroy neoplastic cells [27]. Histone H1 has
further been demonstrated to have strong antibiotic
properties, particularly against pathogenic Gram-
negative bacteria [28]. All histone H1 molecules
share the same three-domain structure demonstrat-
ing an unmatched degree of evolutionary conserva-
tion. Therefore, the secondary H1-structure seems to
be of major importance for its biological properties.
If histone H1, or fragments of H1 are systemically
administered into patients its effects on blood and
its components must be thoroughly understood.
The aim of the present study was to further expand
on our observation that histone H1 can inhibit
g-thrombin-induced platelet aggregation [14]. Here
we show that recombinant subtypes of H1 and one
proteolytic fragment of the H1.3 molecule all
selectively inhibit g-thrombin activation of human
platelets via the PAR-4 receptor. This selective
inhibition of the platelet aggregation – PAR-4 path-
way, especially by a fragment of the H1.3 molecule,
provides a basis for the potential development
of specific drugs to differentially inhibit platelet
thrombin receptors.
Materials and methods
Recombinant histone H1 subtypes (H1.0, H1.2,
H1.3, H1.4 and H1.5) were purchased from Alexis
Biochemicals (San Diego, CA, USA). Five different
fragments of the H1.3 subtype (N.1, N.2, NG, G,
GC) were prepared by enzymatic cleavage [29]. The
five H1.3 fragments refer to different regions of the
peptide (N-terminal, C-terminal, globular regions),
with N.1 and N.2 representing complementary
pieces resulting from a single proteolytic cut (see
Figure 1). Fragments N.1 and N.2 were prepared by
digesting rhH1.3 with endoproteinase AspN (Sigma,
St. Louis, USA), (2 U/mg protein) over night at 37�C
and purified by HPLC using a Vydac RP 214TP
column and eluted with water and acetonitrile.
The identification and characterization of the
proteolytically generated H1.3 fragments, N.1 and
N.2, was critical since the N.1 fragment is the only
peptide that significantly inhibited g-thrombin-
induced platelet aggregation. The peptide eluting
early from an HPLC column should be the lower
molecular weight N-terminal fragment (m wt 7009)
and the second peak should be the C-terminal
(15,230 m wt) peptide. However, when these HPLC
fractions were subjected to PAGE (polyacrylamide
gel electrophoresis) analysis the early eluting fraction
migrated slower as a single band and the second
fraction migrated faster as a lower molecular weight
band. This occurred with both a 15% gel and a
10–20% gel and neither fraction migrated at the
appropriate apparent molecular weight (data not
shown). Samples were therefore analysed by MALDI
mass spectrometry to correctly identify the proteoly-
tic fractions. The MALDI analysis was conducted by
the Washington University Mass Spectrometry
Service (St. Louis, MO, USA). MALDI analysis
clearly demonstrated that the early and late eluting
peaks from HPLC were indeed the N-terminal and
C-terminal peptides, respectively (Figure 2a, b). N.1
is designated the 15,237 C-terminal fragment while
N.2 is the 7009 N-terminal fragment. It is interesting
to note that the N-terminal fragment consists of two
species, one with no N-terminal methionine group
(m wt 7009 daltons) and one with 2 N-terminal
methionine groups (m wt 7272 daltons) in a 2:5
ratio. The second peak in Figure 2b at an apparent
m wt of 7612 daltons is actually a dimeric form of the
15,237 dalton species. This was proven by analysing
the sample by deconvolution of the electrospray mass
spectrometry data and by mass spectrometry of
tryptic digests. The final spectrum shows a single
peak at 15,229 daltons and a 7614 species that is an
artifact of the deconvolution program (Figure 2c).
The tryptic data is not shown.
Blood was obtained from donors who signed
consent forms under an approved IRB protocol, by
venipuncture and collected in 4.5 ml sodium citrate
vacutainers. Washed platelets were prepared as
350 G. Soslau et al.
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

previously described [30] where the final
platelet pellet was gently disrupted with 10 ml of
pH 6.0 sodium citrate solution (1 mM sodium
citrate in 20 mM Tris, 150 mM NaCl) followed
by their resuspension in hepes Tyrode’s buffer,
(136 mM NaCl, 2.7 mM KCl, 3.3 mM NaH2PO4,
1.0 mM MgCl2, 3.8 mM Hepes, pH 7.4) with
2 mg/ml dextrose and 1mg/ml bovine serum albu-
min at the initial PRP (platelet rich plasma) volume
at 2 – 3� 105 platelets/ml.
