Discovery and Function of a General Core Hormetic Stress ... · by Sublethal Concentrations of...
-
Upload
duongquynh -
Category
Documents
-
view
219 -
download
2
Transcript of Discovery and Function of a General Core Hormetic Stress ... · by Sublethal Concentrations of...

Cell Reports, Volume 17
Supplemental Information
Discovery and Function of a General Core
Hormetic Stress Response in E. coli Induced
by Sublethal Concentrations of Antibiotics
Aurélie Mathieu, Sébastien Fleurier, Antoine Frénoy, Julien Dairou, Marie-FlorenceBredeche, Pilar Sanchez-Vizuete, Xiaohu Song, and Ivan Matic

! 1
Supplemental*Information**Supplemental*Figures*!!
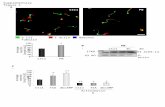
!!!Figure S1, Examples of Expression Profiles of Different Promoter-gfp Fusions During
Growth With or Without 50% MIC of Ampicillin, related to Table 1.
(LB): liquid LB medium. (Amp): liquid LB medium with 50% MIC of ampicillin. Expression
profiles of (A) rpsM promoter-gfp fusion, (B) tnaC promoter-gfp fusion, (C) mlc promoter-
gfp fusion, and (D) acnB promoter-gfp fusion. Fluorescence is expressed in arbitrary units
(A.U). Data are presented as the mean (±SD) values from three independent experiments. For
more information, see the Extended Experimental Procedures.
0.2 0.3 0.40
2000
4000
6000
8000
OD600nm
P mlc-G
FP s
igna
l (A.
U)
LB Amp
0.2 0.3 0.40
2000
4000
6000
8000
10000
12000
OD600nm
P acnB
-GFP
sig
nal (
A.U
)
LB Amp
0.2 0.3 0.4
10000
20000
30000
40000
50000
OD600nm
P rpsM
-GFP
sig
nal (
A.U
)
LBAmp
A
0.2 0.3 0.40
10000
20000
30000
OD600nm
P tnaC
-GFP
sig
nal (
A.U
)
LB Amp
B
C D
rpsM tnaC
mlc acnB

! 2
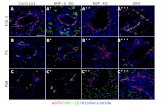
Figure S2. ROS Production and Oxidative Stress Response in Cells Exposed to 50%
MIC of Ampicillin, related to Table 1.
The following strains were used: MG1655 (wt) and isogenic derivative strains containing
reporter plasmids from the promoter library. (LB): liquid LB medium. (Amp): liquid LB
medium with 50% MIC of ampicillin. The vertical dashed line indicates the time of the
growth shift for treated relative to untreated cells. (A) ROS production was measured using a
flow cytometer and dihydrorhodamine 123 dye (DHR). For each time point, the fold increase
relative to the background signal is presented. (B) Expression profiles of sodA, soxS, ahpC
and oxyR promoter-gfp fusions during growth with or without 50% MIC of ampicillin. GFP
fluorescence was quantified using a flow cytometer at OD600≈0.3. For each strain, the fold
increase relative to the untreated corresponding cells is presented. An upregulation of all of
these gene promoters confirmed that ROS production increases in treated cells. Data are
presented as the mean (±SD) values from three independent experiments.
!!!
200 400 600 8000
2
4
6
Time (min)
Fold
incr
ease
in D
HR
sig
nal
rela
tive
to b
ackg
roun
d si
gnal
A
B
wt, LBwt, Amp
P sodA-gfp
P soxS-gfp
P ahpC-gfp
P oxyR-gfp
0
2
4
6
8
Fold
incr
ease
in G
FP s
igna
l re
lativ
e to
unt
reat
ed s
train
s

! 3
Figure S3. β-galactosidase Activity and Stringent Response Induction in Populations
Exposed to 50% MIC of Different Antibiotics, related to Figure 3.
(A) β-galactosidase activity during growth. The MG1655 strain was used. (LB): liquid LB
medium. LB with 50% MIC of trimethoprim (Tmp), gentamicin (Gm), streptomycin (Strp)
and norfloxacin (Nor). The β-galactosidase activity was measured during growth in medium
supplemented with IPTG. For each condition, bars represent β-galactosidase activity, which is
expressed in Miller units (M.U), between OD 3 and 1 or (+) between OD 2.5 and 0.5 for
norfloxacin treatment because norfloxacin-treated populations did not grow to OD 3. t-tests
were performed to compare treated and untreated conditions: * p <0.05 and ** p<0.005 (B)
0 2 4 60
5000
10000
15000
20000
25000
OD600nm
P hisG
-mKa
te s
igna
l (A.
U)
LBAmp
0.0 0.2 0.4 0.6 0.80
5000
10000
15000
20000
25000
OD600nm
P1rrn
B-G
FP s
igna
l (A.
U)
LBStrp 2Strp 2.5
0.0 0.2 0.4 0.6 0.80
5000
10000
15000
20000
25000P1
rrnB-
GFP
sig
nal (
A.U
)
OD600nm
LBGm 0.5Gm 1
0.0 0.2 0.4 0.6 0.80
5000
10000
15000
20000
25000
OD600nm
P1rrn
B-G
FP s
igna
l (A.
U)
LBNor 0.04Nor 0.05
0.0 0.2 0.4 0.6 0.80
5000
10000
15000
20000
25000
OD600nm
P1rrn
B-G
FP s
igna
l (A.
U) LB
Tmp 0.05Tmp 0.1
B
CLB
TmpGm
StrpNor
+0
5
10
15
Rat
io o
f β-g
alac
tosi
dase
(M.U
)ac
tivity
bet
wee
n O
D 1
and
3
**
** **
A

