The role of DNA methylation in mediating the effects of estrogens in oysters
description
Transcript of The role of DNA methylation in mediating the effects of estrogens in oysters

The role of DNA methylation in
mediating the effects of estrogens
in oysters
Mackenzie Gavery & Steven Roberts
School of Aquatic and Fishery Sciences
University of Washington

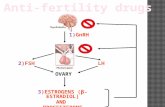
Outline17α ethinylestradiol
(EE2)

Outline
Background
DNA methylation
EDCs & bivalves
Results
Implications
17α ethinylestradiol
(EE2)

GENES (DNA)
TRAITS
color
growth
disease resistance
ENVIRONMENT
Background

GENES (DNA)
EPIGENOME(DNA methylation)
TRAITS
color
growth
disease resistance
ENVIRONMENT
Background

GENES (DNA)
EPIGENOME(DNA methylation)
TRAITS
color
growth
disease resistance
ENVIRONMENT
Background

GENES (DNA)
EPIGENOME(DNA methylation)
TRAITS
color
growth
disease resistance
ENVIRONMENT
Background

Me
C
GC
G

Me
C
GC
G
Gene A
TF X

Me
C
GC
G
Gene A
TF X

Me
C
GC
G
Gene A
TF X

Me
C
GC
G
Gene A
TF X

Reproduction in oysters
Pacific oysters are sequential hermaphrodites
Sex determination has a genetic component, but
influenced by environmental factors

Reproduction in oysters
Nonylphenol
• offspring of exposed
larvae had intersex (Nice et al. 2003)
17α ethinylestradiol (EE2)
• rate of oocyte
development (Andrew 2010)
Estradiol
• induces sex reversal (Mori
1969)
Pacific oysters are sequential hermaphrodites
Sex determination has a genetic component, but
influenced by environmental factors

Hypotheses EE2 exposure will result in phenotypes such as skewed
sex ratios and increased rate of gonad development
DNA methylation patterns will be altered in oysters
exposed to EE2

Hypotheses EE2 exposure will result in phenotypes such as skewed
sex ratios and increased rate of gonad development
DNA methylation patterns will be altered in oysters in
upon exposure to EE2

Estrogen Experiment
500 ng/L EE2: 150 oysters (n=50/tank)
Control: 150 oysters (n=50/tank)

Estrogen Experiment
Control: 150 oysters (n=50/tank)
Day 0 Day 7 Day 60
Samples: histology, gonad tissue
500 ng/L EE2: 150 oysters (n=50/tank)

Estrogen Experiment
Day 0 Day 7 Day 60
500 ng/L EE2: 150 oysters (n=50/tank)
Control: 150 oysters (n=50/tank)
Samples: histology, gonad tissue

Results: Day 60
sex determination
0%
20%
40%
60%
80%
100%
control EE2
Pe
rce
nt o
f To
tal O
yste
rs
Hermaphrodite
Unknown
Male
Female

Results: Day 60
size of females
30
35
40
45
50
55
60
length width
mm
shell size (mm)
control
EE2
17
18
19
20
21
22
23
24
control EE2
gra
ms
total weight (g)

Results: DNA methylation
control 150 oysters (n=50/tank)
Day 0 Day 7 Day 60
EE2 (500ng/L) 150 oysters (n=50/tank)

Results: DNA methylation
control 150 oysters (n=50/tank)
Day 0 Day 7 Day 60
EE2 (500ng/L) 150 oysters (n=50/tank)

MBD-Chip
NimbleGen
3 x 720k Array
compare
DNA methylation
control
EE2
exposed
MBD
MBD
Results: DNA methylation

Results:
45 differentially methylated regions (DMR)
DMRs were located in 38 different genes
Results: DNA methylation