Platelet aggregations were conducted on a dual
channel lumi aggregometer (Chronolog corp.
Haverford, PA, USA) with 480 ml of washed platelets
plus 5 ml human fibrinogen (final concentration of
100 mg/ml). Platelet samples were incubated for
2–4 min with the histones/histone fragments prior
to the addition of agonist. In some experiments the
histone fragments were pre-treated with N-terminal
aminopeptidase (AP) from Aeromonas proteolytica
(Sigma) for varying times prior to the addition to
Figure 2. MALDI mass spectrometric analysis of the histone H1.3 fragments N.1 and N.2. (a) Fragment N.2; Panel (b) fragment N.1;
Panel (c) spectrum of fragment N.1 after deconvolution of the electrospray mass spectrometric data; (d) fragment N.1 treated with
aminopeptidase as described in Methods.
Figure 1. Schematic of the structural regions of the histone H1.3 molecule and its fragments.
Inhibition of �-thrombin-induced human platelet aggregation by histone H1subtypes and H1.3 fragments 351
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

washed platelets. Aggregations with these samples
were then conducted as above. In all cases when
g-thrombin was employed as the agonist the
concentration was experimentally set to yield
about 60–80% aggregation (final concentration of
g-thrombin 8–25 nM).
Histone H1.3 inhibition studies of thrombin
activity with the synthetic substrate, H-D-CHG-
Ala-Arg-pNA 2AcOH (Pefachrome TH, Cen-
terchem, Inc., Norwalk CT, USA), were conducted
as previously described [31]. The generation of a
colored product by the hydrolysis of the substrate in
the reaction buffer (50 mM Tris-HCL, 50 mM
imidazole, 150 mM NaCl, pH 8.0) was monitored
at 405 nm over a 4 min time period on a Hitachi
U-2000 spectrophotometer. Reactions were run in
triplicate with 10 and 20 nM g-thrombin or with
0.5 U/ml �-thrombin (Chronolog, Corp, Havertown,
PA, USA). The Pefachrome TH substrate concen-
trations employed were 0.05, 0.1 and 0.2 mM.
Results
We have previously demonstrated that histone H1
selectively inhibits g-thrombin – induced platelet
aggregation and has no effect on the �-thrombin –
induced reaction [14]. The current studies focused
on potential structural aspects of histone1 inhibition
of g-thrombin-induced platelet aggregation. Previous
studies were performed with tissue-derived bovine
histone H1, which represents a mixture of several H1
subtypes. At least seven different subtypes or variants
of H1 have been described in higher eukaryotes (for
review see [32]). The globular core domain is highly
conserved between the subtypes; whereas the N- and
C-terminal regions are more divergent, which may
result in functional differences. For example,
only the H1.2 subtype has been shown to trigger
apoptosis [22]. To further dissect the contribution of
different subtypes to inhibiting g-thrombin–induced
platelet aggregation, we compared several human
full-length H1 subtypes in a typical platelet aggrega-
tion assay. All 5 subtypes of histone H1 tested
inhibited g-thrombin–induced platelet aggregation
with no significant difference at 1mM (Figure 3). The
degree of inhibition of platelet aggregation was
dose dependent with no statistical difference in the
inhibitory activities between the subtypes tested
(Figure 4). For further studies, H1.3 was used to
generate fragments of the parent molecule.
Fragments of the histone H1.3 molecule, as depicted
in Methods, were tested to determine if the inhibi-
tory activity resided in a portion of the molecule or
required the intact molecule. Of the fragments
tested, only the N.1 fragment of histone H1.3
inhibited platelet aggregation (Figure 5). At the
concentration tested (1mM), inhibition was compa-
rable to the full-length H1.3 while the N.2 fragment
had little to no effect at the same concentration, and
sometimes appeared to slightly stimulate the reac-
tion. Other fragments, NG, GC, G alone, or in
combination with N.2, also had little or no effect at
1 mM concentrations (Figure 5). This suggests that
the inhibitory activity resides within the C-terminal
portion of the molecule and, possibly, the third helix
of the globular domain. However, since the GC
fragment had no effect, it has to be assumed that the
C-terminal domains within these two molecules (N.1
and GC) have different conformations, the C-
terminus of N.1 presumably being similar to that of
full-length H1.3.