! 4
Expression profiles of the hisG promoter-mKate fusion during growth. (LB): in liquid LB
medium. (Amp): in LB medium with 50% MIC of ampicillin. The vertical dashed line
indicates the time of the growth shift for treated cells. (B) Expression profiles of the rrnB P1
promoter-gfp fusion during growth in LB medium (LB) and in LB with sub-MICs of the
following antibiotics: norfloxacin (Nor) at 0.04 and 0.05 µg/ml, gentamicin (Gm) at 0.5 and 1
µg/ml, trimethoprim (Tmp) at 0.05 and 0.1 µg/ml and streptomycin (Strp) at 2 and 2.5 µg/ml.
Data are presented as the mean (±SD) values from three independent experiments.

! 5
!!Figure S4. Induction of the RpoS Regulon in Populations Exposed to 50% MIC of
Ampicillin, related to Figure 4D.
(LB): liquid LB medium. (Amp): liquid LB medium with 50% MIC of ampicillin. The
vertical dashed line indicates the time of the growth shift for treated cells. Expression of the
PadaZ fusion in the wt strain (wt) and isogenic derivative strains was evaluated by measuring
the β-galactosidase activity, which is expressed in Miller units (M.U). (A) PadaZ expression
during growth and (B) at OD600=3. Bars represent β-galactosidase activity calculated as
follows: mutant (Amp - LB)/wt (Amp - LB). Data are presented as the mean (±SD) values from three
independent experiments. See also Table S6. !! !
0 1 2 3 4 50
20
40
60
80
100
OD600nm
Pada
Z ex
pres
sion
β-
gala
ctos
idas
e ac
tivity
(M.U
)
ΔrpoS, LBΔrpoS, Amp
wt, Ampwt, LB
A
0.0 0.5 1.0 1.5
ΔrsdΔiraMΔiraDΔiraPΔrssBΔcspEΔcspCΔdeaDΔrcsFΔrcsDΔrcsCΔrcsBΔrcsAΔhfqΔcrpΔuvrYΔbarAΔarcAΔrpoS
wt
PadaZ expression at OD600nm= 3 (M.U)Mutant (Amp - LB) / wt (Amp - LB)B

! 6
!
!Figure S5. Media pH Variations, Production of Extracellular H2S and Growth Kinetics
of Populations Exposed to 50% MIC of Ampicillin, related to Figure 4E.
The following strains were used: PadaZ (wt) and the isogenic derivative ΔtnaA and ΔaspC
strains. (LB): liquid LB medium. (Amp): liquid LB medium with 50% MIC of ampicillin.
(pyr): with 20 mM pyruvate. (A) pH variations during growth in LB with and without
ampicillin. Growth kinetics of (B) the wt strain and (C) the ΔtnaA mutant strain grown in LB
with or without ampicillin, supplemented or not with pyruvate. (D) Extracellular H2S
production. Strains were grown in LB with or without ampicillin. Paper strips show a brown
or black stain as a result of their reaction with gaseous extracellular! H2S. The strips were
scanned, and grey levels were quantified with ImageJ software. (%): shows the H2S
production relative to the wt in LB. Data are presented as the mean (±SD) values from three
independent experiments.!!! !
0 1 2 36.5
7.0
7.5
8.0
OD600nm
pH
wt, LBwt, AmpΔtnaA, LBΔtnaA, Amp
50 100 150 200 250 300 3500
1
2
3
4
Time (min)
OD
600n
m
wt, LBwt, Ampwt, LB + pyrwt, Amp + pyr
50 100 150 200 250 300 3500
1
2
3
4
Time (min)
OD
600n
m
ΔtnaA, LB + pyrΔtnaA, Amp + pyr
ΔtnaA, LBΔtnaA, Amp
BA
C D!"# !"#$%&#
'(#
!"#$%&
!$'()&
)(*+,-.# /01,+#2.01#
!" #$$" %#"&"'%"
!" %("&"'%" ')"&"#*"
!" %)"&"%" ))"&"''"

! 7
!!
!!
!!!!"
"
#"
"
$"
""

! 8
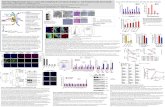
Figure S6. Cell Growth in LB With or Without 50% MIC of Ampicillin in a
Microfluidic Device, related to Movie S1 and Discussion.
The MG1655 strain was used. The microfluidic device is a series of growth channels through
which the LB medium with 50% MIC of ampicillin is passed at a constant rate (Wang et al.,
2010). Images were acquired every 5 min. (A) Temporal montage of images in a single
growth channel. One end of the growth channel is closed (top of the image). The open end of
the channel is at the bottom of the image. Individual cells were segmented. Normal-sized cells
are marked by yellow rectangles. Filamentous cells are marked by purple rectangles. Cells
were defined as filamentous when their length was twice as long as the median length of all
cells. Lines indicate cell lineages. This panel shows that all of the cells grow and divide.
Filamentous cells continue to produce viable progeny. (B) Changes in the progeny cell length
of the first cell, which is trapped at the closed end of the growth channel, over time (solid
line). Filamentous cells produce normal-sized cells, which push the filamentous cell down the
channel and continue to produce normal-sized cells. The filamentous cell, which is now at the
bottom of the channel, continues to produce normal-sized cells (dashed line). (C) The changes
in the generation time of the first cell, which is trapped at the closed end of the growth
channel, over time. Therefore, despite the fact that treated cells were longer than untreated
cells, which may be due to antibiotic-induced morphological alteration, they maintained the
same division rates as untreated cells. We also observed that some rare cells were
filamentous. Nevertheless, filamentous cells continued to produce viable progeny, i.e., cells
budding from the filamentous cells have same size and have same growth rates as other, non-
filamentous cells.
Using this microfluidic device, we measured size and division time of cells growing in
LB without (NT) or with (T) 50% MIC of Ampicillin. For the measurement of the cell size,
we analyzed 28,788 NT and 22,465 T cells. The median of the cell size was 3.71 µm for NT
and 4.61 µm for T cells. Mean ± SD of the cell size was 3.83 ± 1.05 µm for NT and 4.83 ±
1.29 µm for T cells. For the measurement of the division time, we analyzed 5,699 NT and
4,795 T cells. The median of the division time was 15 min for NT and 15 min for T cells.
Mean ± SD of the division time was 16.4 ± 4.21 min for NT and 15.9 ± 3.99 min for T cells.
Therefore, although the treated cells were longer than untreated cells, which may be due to
antibiotic-induced morphological alteration, they maintained the same median division rate as
untreated cells during the experiment.