Organism Proteinnames
Didelphisvirginiana 5-hydroxytryptaminereceptor1B
Homosapiens ATP-bindingcassettesub-familyGmember1
Gallusgallus Angiotensin-convertingenzyme
Rattusnorvegicus Neuronalacetylcholinereceptorsubunitalpha-6
Homosapiens Anaphase-promotingcomplexsubunit1
Musmusculus Arrestindomain-containingprotein3
Patinopectensp. Calmodulin
Xenopuslaevis Corticotropin-releasingfactorreceptor2
Homosapiens Carnosinesynthase1
Homosapiens E3ubiquitin-proteinligaseDTX3L
Chlamydomonasreinhardtii Dyneingammachain,flagellarouterarmDaniorerio Elongatorcomplexprotein2
Cerberusrynchops Ryncolin-1
Acomyscahirinus Glutaminesynthetase
Daniorerio Glutaredoxin3
Homosapiens Granulins
Musmusculus TranslationfactorGuf1,mitochondrial
Cydiapomonellagranulosisvirus ApoptosisinhibitorIAP
Musmusculus Interferon-inducedprotein44
Rattusnorvegicus Kelch-likeprotein24
Musmusculus Liprin-beta-1
Musmusculus Low-densitylipoproteinreceptor-relatedprotein6
Homosapiens Unconventionalmyosin-Vb
Pongopygmaeus NADHdehydrogenase[ubiquinone]flavoprotein1,mitochondrialCaenorhabditiselegans Noseresistanttofluoxetineprotein6
Xenopustropicalis PeptidaseM20domain-containingprotein2
Homosapiens 60kDaSS-A/Roribonucleoprotein
Musmusculus Solutecarrierfamily28member3
Macacafascicularis Solutecarrierfamily45member3
Homosapiens ProteintransportproteinSec16ABostaurus Smallintegralmembraneprotein14Xenopuslaevis Srckinase-associatedphosphoprotein2-B
Xenopuslaevis DNAtopoisomerase1Musmusculus tRNApseudouridinesynthaseA,mitochondrialMusmusculus VasorinHomosapiens Vacuolarproteinsorting-associatedprotein13CHomosapiens WASHcomplexsubunit7

Organism Proteinnames
Didelphisvirginiana 5-hydroxytryptaminereceptor1B
Homosapiens ATP-bindingcassettesub-familyGmember1
Gallusgallus Angiotensin-convertingenzyme
Rattusnorvegicus Neuronalacetylcholinereceptorsubunitalpha-6
Homosapiens Anaphase-promotingcomplexsubunit1
Musmusculus Arrestindomain-containingprotein3
Patinopectensp. Calmodulin
Xenopuslaevis Corticotropin-releasingfactorreceptor2
Homosapiens Carnosinesynthase1
Homosapiens E3ubiquitin-proteinligaseDTX3L
Chlamydomonasreinhardtii Dyneingammachain,flagellarouterarmDaniorerio Elongatorcomplexprotein2
Cerberusrynchops Ryncolin-1
Acomyscahirinus Glutaminesynthetase
Daniorerio Glutaredoxin3
Homosapiens Granulins
Musmusculus TranslationfactorGuf1,mitochondrial
Cydiapomonellagranulosisvirus ApoptosisinhibitorIAP
Musmusculus Interferon-inducedprotein44
Rattusnorvegicus Kelch-likeprotein24
Musmusculus Liprin-beta-1
Musmusculus Low-densitylipoproteinreceptor-relatedprotein6
Homosapiens Unconventionalmyosin-Vb
Pongopygmaeus NADHdehydrogenase[ubiquinone]flavoprotein1,mitochondrialCaenorhabditiselegans Noseresistanttofluoxetineprotein6
Xenopustropicalis PeptidaseM20domain-containingprotein2
Homosapiens 60kDaSS-A/Roribonucleoprotein
Musmusculus Solutecarrierfamily28member3
Macacafascicularis Solutecarrierfamily45member3
Homosapiens ProteintransportproteinSec16ABostaurus Smallintegralmembraneprotein14Xenopuslaevis Srckinase-associatedphosphoprotein2-B
Xenopuslaevis DNAtopoisomerase1Musmusculus tRNApseudouridinesynthaseA,mitochondrialMusmusculus VasorinHomosapiens Vacuolarproteinsorting-associatedprotein13CHomosapiens WASHcomplexsubunit7
Gene Ontology (GO Slim) Count
transport 10
cell organization and biogenesis 8
other metabolic processes 7
signal transduction 6
protein metabolism 5
RNA metabolism 5
developmental processes 4
cell cycle and proliferation 3
death 2
stress response 2
cell-cell signaling 1
DNA metabolism 1