Next we investigated whether adding the N.1
fragment to the non-inhibitory fragment N.2 would
reveal a regulatory role of inhibition to N.2, i.e.
whether the two parts of the protein could function
when separated into two different subunits. In these
assays, N.1 was added at 0.1, or 0.2 uM as opposed
to the maximum concentration tested (1 uM).
Gamma-thrombin inhibition
0.00 0.25 0.50 0.75 1.00 1.250
102030405060708090
100110
H1.0
H1.2
H1.3
H1.4
H1.5
Histone conc. (uM)
% In
hib
itio
n
Figure. 4. Dose-response of g–thrombin-induced platelet aggre-
gation inhibited with different histone 1 subtypes. The inhibition
of platelet aggregation was conducted with four different
concentrations of each H1 subtype, in triplicate, with the
meansþ/-standard deviations graphed. There was no statistical
difference between groups of data by an ANOVA and Kruskal-
Wallis test.
H1.0 H1.2 H1.3 H1.4 H1.560
70
80
90
100
Histone 1 subtype
% In
hib
itio
n
Figure 3. The inhibition of g–thrombin-induced platelet aggre-
gation by histone 1 subtypes. The final concentration of each
histone 1 subtype was 1mM and the bar represents the mean for
each subtype. Each point represents an individual experiment.
352 G. Soslau et al.
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

Inhibition by N.1 alone was kept in the approxi-
mately 40% range by adjusting the concentration to
either 0.1 or 0.2 uM in order to determine if the
addition of N.2 significantly altered the final level of
inhibition. When N.1 was added to 1 mM N.2, which
by itself shows no inhibitory activity, inhibition
increased to about 75%, indicating a synergistic
effect of the two molecules. This appears to demon-
strate that the two fragments can function as subunits
and mimic the full-length protein (Figure 6).
Our previous studies indicated that H1 inhibited
g-thrombin-induced platelet aggregation at two
levels. There appeared to be a direct effect on
both the thrombin molecule and the PAR-4
receptor [14]. However, under the experimental
conditions described in Methods, no direct inhibi-
tion of �-, or g-thrombin could be detected using a
chromogenic peptide substrate for thrombin.
Therefore, independent of a direct effect on
g-thrombin, it was important to demonstrate that
the N.1 inhibition, like H1, again included a
specific direct effect on the PAR-4 receptor without
involving other agonist receptors. Figure 7 clearly
demonstrates that platelets, preincubated with
1.5 uM N.1 for 4 min prior to the addition of
agonist, were only inhibited at the PAR-4 receptor.
This inhibition was observed with either g-throm-
bin or TRAP-4A (AYPGKF). The reaction with
TRAP-4A was significantly inhibited by N.1 but
not as completely inhibited as was the g-thrombin
reaction.
Finally, we investigated the potential role of the
N-terminal residues of N.1 for platelet inhibition by
enzymatic removal of the N-terminus amino acids
with AP. The amount of AP employed for these
experiments was determined by titrating the amount
of enzyme added to platelets for 2 min prior to the
addition of g-thrombin. The maximum amount of
AP that resulted in no, or minimal levels of,
inhibition of platelet aggregation was employed. AP
concentrations above 0.1 U AP/ml significantly
inhibited the reaction with 1.0 U AP/ml being com-
pletely inhibitory. The random removal of
N-terminal amino acids of platelet surface proteins
with 1.0 U AP/ml also inhibited PAR-1-, collagen-
and ADP-induced platelet aggregations (data not
shown). Figure 8 shows a representative time course
N1 effect on aggregation
g-T PAR-1 PAR-4 U-4 Coll ADP0
25
50
75
100
Control
+1.5uM N1
Agonist
% A
gg
reg
atio
n
Figure. 7. The effect of N.1 on platelet aggregation induced by
various agonists. The platelets were either untreated (control) or
preincubated for 4 min with 1.5 uM N.1 followed by agonists at a
concentration sufficient to induce 60–80% aggregations. g-T is
g-thrombin (15 nM), PAR-1 activated with TRAP-1 (0.45 uM
SFLLRNP), PAR-4 activated with TRAP-4A (60 uM AYPGKF),
U-4 is the thromboxane (T�A2) analog, U46619 (50 uM), coll is
collagen (1–5 ug/ml), and ADP (1–10 uM). The ADP aggrega-
tions were conducted with PRP while all other aggregations were
with washed platelets. Each data point is a repeat of three separate
experiments with the meanþ/-standard deviations depicted. Only
the results with N.1 plus g-thrombin or TRAP-4A were
significantly different than their controls, as analyzed by a
student-t test.