! 9
!!Movie S1. Monitoring Stringent Response Induction in Cells Growing in a Microfluidic
Device, related to Figure 3D.! For this experiment, we used the MG1655 strain carrying a P1rrnB::gfpunstable reporter fusion.
The left column panels show phase contrast images. The right column panels show
fluorescence images. The upper row panels show cells growing, first in LB (red) for 2.5 h,
then in the conditioned LB (green). The lower row panels show cells growing, first in
LB+50% MIC of ampicillin (red) for 2.5 h, then in the conditioned LB+50% MIC of
ampicillin (green). The time point when media were changed is indicated by the change in the
color of labels from red to green. Conditioned media are cell-free supernatants obtained by
filtering cultures grown for 6 h without (LB) or with ampicillin (LB+Amp) in batch cultures.
Images were acquired every 15 min.
When the LB supply is constant, cells growing with or without ampicillin are bright because
the rrnB gene P1 promoter is highly active (red). However, when cells are exposed to
conditioned media, they slow growth, shrink in size and loose fluorescence (green). Time-
lapse films show that the treated cell fluorescence loss is faster than untreated cells. This
confirms that amino acid depletion is faster in batch cultures with ampicillin compared with
untreated conditions. See also Figure S6.

! 10
Supplemental*Tables*
*
*
WT
!LB Amp
Time (h)
ATP (µM)
ADP (µM)
AMP (µM) AEC ATP
(µM) ADP (µM)
AMP (µM) AEC AEC
Ratio
2 28.28 96.02 46.82 0.45 25.35 94.92 59.44 0.41 0.91 ± 3.94 ± 16.06 ± 4.87 ± 0.02 ± 4.51 ± 13.77 ± 8.56 ± 0.03 ± 0.04
3 52.58 196.25 127.80 0.40 62.93 209.29 ± 13.55
143.17 ± 19.97
0.41 1.01 ± 2.42 ± 22.37 ± 15.26 ± 0.03 ± 6.17 ± 0.02 ± 0.10
4 299.52 ± 59.85
860.71 1158.39 ± 113.13
0.31 214.25 ± 20.91
538.11 ± 21.54
459.81 ± 33.91
0.40 1.27 ± 58.36 ± 0.01 ± 0.03 ± 0.08
5 672.90 ± 0,06
1402.75 ± 135.56
1915.21 ± 162.69
0.34 526.78 ± 19.63
964.19 ± 79.40
859.19 ± 83.83
0.43 1.25 ± 0.05 ± 0.03 ± 0.20
ΔtnaA
!LB Amp
Time (h)
ATP (µM)
ADP (µM)
AMP (µM) AEC ATP
(µM) ADP (µM)
AMP (µM) AEC AEC
Ratio
2 27.21 87.35 48.28 0.44 20.10 78.64 45.62 0.42 0.95 ± 2.11 ± 14.00 ± 3.37 ± 0.03 ± 1.37 ± 12.97 ± 5.11 ± 0.01 ± 0.07
3 60.33 193.28 ± 15.02
101.61 ± 5.78
0.44 64.04 ± 2.61
204.39 ± 4.27
139.93 ± 15.17
0.41 0.92 ± 1.35 ± 0.01 ± 0.02 ± 0.04
4 311.71 ± 22.35
887.23 ± 55.34
1007.4 ± 38.18
0.34 173.10 ±18.08
476.24 ± 29.61
261.98 ± 24.61
0.45 1.32 ± 0.02 ± 0.06 ± 0.17
5 862.02 ±114.36
1344.6 ± 218.50
1125.2 ± 78.74
0.46 621.10 ±101.1
918.40 ±119.72
558.11 ± 66.27
0.51 1.12 ± 0.02 ± 0.06 ± 0.10
*
Table S1. ATP, ADP, AMP Quantification by HPLC and Adenylate Energy Charge
Values (AEC) for Treated (Amp) and Untreated (LB) Wild-Type and tnaA Mutant
Cells, related to Figure 1B. Data are presented as the mean (±SD) values from three
independent experiments.
*
*
*
*
*
*
*
*
*