Organism Proteinnames
Didelphisvirginiana 5-hydroxytryptaminereceptor1B
Homosapiens ATP-bindingcassettesub-familyGmember1
Gallusgallus Angiotensin-convertingenzyme
Rattusnorvegicus Neuronalacetylcholinereceptorsubunitalpha-6
Homosapiens Anaphase-promotingcomplexsubunit1
Musmusculus Arrestindomain-containingprotein3
Patinopectensp. Calmodulin
Xenopuslaevis Corticotropin-releasingfactorreceptor2
Homosapiens Carnosinesynthase1
Homosapiens E3ubiquitin-proteinligaseDTX3L
Chlamydomonasreinhardtii Dyneingammachain,flagellarouterarmDaniorerio Elongatorcomplexprotein2
Cerberusrynchops Ryncolin-1
Acomyscahirinus Glutaminesynthetase
Daniorerio Glutaredoxin3
Homosapiens Granulins
Musmusculus TranslationfactorGuf1,mitochondrial
Cydiapomonellagranulosisvirus ApoptosisinhibitorIAP
Musmusculus Interferon-inducedprotein44
Rattusnorvegicus Kelch-likeprotein24
Musmusculus Liprin-beta-1
Musmusculus Low-densitylipoproteinreceptor-relatedprotein6
Homosapiens Unconventionalmyosin-Vb
Pongopygmaeus NADHdehydrogenase[ubiquinone]flavoprotein1,mitochondrialCaenorhabditiselegans Noseresistanttofluoxetineprotein6
Xenopustropicalis PeptidaseM20domain-containingprotein2
Homosapiens 60kDaSS-A/Roribonucleoprotein
Musmusculus Solutecarrierfamily28member3
Macacafascicularis Solutecarrierfamily45member3
Homosapiens ProteintransportproteinSec16ABostaurus Smallintegralmembraneprotein14Xenopuslaevis Srckinase-associatedphosphoprotein2-B
Xenopuslaevis DNAtopoisomerase1Musmusculus tRNApseudouridinesynthaseA,mitochondrialMusmusculus VasorinHomosapiens Vacuolarproteinsorting-associatedprotein13CHomosapiens WASHcomplexsubunit7
Gene Ontology (GO Slim) Count
transport 10
cell organization and biogenesis 8
other metabolic processes 7
signal transduction 6
protein metabolism 5
RNA metabolism 5
developmental processes 4
cell cycle and proliferation 3
death 2
stress response 2
cell-cell signaling 1
DNA metabolism 1
ATP-binding cassette protein
Serotonin receptor
Low density lipoprotein receptor
Granulin

Summary
EE2 treatment did not affect
sex ratios, but exposed
females were larger than
controls
DMRs were identified within
1 week of EE2 exposure
Genes with DMRs are
functionally diverse (e.g.
growth, immune, reproduction)

Implications
DNA methylation may play a role in mediating
responses to EDCs in bivalves
Epigenetic marks may provide early indicators of EDC
exposure in aquatic species

Acknowledgements Roberts Lab:
Steven RobertsSamuel White Emma Timmins-SchiffmanClaire EllisBrent VadopalasJake Heare
Taylor Shellfish:
Joth DavisMolly Jackson
Irv Shultz (Battelle PNNL)