Figure. 6. The synergistic inhibitory effect of histone H1.3
fragments N.1 plus N.2 on g-thrombin-induced platelet aggrega-
tion. The designation of the fragments are as in Figure 1. The final
concentration of N.1 employed was either 0.1 or 0.2 uM plus
1 uM N.2. Each point represents an individual experiment.
Figure 5. The inhibition of g–thrombin-induced platelet aggre-
gation by H1.3 fragments and combination of fragments
compared to full-length histone H1.3. The designation of the
fragments are the same as in Figure 1. All fragments were tested at
a final concentration of 1 uM. Each point represents an individual
experiment.
Inhibition of �-thrombin-induced human platelet aggregation by histone H1subtypes and H1.3 fragments 353
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

experiment in which inhibition of g-thrombin-
induced platelet aggregation was measured after
digesting 0.1 mM N.1 with 0.075 U AP for
0–110 min. Platelets were preincubated with the
modified N.1 for 2 min prior to the addition of the
g-thrombin. Platelet receptors, other than PAR-4,
are not affected by 0.075U AP under these condi-
tions. Interestingly, the capacity of N.1 to inhibit
platelet aggregation increased with the first 10 min of
digestion time and then reached a plateau indicating
that any modification of N.1 by AP was completed by
about 10 min.
MALDI analysis of the AP-treated N.1 fragment
(5 ml 100 mM N.1 plus 1 U AP at 37�C for 90 min)
demonstrated that two N-terminal amino acids (Asp-
Val) were hydrolyzed off and the reaction did not go
any further (Figure 2b, d). This conclusion was
based upon the difference between the digested and
undigested N.1 fragment, which was about 212
daltons (the molecular weight of peptidyl-linked
Asp-Val amino acids), resulting in the 15,025
dalton fragment. Removal of these two amino acids
appears to impact on the conformation of the peptide
resulting in a more active molecule by maintaining
the peptide in a conformation with greater inhibitory
function.
Discussion
The presence of three different thrombin receptors
on human platelets that differentially respond to
varied concentrations of �-thrombin and to �- versus
�- and g-thrombin may have important physiological
regulatory roles in normal and abnormal hemostasis
[6, 13, 14]. We have extended our earlier studies that
demonstrated that histone 1 selectively inhibits
human platelet PAR-4 at the membrane level [14]
by pinpointing the region of one histone H1 subtype
responsible for inhibitory activity to the C-terminal
portion of the molecule. This may serve as a
blueprint for generating therapeutic molecules for
treatment of abnormal hemostasis.
We found that all five subtypes of the recombinant
H1 protein tested were approximately equally active
as selective inhibitors of PAR-4. We further dissected
one subtype, recombinant human H1.3, with respect
to functional domains by creating several fragments
by enzymatic cleavage. Of all fragments tested, only
the C-terminal 15 k N.1 fragment retained inhibitory
activity. This fragment is composed of the complete
C-terminus and the third helix of the globular
domain. The fact that another fragment, GC,
which includes the C-terminus and the whole glob-
ular domain, did not have significant inhibitory
activity, suggests that the other two helices of the
globular domain disrupt the conformation of the
C-terminus required for functional activity. It is also
possible that these two other helical regions are
exposed in such a way to prevent the third helix-C-
terminal structure, found in N.1, from interacting
with the PAR-4 receptor. The conformation and/or
amino acid sequence of the third helix may also have
a slight impact on the overall conformation or
binding properties of N.1, since removal of the two
N-terminal amino acids of the N.1 fragment further
increased its inhibitory activity.