! 11
*
*
*
*
* LB Amp
Area Under Curve (AUC) Time (h) 3 4 5 3 4 5
Val 4.69E+07 ±1.67E+06
4.84E+07 ±2.71E+06
7.79E+07 ±5.40E+06
3.91E+07 ±2.35E+06
3.99E+07 ±2.24E+06
5.11E+07 ±5.29E+06
Leu 6.54E+06 ±3.42E+05
1.04E+07 ±6.72E+05
1.88E+07 ±9.71E+05
6.14E+06 ±3.52E+05
9.26E+06 ±9.79E+05
1.45E+07 ±9.68E+05
Lys 2.09E+07 ±4.24E+05
2.46E+07 ±9.94E+05
2.61E+07 ±1.37E+06
1.79E+07 ±1.20E+06
1.75E+07 ±9.33E+05
2.00E+07 ±2.22E+06
Met 4.58E+06 ±1.69E+05
4.73E+06 ±8.70E+04
6.63E+06 ±8.85E+04
3.96E+06 ±1.07E+05
3.41E+06 ±8.53E+04
3.34E+06 ±4.74E+05
Tyr 2.92E+06 ±7.33E+04
3.86E+06 ±2.49E+05
5.62E+06 ±7.36E+04
2.26E+06 ±2.31E+05
3.30E+06 ±2.61E+05
4.12E+06 ±2.63E+05
Gln 3.15E+06 ±7.19E+04
2.30E+06 ±1.36E+05
2.77E+06 ±1.10E+05
2.78E+06 ±1.19E+05
1.56E+06 ±1.23E+05
1.70E+06 ±1.72E+05
Glu 4.86E+07 ±3.26E+06
4.92E+07 ±1.09E+06
8.09E+07 ±1.77E+06
3.65E+07 ±2.88E+06
3.96E+07 ±2.24E+06
5.21E+07 ±3.76E+06
Asp 6.75E+07 ±4.09E+06
7.83E+07 ±5.70E+06
9.56E+07 ±2.46E+06
6.49E+07 ±3.59E+05
5.87E+07 ±4.66E+06
5.90E+07 ±1.62E+06
Thr 5.21E+06 ±1.67E+05
5.71E+06 ±1.45E+05
9.79E+06 ±7.08E+05
4.70E+06 ±2.95E+05
4.83E+06 ±3.37E+05
5.88E+06 ±9.48E+05
Asn 7.24E+05 ±5.49E+04
8.18E+05 ±2.42E+04
1.28E+06 ±2.11E+05
6.83E+05 ±4.85E+04
6.07E+05 ±5.49E+04
7.83E+05 ±8.00E+04
Ile 1.76E+07 ±2.11E+06
1.91E+07 ±7.17E+05
2.26E+07 ±2.71E+06
1.43E+07 ±1.68E+06
1.42E+07 ±2.00E+06
1.58E+07 ±2.64E+06
His 1.01E+06 ±5.04E+04
1.02E+06 ±9.03E+04
1.04E+06 ±1.29E+05
9.37E+05 ±7.15E+04
6.91E+05 ±9.03E+04
8.78E+05 ±1.29E+05
Ala 5.80E+06 ±1.21E+05
5.09E+06 ±1.70E+05
3.65E+06 ±2.52E+05
5.06E+06 ±6.53E+05
4.36E+06 ±5.78E+05
3.18E+06 ±3.14E+05
*
Table S2. Quantification of the Intracellular Concentration of Thirteen Amino Acids by
HPLC in Treated (Amp) and Untreated (LB) Wild-Type Cells, related to Figure 1E. Data
are presented as the mean (±SD) values from three independent experiments.
*
*
*
*
*
*
*
*

! 12
*
*
*
*
*
*
*
*
! ! Relevant!genotype!
! ! PhtpG! PrpoH!!!!!!!!Stressor!family! Stressor! wt! wt!
! No!stress!! 5! 5!
Antibiotics!
Ampicillin! +! +!
Gentamicin! +! +!
Norfloxacin! +! +!
Streptomycin! +! +!
Tetracyclin! +! +!
Trimethoprim! +! +!
Alkylating!agent! MMS! +! +!
Oxidizing!agent! H2O2! +! +!
Low!pH! Acetic!acid!! +! +!
Metal! Copper!!! +! +!
Table S3. Induction of the RpoE- and RpoH-Controlled Reporters by Sub-Inhibitory
Doses of Different Stressors in the Wild-Type Strain, related to Table 2.
Induction of the RpoH-controlled PhtpG-lacZ and RpoE-controlled PrpoH-lacZ reporters was
monitored using a disk diffusion assay on LB plates supplemented with X-Gal. Induction was
visualized as blue rings close to the growth inhibition zone (see Figure 4A as example). (+):
blue rings. (-): no blue rings. (MMS): methyl methanesulfonate. (H2O2): hydrogen peroxide.
(Copper): copper (II) chloride.
* *