The N-terminal amino peptidase (AP) from
Aeromonas proteolytica generally will not hydrolyze
aspartic or glutamic residues. The fact that the newly
generated N-terminal aspartate is hydrolyzed off
from the N.1 fragment indicates that it may have
been modified during hydrolysis of the parent H1.3
molecule. The AP was then able to rapidly hydrolyze
the valine residue and stopped at the newly generated
N-terminal glutamic acid residue which is resistant to
further hydrolysis. The alteration of the N-terminal
region of N.1 seems to play a significant role in
increasing the inhibitory potential of the H1.3
fragment. However, any structural modification in
the N.1 fragment due to removal of the Asp-Val that
may have occurred could not be detected by circular
dichroism (data not shown), which would indicate
little, or no alteration in the tertiary structural
elements. The C-terminal N.1 fragment contains
the amphipathic helix III region of the globular
domain and the unstructured C-terminal tail. The
helix III region is thought to be the major DNA
binding region of the H1 molecule. Also, the
‘‘unstructured’’ C-terminal region appears to be
able to assume �-helical structures in the appropriate
environment (such as the electrostatic environment
of DNA) [15]. It may be that these properties
facilitate binding of the N.1 fragment to the
platelet membrane/PAR-4 receptor as well as to the
g-thrombin molecule. As noted in our previous
publication [14], the crystalline structure and bind-
ing properties of �-thrombin and g-thrombin are
Figure. 8. The inhibition of g-thrombin-induced platelet aggre-
gation by the histone H1.3 fragment N.1 pre-treated with
aminopeptidase for varying times. This is a representative exper-
iment with 0.1 uM N.1 pre-treated with 0.075 U aminopeptidase
for 0–110 min (‘‘Time’’ on X-axis) at 37�C.
354 G. Soslau et al.
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

quite distinct and likely account for some, or all of
the observed differential inhibition by N.1 of these
two forms of thrombin.
A second DNA binding region resides in the
globular domain associated with the N-terminal
fragment N.2 [15]. While there is little or no
inhibitory effect of N.2 by itself, it is interesting
that the addition of low levels of N.1 plus N.2 result
in a synergistic inhibitory response. This was not
observed with combinations of N.2 plus other H1.3
fragments. It is possible that the synergistic response
results from a conformational change induced by the
association of a region of N.2 with N.1 that increases
the exposure of the N.1 binding site for PAR-4 or a
membrane region near PAR-4 and/or a region on the
g-thrombin molecule. This enhanced binding may
mimic conformational changes of N.1 induced by AP
treatment that removes a charged Asp and a
lipophilic Val group.
It is clear that the N.1 fragment, like its parent H1
molecule, inhibits g-thrombin-induced platelet
aggregation at the PAR-4 receptor level. This con-
clusion is based upon the almost complete inhibition
by N.1 of the TRAP-4A-induced platelet aggregation
reaction. However, as previously observed [14],
the TRAP-4A reaction was not as fully sensitive as
the g-thrombin reaction and may indicate that N.1
is inhibiting aggregation by interacting both with the
g-thrombin molecule and the PAR-4 receptor.
However, a direct effect of H1.3 on the thrombin
molecule is not indicated by our studies with a
chromogenic thrombin substrate. It is possible that
there is, in fact, a small direct H1.3/N.1 effect on
thrombin that can not be detected by this system. A
small substrate molecule may still be able to interact
with the catalytic site on thrombin while interactions
with PAR-4 would be spatially blocked by the large
H1.3/N.1 bound to/associated with the thrombin
molecule. Inhibition was specific for the PAR-4
receptor and N.1 had no significant effect on any of
the other platelet receptors tested. The slight inhibi-
tion at the ADP receptor previously observed with
very high levels (10�) of H1(14) were not detected
here since platelets were preincubated with N.1 for
only 4 min versus the 30 min in our previous studies
and the high concentrations of N.1 compared to H1
were not used. The 4 min time interval was found to
give maximal levels of inhibition.
Much work remains to fully understand what
portion(s) of the H1 molecule selectively binds
to, and inhibits the human PAR-4 receptor and/or
the g-thrombin molecule. It would also be interesting
to compare the effect of N.1 fragments generated
from different H1 subtypes. It is possible that the
variable C-terminal region modifies/regulates the
binding/inhibitory potential of the ‘‘invariant’’ third
helical region. It is unlikely that H1, released into the
circulation from degraded cells, would reach a
sufficient concentration to have a significant clinical
impact on hemostasis. However, the full elucidation
of what sequence(s) of the molecule selectively binds
to and inhibits g-thrombin-induced platelet aggrega-
tion at PAR-4 may serve as a paradigm for the
production of drugs that have clinical significance in
regulating hemostasis at this receptor site.
Declaration of interest: The authors report no
conflicts of interest. The authors alone are respon-
sible for the content and writing of the paper.
References
1. Davie EW, Fujikawa K, Kisiel W. The coagulation cascade:
initiation, maintenance, and regulation. Biochemistry 1991;
30:10363–10370.
2. Boissel J-P, Le Bonniec B, Rabiet M-J, Labie D, Elion J.
Covalent structure of � and g autolytic derivatives of human
�-thrombin. J Biol Chem 1984;259:5691–5697.
3. Chang JY. The structures and protelytic specifities of
autolysed human thrombin. Biochem J 1986;240:797–802.
4. Vu TK, Hung DT, Wheaton VI, Coughlin SR. Molecular
cloning of a functional thrombin receptor receals a novel
proteolytic mechanism of receptor activation. Cell 1991;
64:1057–1068.
5. Kahn ML, Zheng YW, Huang W, Bigornia V, Zheng D, Moff
S, Farese RV, Tam C, Coughlin SR. A dual thrombin
receptor system for platelet activation. Nature
1998;394:690–694.
6. Soslau G, Class R, Morgan DA, Foster C, Lord ST,
Marchese P, Ruggeri ZM. Unique pathway of thrombin-
induced platelet aggregation mediated by glycoprotein Ib.
J Biol Chem 2001;276:21173–21183.
7. Ramakrishnan V, DeGuzman F, Bao M, Hall SW, Leung LL,
Phillips DR. A thrombin receptor function for platelet
glycoprotein Ib-IX unmasked by cleavage of glycoprotein V.
Proc Natl Acad Sci 2001;98:1823–1828.
8. Brass LF. Thrombin and Platelet activation. Chest
2006;124:18S–25S.
9. Jarvis GE, Atkinson BT, Frampton J, Watson SP. Thrombin-
induced conversion of fibrinogen to fibrin results in rapid
platelet trapping which is not dependent on platelet activation
or GPIb. Br J Pharmacol 2003;138:574–583.
10. Soslau G, Favero M. The GPIb-thrombin pathway: Evidence
for a novel role of fibrin in platelet aggregation. J Thromb
Haemost 2004;2:522–524.
11. Adam F, Guillin MC, Jandrot-Perrus M. Glycoprotein Ib-
mediated platelet activation. A signaling pathway triggered by
thrombin. Eur J Biochem 2003;270:2959–2970.
12. Dubois C, Steiner B, Kieffer N, Meyer-Reigner SC.
Thrombin binding to GPIb� induces platelet aggregation
and fibrin clot retraction supported by resting �IIb�3 inter-
action with polymerized fibrin. Thromb Haemost 2003;
89:853–865.
13. Soslau G, Goldenberg SJ, Class R. The three thrombin
receptors on human platelets respond differentially to alpha-,
beta-, and gamma-thrombin. FASEB J 2001;15:A896.
14. Soslau G, Goldenberg SJ, Class R, Jameson B. Differential
activation and inhibition of human platelet thrombin recep-
tors by structurally distinct �-, �- and g-thrombin. Platelets
2004;15:155–166.
15. Hill DA. Influence of linker histone H1 on chromatin
remodeling. Biochem Cell Biol 2001;79:317–324.
16. Simpson RT. Structure of the chromatosome, a chromatin
particle containing 160 base pairs of DNA and all the
histones. Biochemistry 1978;17:5524–5531.