! 13
**
! Type!of!Stressors!
! Antibiotic! Low!pH! Alkylating!
! Bactericidal! !Bacteriostatic! ! !
Genes! Amp! !Gm! Nor! Cm! !!Acetic!Acid! MMS!acnB% !! !! !! !! !! !!cysP% !! !! !! !! !! !!deoC% !! !! !! !! !! !!entF% !! !! !! !! !! !!erpA% !! !! !! !! !! !!evgA% !! !! !! !! !! !!htpX% !! !! !! !! !! !!mdoB% !! !! !! !! !! !!mlc% !! !! !! !! !! !!msrB% !! !! !! !! !! !!pheL% !! !! !! !! !! !!pyrB% !! !! !! !! !! !!pyrG% !! !! !! !! !! !!rpsM%% !! !! !! !! !! !!rsd% !! !! !! !! !! !!ssb% !! !! !! !! !! !!tig% !! !! !! !! !! !!tnaC% !! !! !! !! !! !!ybiH% !! !! !! !! !! !!ydjN% !! !! !! !! !! !!yedP% !! !! !! !! !! !!ygaZ% !! !! !! !! !! !!yheO% !! !! !! !! !! !!yhhQ% !! !! !! !! !! !!yniA% !! !! !! !! !! !!
Table S4. Expression of the 25 Promoters in Cells Treated With Sublethal
Concentrations of Ampicillin, Gentamicin, Norfloxacin, Chloramphenicol, Acetic Acid
and MMS, related to Figure 2.
The 25 promoters were selected because they were significantly differentially expressed in
cells treated with 50% MIC of ampicillin, norfloxacin and gentamicin relative to untreated
controls. Ampicillin (1 µg/ml), gentamicin (0.5 µg/ml), norfloxacin (0.04 µg/ml),
chloramphenicol (0.5 µg/ml), acetic acid (0.006 M) and MMS (0.04 %). (Blue): upregulated.
(Red): downregulated. (White): like untreated control.

! 14
!!
!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!Competitive!Index!(CI)!
Medium! LB! Amp!
Relevant!genotype! Mean! SD! Mean! SD!
ΔgalS% 0.845! 0.062! 0.723! 0.102!ΔacnB% 0.217! 0.095! 0.059! 0.026!ΔtnaA% 0.234! 0.103! 0.042! 0.020!
!!!!!!!Table S5. Fitness of the acnB and tnaA Mutants Relative to the Wild-Type Strain in the
Presence or Absence of 50% MIC of Ampicillin, related to Table 1.
MG1655 and isogenic derivative mutants resistant to kanamycin were inoculated at an
approximately 1:1 initial ratio in LB medium supplemented with or without 1 µg/ml of
ampicillin and incubated for 16 h at 37°C. The competitive index (CI) was calculated for each
competition. Data are presented as the mean (±SD) values from three independent
experiments. The galS mutant strain was randomly chosen and used to evaluate the possible
fitness cost of the kanamycin resistance cassette. !! !

! 15
β5galactosidase!activity!(M.U)!
!!!!!!!!!! OD600nm=3!
Strains!! LB! Amp!
wt! 34.4!±!6.4! 62.0!±!2.1!ΔrpoS% 2.5!±!0.2! 3.6!±!0.7!ΔarcA% 26.0!±!3.8! 52.4!±!7.0!ΔbarA% 20.5!±!7.3! 43.0!±!9.9!ΔuvrY% 23.8!±!9.6! 41.6!±!12.5!Δcrp% 33.0!±!11.3! 47.8!±!7.6!Δhfq*% 2.3!±!1.5! 3.9!±!2.4!ΔrcsA% 23.4!±!2.7! 37.1!±!7.0!ΔrcsB% 19.7!±!2.4! 37.3!±!3.8!ΔrcsC% 70.5!±!17.3! 88.0!±!21.8!ΔrcsD% 32.3!±!2.7! 57.8!±!3.9!ΔrcsF% 37.2!±!5.1! 54.4!±!5.7!ΔdeaD% 22.9!±!5.7! 37.5!±!3.9!ΔcspC% 47.0!±!13.8! 67.9!±!2.4!ΔcspE% 19.8!±!4.0! 36.5!±!5.6!ΔrssB% 55.0!±!6.1! 99.0!±!15.0!ΔiraP% 21.8!±!5.2! 28.4!±!5.5!ΔiraD% 27.2!±!1.2! 51.5!±!9.2!ΔiraM% 21.5!±!6.1! 33.4!±!4.7!Δrsd% 23.4!±!4.3! 31.1!±!2.0!ΔtnaA% 22.4!±!2.1! 27.9!±!4.0!ΔaspC% 27.5!±!8.1! 35.3!±!6.8!ΔrelA%spoT% !!!!!!!2.0!±!1.0! 1.4!±!0.8!ΔdksA% 2.1!±!2.0! 3.6!±!2.1!wt!+!indole!(2mM)% 30.9!±!6.7! 47.7!±!12.3!wt!+!NaHS!!(0.2mM)% 33.7!±!0.4! 73.2!±!6.6!wt!+!pyruvate!(20mM)% 32.2!±!5.0! 61.7!±!6.6!wt!+!AA!% !33.3!±!5.9!! 33.7!±!5.5!wt!+!pyruvate!(20mM)!+!AA% 39.5!±!5.3! 57.3!±!1.5!ΔtnaA%+!indole!(2mM)! 19.6!±!2.9! 29.4!±!0.9!ΔtnaA!+!NaHS!(0.2mM)! 29.1!±!3.1! 38.4!±!6.7!ΔtnaA%+!pyruvate!(20mM)! 25.7!±!3.9! 55.9!±!3.5!ΔtnaA!+!AA! 30.4!±!7.3! 30.6!±!7.8!ΔtnaA!+!pyruvate!(20mM)!+!AA!! 31.0!±!1.1! 53.0!±!3.8!! ! !! ! !
Table S6. Genetic Control of RpoS Regulon Induction by 50% MIC of Ampicillin in
Different Strains and Growth Conditions, related to Figures 4D, 4E, and S4B.
All strains used were isogenic derivatives of the PadaZ (wt) strain. The strains were grown in
liquid LB medium without (LB) or with 1 µg/ml of ampicillin (Amp). PadaZ expression was
measured by quantifying the β-galactosidase activity at OD600=3. (M.U): Miller units. (AA):
casamino acids complemented with tryptophan, cysteine, glutamine and asparagine. *Δhfq
has a severe growth defect. Data are presented as the mean (±SD) values from three
independent experiments.!

! 16
*
Strains! Genotype! Sources!
E%coli!MG1655!!! rphM1% U1001!lab!collection!
PadaZ! PadaMlacZ,Y,A%∆lacI::cm! (Gutierrez!et!al.,!2013)!PadaZ%∆rpoS! PadaZ%%∆rpoS::kan! This!study!PadaZ%∆dksA! PadaZ%%∆dksA::kan!! This!study!!PadaZ%∆arcA! PadaZ%%∆arcA::kan!! This!study!PadaZ%∆barA! PadaZ%%∆barA::kan!! This!study!!PadaZ%∆uvrY! PadaZ%%∆uvrY::kan!! This!study!PadaZ%∆crp! PadaZ%%∆crp::kan%! This!study!!PadaZ%∆hfq! PadaZ%%∆hfq::kan! This!study!PadaZ%∆rcsA! PadaZ%%∆rcsA::kan! This!study!!PadaZ%∆rcsB%! PadaZ%%∆rcsB::kan! This!study!PadaZ%∆rcsC!! PadaZ%%∆rcsC::kan! This!study!!PadaZ%∆rcsD%! PadaZ%%∆rcsD::kan! This!study!PadaZ%∆rcsF%! PadaZ%%∆rcsF::kan! This!study!!PadaZ%∆csdA%! PadaZ%%∆csdA::kan! This!study!PadaZ%∆cspC!!! PadaZ%%∆cspC::kan! This!study!!PadaZ%∆cspE%!! PadaZ%%∆cspE::kan! This!study!PadaZ%∆rssB! PadaZ%%∆rssB::kan! This!study!!PadaZ%∆iraP!! PadaZ%%∆iraP::kan! This!study!PadaZ%∆iraM%%! PadaZ%%∆iraM::kan! This!study!!PadaZ%∆iraD! PadaZ%%∆iraD::kan! This!study!PadaZ%∆rsd! PadaZ%%∆rsd::kan! This!study!!PadaZ%∆sseA! PadaZ%%∆sseA::kan! This!study!!PadaZ%∆aspC! PadaZ%%∆aspC::kan! This!study!!PadaZ%∆tnaA! PadaZ%%∆tnaA::kan! This!study!PadaZ%∆relA%∆spoT! PadaZ%%∆relA::FRT%∆spoT::kan! This!study!MG1655!∆acnB! ∆acnB::kan% This!study!MG1655!∆galS! ∆galS::kan% This!study!MG1655!∆tnaA! ∆tnaA%::kan% This!study!MG1655!PhisG]mKate! rphM1!intC::PhisGMmKate]kan! U1001!lab!collection!MG1655!P1rrnB]GFP! rphM1%attB::rrnB!P1]gfpASV! (Shah!et!al.,!2006)!MG1655!L19]EGFP! rplsMEGFPMkan% (Nikolay!et!al.,!2014)!MG1655!PrpoH]lacZ! rphM1!∆lac%rpoHP3::lacZ! (Grace!ED!et!al.,!2015)!MG1655!PHtpG]lacZ! rphM1!∆lacX174%PHtpG::LacZ! (Nonaka!G!et!al.,!2016)!!Plasmids! !! !!
DH5alpha!pCP20!
Δ!(argFMlac)169!,!phi80dlacZ58!(M15),!
glnV44(AS),!λ],!rfbD1,!gyrA96!(NalR),!recA1,!
endA1,!spoT1,!thiM1,!hsd!
(Datsenko and Wanner, 2000)
Promoter!collection!! Promoter!specific]GFP;!kan! (Zaslaver!et!al.,!2006)!
*
Table S7. Strains and Plasmids.

! 17
All PadaZ derivative strains were constructed using P1 transduction of alleles from the Keio
collection (Baba et al., 2006) (except the spoT allele, which was obtained from Christophe
Beloin’s strain collection, Pasteur Institute, Paris, France).
Supplemental Experimental Procedures
Determination of the Minimum Inhibitory Concentration of Antibiotics and Other
Stressors
The MIC was defined as the antibiotic concentration that inhibited growth after 24 h of
incubation in LB liquid medium at 37°C. Cells from fresh overnight cultures were diluted
1/1,000 (v/v) and inoculated into 96-well plates (Greiner Bio-One 96 rounded) containing LB
medium and different concentrations of antibiotics. Microtiter plates were incubated for 24 h
at 37°C in a microplate reader and incubator (Infinite M200 PRO, Tecan). The optical density
(OD) at 600 nm was measured every 10 min.
The MICs for MG1655 strain are: 2 µg/ml for Ampicillin, 0.1 µg/ml for Norfloxacin,
0.2 µg/ml for Trimethoprim, 2 µg/ml for Gentamicin, 5 µg/ml for Streptomycin, 1 µg/ml for
Chloramphenicol, 0.08% for MMS and 0.012 M for Acetic acid. The sub-MICs of antibiotics
and other stressors used for experiments are: 1 – 1.25 µg/ml for Ampicillin, 0.04 – 0.05 µg/ml
for Norfloxacin, 0.05 – 0.1 µg/ml for Trimethoprim, 0.5 - 1 µg/ml for Gentamicin, 2 – 2.5
µg/ml for Streptomycin, 0.5 µg/ml for Chloramphenicol, 0.04% for MMS and 0.006 M for
Acetic acid.
Quantification of the Extracellular H2S
Lead acetate-soaked paper strips (Sigma) show a brown or black PbS stain as a result of the
reaction with H2S. Strips were affixed to the caps of Erlenmeyer flasks, above the level of the
liquid. Cultures were incubated for 15 h. The stained paper strips were scanned, and the grey
level was quantified using ImageJ software.
ROS Measurements by Flow Cytometry and DHR
These measurements were performed using a 96-well plate and robotic system (Freedom Evo,
Tecan). During growth in LB medium, with or without 1 µg/ml of ampicillin, 1 ml aliquots of
cell cultures were collected sequentially. After measuring the OD600, cells were incubated for
30 min in the dark at 4°C in PBS (pH 7.4) containing 1%mM dihydrorhodamine 123 (DHR).
Fluorescence was measured using flow cytometry at 488%nm excitation and 530%nm emission.