Inhibition of �-thrombin-induced human platelet aggregation by histone H1subtypes and H1.3 fragments 355
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.

17. Thoma F, Koller T, Klug A. Involvement of histone H1 in the
organization of the nucleosome and of the salt dependent
superstructures of chromatin. J Cell Biol 1979;83:403–427.
18. Lantner AL, Turner GA, Longstaff E. Thymus histone
treated BHK21 cells; malignant characteristics in vitro but
reduced tumorigenicity. Europ J Cancer 1974;10:601–609.
19. Svironovsky AI, Levin VI, Shimanskaya TV, Nikolaichick VV,
Zagorskeya LS. The influence of RNA and histone prepara-
tion on some properties of lymphoid cells. Tsitologia
1979;21:1337–1341.
20. Kolodny GM. Histone mediated changes in morphology and
confluent density of cultured cells. J Cell Physiol
1973;81:233–240.
21. Levine AS, Nesbit ME, White JG, Yarbro JW. Effects of
fractionated histones on nucleic acid synthesis in 6C3HED
mouse ascites tumor cells and in normal spleen cells. Can Res
1968;28:831–844.
22. Yan N, Shi Y. Histone H1.2 as a Trigger for Apoptosis.
Nature Structural Biology 2003;10:983–985.
23. Elferink JGR. Cytolytic effect of polylysine on rabbit
polymorphonuclear leukocytes. Inflammation 1985;9:
321–331.
24. Vani G, Devi CSG. Tumor suppressor activity of histone H1
in experimental mammary carcinoma. Indian J Pharmacol
2000;32:210–213.
25. Class R, Lindman S, Fassbender C, Leinenbach H-P, Rawer
S, Emrich JG, Brady LW, Zeppezauer M. Histone H1
suppresses tumor growth of leukemia cells in vitro, ex vitro
and in an animal model suggesting extracellular functions of
histones. Am J Clin Oncol 1996;28:831–844.
26. Cui S, Reicher JS, Matero RB, Albina JE. Activated murine
macrophages induce apoptosis in tumor cells through nitric
oxide dependent or independent mechanisms. Can Res
1994;54:2462–2467.
27. Hiemstra PS, Eisenhauen PB, Harwig SS,
van den Barselaar MT, van Furth R, Lehrer RI.
Antimicrobial proteins of murine macrophages. Infect
Immun 1993;61:3038–3046.
28. Rose FRAJ, Bailey K, Keyte JW, Chan WC, Greenwood D,
Mahida YR. Potential role of epithelial-derived histone H1
proteins in innate antimicrobial defense in the human
gastrointestinal tract. Infect Immun 1998;66:3255–3263.
29. Monestier M, Fasy TM, Bohm L. Monoclonal anti-histone
H1 autoantibodies from MRL lpr/lpr mice. Molec Immun
1998;66:749–758.
30. Basheer AR, El-Asmar MF, Soslau G. Characterization of
potent platelet aggregation inducer from Cerastes cerastes
(Egyptian sand viper) venom. Biochim Biophys Acta
1995;1250:97–109.
31. Soslau G, Ando A, Floyd L, Hong T, Mathew L, Yen Y.
Effect of low and high dose melagatran and other antithrom-
botic drugs on platelet aggregation. J Thromb Thrombolysis
2008;25:198–203.
32. Izzo A, Kamieniarz K, Schneider R. The histone H1 family:
Specific members, specific functions? Biol Chem
2008;389:333–343.
356 G. Soslau et al.
Plat
elet
s D
ownl
oade
d fr
om in
form
ahea
lthca
re.c
om b
y U
nive
rsity
of
Bat
h on
11/
09/1
4Fo
r pe
rson
al u
se o
nly.







![OPEN ACCESS International Journal of Molecular Sciences...hair growth [2].Platelet-derived growth factor (PDGF) isoforms reportedlyinduce and maintain theanagen phase of the murine](https://static.fdocument.org/doc/165x107/60f85444d7faee31306fdb0e/open-access-international-journal-of-molecular-sciences-hair-growth-2platelet-derived.jpg)