! 18
A minimum of 50,000 cells was analyzed for each time point with a maximum of 2,000
events counted per second. The obtained data were corrected by the autofluorescence
measured without DHR.
GFP Fluorescence Measurements by Flow Cytometry
These measurements were performed using a 96-well plate and robotic system (Freedom Evo,
Tecan). Cells were grown in LB medium with or without 1 µg/ml of ampicillin. At OD600=3,
aliquots were collected, and GFP fluorescence was quantified by flow cytometry at 488%nm
excitation and 530%nm emission. A minimum of 50,000 cells was analyzed. For each strain,
after background correction to cells without GFP fluorescence, the fold increase in the GFP
signal was calculated for strains treated with 1 µg/ml of ampicillin relative to their untreated
controls.
Expression Profiles of Various Fluorescence Transcriptional Reporters in Cells treated
with Sublethal Concentrations of Gentamicin, Norfloxacin, Chloramphenicol, Acetic
Acid and MMS
Ninety-six-well plates containing various reporter strains (listed in Table 1) were used to
study the impact of sublethal concentrations of norfloxacin (0.04 µg/ml), gentamicin (0.5
µg/ml), chloramphenicol (0.5 µg/ml), acetic acid (0 .006 M) and MMS (0.04%) on gene
expression profiles. We used the same robotic system, the same experimental procedure and
the same data analysis as those used for the study of the impact of the sub-MIC of ampicillin
(see Experimental Procedures and Extended Experimental Procedure).
Microfluidics Experiment
Cells from the overnight cultures were concentrated 10 × by centrifugation. They were then
loaded into a microfluidic mother machine device (Wang et al., 2010) by diffusion. Fresh
filtered LB medium with or without 1 µg/ml of ampicillin was infused with a syringe pump
(diffusion rate = 2 ml/h). For each experiment, images were acquired for 10 h at 5 min
intervals with the MG1655 strain or 15 min intervals with the MG1655 strain carrying the
P1rrnb::gfpunstable reporter fusion using Metamorph software and a Nikon Eclipse Ti
fluorescence microscope equipped with a motorized stage and a CCD camera (Photometrics
CoolSnap HQ2). To extract information about the cells from the images, each channel
containing cells was cut and pasted on a new image and aligned from left to right according to

! 19
the acquisition time (Figure S6). The resulting image contained cell segmentation information
and was analyzed by ImageJ. The cell lineage was determined by visualizing the arrows
connecting the cells. The division time of one cell is defined by the time gap between the birth
of one cell and the first division of this cell. Filamentous bacteria, defined when the cell size
exceeds twice the length of the median cell size in all the cells in the movie, are indicated by
purple.
Fitness Measurements
Overnight cultures of MG1655 (wt) and mutant Kan (m) strains were diluted to
approximately 104 cells per ml and were inoculated at an approximately 1:1 initial ratio in LB
medium supplemented with or without 1 µg/ml of ampicillin and incubated for 16 h at
37°C. The relative proportion of each strain at t0h and t16h was evaluated by plating dilutions
on selective and non-selective LB agar plates. The competitive index (CI) was calculated as
follows. CI = CFU [(m/wt) t16h / (m/wt) t0h].
Survival of the Lethal Ampicillin Treatment
Overnight cultures were diluted 1/1,000 in LB medium and grown for 3 h until exponential
phase (OD600 ≈ 0.6). Cultures were subsequently diluted 1/1,000 and grown with or without 1
µg/ml of ampicillin for 4 h. Cells were then treated with lethal concentrations of ampicillin
(100 × MIC = 200 µg/ml). After 5 h of treatment, cells were collected by centrifugation and
washed with PBS. The number of colony forming units (CFU) was quantified before and after
antibiotic treatment by plating serial dilutions of cells on LB plates.

! 20
Extended Experimental Procedures
Automated Analysis of the Expression Profiles of the Ampicillin-Treated Cells using the
Library of Fluorescence Transcriptional Reporters
For each of the 21 microplates containing a total of 1,920 promoters, we performed 4
replicate treatments with 1 µg/ml of ampicillin and 4 replicate controls without ampicillin for
the growth kinetics. The OD600 and fluorescence were measured approximately every 37 min.
We denoted t(n) as the time at which the nth data point was measured. For each replicate
microplate, we extracted two time series from the monochromator: optical density and
fluorescence at each time point, which we denoted ODraw (t(n)) and fluoraw (t(n)).
To identify candidate promoters responding to the presence of a sub-inhibitory dose of
ampicillin, we developed a collection of programs performing the following analyses.
For each well of each replicate plate, we subtracted the optical density and fluorescence
of the well containing a promoterless construct from the ODraw (t(n)) and fluoraw (t(n)) and
obtained the corrected optical density and fluorescence, which we denoted OD(t(n)) and
fluo(t(n)).
To compare specific promoter activity between populations of bacteria even when they
had different growth kinetics (due to the presence or absence of ampicillin), we converted the
corrected time-series data extracted from the monochromator (optical density and
fluorescence at each time point, corrected with background) into fluorescence as a function of
optical density. To do so, for each target optical density value x between 0 and 0.5 with an
increment of 0.02, we found the smallest n, such that OD(t(n)) < x < OD(t(n + 1)). We
performed a linear interpolation between fluo (t(n)) and fluo (t(n + 1)) to determine the
fluorescence value associated with optical density x. We denoted fluoi (OD) to be the obtained
series of fluorescence as a function of optical density.
This transformation is logical because we identified the optical density of the growth
state of the bacterial population and the fluorescence of the promoter activity. We thus
considered that a given promoter responded to ampicillin at OD=x when fluoi (x) differed
significantly between the 4 ampicillin-treated replicates and the 4 non-treated replicates. We
stopped the analysis when cells were at stationary phase, i.e., at OD 0.5. We defined fluoi
[n](x) as the fluorescence as a function of optical density for replicate n. To identify
candidates exhibiting such responses to ampicillin, for each value x of optical density between
0 and 0.5 with increment 0.02 (we defined this interval as [0 : 0.02 : 0.5]), we performed a
two-sided unpaired Student’s t-test on fluoi (x) between the ampicillin treatments and the

! 21
controls and defined pv(x) as the p-value obtained at OD = x. In addition to the significance
of the difference in promoter activity between the treatment and the control, we also recorded
the magnitude of this difference, expressed as r(x) = log2(Median n�treatments (fluoi
[n](x))/Median n�controls (fluoi [n](x))).
For each promoter p in the library, we determined whether it exists x � [0 : 0.02 : 1]
such that pv(x) < 0.01 and pv(x + 0.02) < 0.01. If so, we included this promoter in our list of
candidates and recorded the value of the smallest x satisfying the above condition. This value
is the optical density at which this promoter p starts behaving differently in the ampicillin
treatment and in the ampicillin-free control.
Because fluoi (OD) sometimes differed significantly between one of the replicates and
the other three, we automatized the detection of potential outliers using the following test. At
a given optical density x, we computed the median distance to the median, expressed as
Mediann (|Mediann (fluoi [n](x)) − fluoi [n](x)|), which we denoted M (x). We considered that
a replicate n at OD x is an outlier if |fluoi [n](x) − M (x)| > 6 ∗ M (x). In other words, a data
point is an outlier if its distance to the median (of all replicates of the same treatment) is 6
times larger than the median distance to the median.
We extracted the list of all promoters for which one of the replicates contained many
outliers and visualized the graphical representation of fluoi [n] for each replicate. We
manually added to our candidate list the promoters for which there is a strong visual
difference between treatment and control that was not detected due to the presence of outliers.
Next, to eliminate false positives due to experimental noise and multiple testing, we
subjected the set of candidate promoters to a second round of screening. These experiments
were performed exactly as described above, except there were 6 treated and 6 untreated
replicates to increase the statistical power of the algorithm. All the promoters whose activity
was no longer detected as impacted by ampicillin treatment in this second set of experiments
were removed from the list.
Finally, for all candidates, the constructs were assessed by PCR, and truncated
promoters were removed, resulting in a final list of 64 promoters (Table 1 and Figure S1 for
examples).
! !

! 22
Supplemental References
Baba, T., Ara, T., Hasegawa, M., Takai, Y., Okumura, Y., Baba, M., Datsenko, K.A., Tomita,
M., Wanner, B.L., and Mori, H. (2006). Construction of Escherichia coli K-12 in-frame,
single-gene knockout mutants: the Keio collection. Molecular systems biology 2, 2006 0008.
Datsenko, K.A., and Wanner, B.L. (2000). One-step inactivation of chromosomal genes in
Escherichia coli K-12 using PCR products. Proceedings of the National Academy of Sciences
of the United States of America 97, 6640-6645.
Gutierrez, A., Laureti, L., Crussard, S., Abida, H., Rodriguez-Rojas, A., Blazquez, J.,
Baharoglu, Z., Mazel, D., Darfeuille, F., Vogel, J., et al. (2013). beta-Lactam antibiotics
promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity. Nature
communications 4, 1610.
Nikolay, R., Schloemer, R., Schmidt, S., Mueller, S., Heubach, A., and Deuerling, E. (2014).
Validation of a fluorescence-based screening concept to identify ribosome assembly defects
in Escherichia coli. Nucleic acids research 42, e100.
Shah, D., Zhang, Z., Khodursky, A., Kaldalu, N., Kurg, K., and Lewis, K. (2006). Persisters:
a distinct physiological state of E. coli. BMC microbiology 6, 53.
Wang, P., Robert, L., Pelletier, J., Dang, W.L., Taddei, F., Wright, A., and Jun, S. (2010).
Robust growth of Escherichia coli. Curr Biol 20, 1099-1103.
Zaslaver, A., Bren, A., Ronen, M., Itzkovitz, S., Kikoin, I., Shavit, S., Liebermeister, W.,
Surette, M.G., and Alon, U. (2006). A comprehensive library of fluorescent transcriptional
reporters for Escherichia coli. Nat Methods 3, 623-628. !