Predisposed αβ T cell antigen receptor recognition of MHC and MHC-I like molecules?
Transcript of Predisposed αβ T cell antigen receptor recognition of MHC and MHC-I like molecules?

Available online at www.sciencedirect.com
ScienceDirect
Predisposed ab T cell antigen re
ceptor recognition ofMHC and MHC-I like molecules?Sidonia BG Eckle1, Stephen J Turner1, Jamie Rossjohn2,3,4 andJames McCluskey1,4The diverse ab T cell receptor (TCR) repertoire exhibits versatility
in its ability to generate antigen (Ag) receptors capable of
interacting with polymorphic Major Histocompatibility Complex
(MHC) molecules and monomorphic MHC-I like molecules,
including the CD1 and MR1 families. Collectively, these
evolutionarily related Ag-presenting molecules present
peptides, lipids and vitamin B metabolites for T cell surveillance.
Interestingly, whilst common TCR gene usage can underpin
recognition of these distinct classes of Ags, it is unclear whether
the ‘rules’ that govern abTCR-Ag MHC interactions are shared.
We highlight recent observations in the context of TCR biases
towards MHC and MHC-I like molecules.
Addresses1 Department of Microbiology & Immunology, Peter Doherty Institute for
Infection and Immunity, University of Melbourne, Parkville, Victoria 3010,
Australia2 Department of Biochemistry and Molecular Biology, School of
Biomedical Sciences, Monash University, Clayton, Victoria 3800,
Australia3 Institute of Infection and Immunity, Cardiff University, School of
Medicine, Heath Park, Cardiff CF14 4XN, United Kingdom
Corresponding authors: Rossjohn, Jamie ([email protected])
and McCluskey,
James ([email protected])
4 Joint senior authors.
Current Opinion in Immunology 2013, 25:653–659
This review comes from a themed issue on Immunogenetics and
transplantation
Edited by Miles Davenport and Deborah K Dunn-Walters
For a complete overview see the Issue and the Editorial
Available online 29th August 2013
0952-7915/$ – see front matter, # 2013 Elsevier Ltd. All rights
reserved.
http://dx.doi.org/10.1016/j.coi.2013.07.010
IntroductionThe ab T cell antigen receptor (TCR) can recognize
antigens presented by products of the MHC gene family.
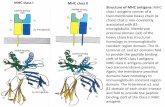
MHC class I (MHC-I) and MHC class II (MHC-II) mol-
ecules are highly polymorphic receptors that bind peptide-
based (p) antigens. MHC-I-like molecules, which are
essentially monomorphic, present non-peptide antigens
to specialized T cell subsets, including Natural Killer T
(NKT) cells, Germline-encoded mycolyl lipid reactive
(GEM) T cells and Mucosal-associated invariant T
www.sciencedirect.com
(MAIT) cells [1–3]. MHC-I like molecules share the same
overall structure as MHC-I, yet exhibit customized chemi-
cal properties suited to bind distinct specific classes of
ligand [4]. For example, the CD1 family and the MHC-I
related molecule MR1, are ideally suited to bind lipid-
based and vitamin B based metabolites respectively [4,5��].Moreover the MHC-I molecule can, in part, accommodate
non-peptide-based ligands, as exemplified by HLA-linked
drug hypersensitivities [6��,7]. The host uses a diverse
repertoire of ab TCRs to engage these various Ag-pres-
entation platforms. Intriguingly however, TCR biases are
observed in the MHC-specific repertoire relevant to pro-
tective immunity and auto-immune-like disorders [8–10].
Moreover, biased TCR usage can underpin CD1d and
MR1-restricted recognition [1]. However, there are also
examples of overlapping TCR gene usage against peptide,
lipid and vitamin B Ags. What factors drive these TCR
biases and what does this mean about the hard wiring of
TCRs towards these distinct Ag-presenting molecules?
Here we review recent advances in the field of TCR
recognition of MHC and MHC-I-like molecules.
T cell repertoire selection and pre-determinedbiasTCR genes rearrange their exons during the develop-
ment of T cells in the thymus, and when combined with
mechanisms such as addition/deletion of nucleotides at
exon junctions, generate a diverse repertoire of Ag-recep-
tors. Although thymic selection was first appreciated in
the context of MHC-restricted immunity [11�], it is now
clear that the ab T cell repertoire is also selected against
MHC-I-like molecules [12,13]. While it has long been a
goal of the field to determine what the rules are that
govern ab TCR recognition of classical pMHC com-
plexes, these have remained elusive. Ironically, more
recent studies of ab TCR recognition of CD1d-Ag and
MR1-Ag complexes have provided key insights into
conserved features of ab TCR recognition. Accordingly,
we shall initially review MHC-restriction and sub-
sequently discuss the emerging findings in CD1d-
restricted and MR1-restricted immunity.
Thymic selection favours T cells that exhibit weak reac-
tivity for self-pMHC and deletes T cells with strong self-
pMHC reactivity, thereby resulting in MHC ‘restriction’
of T cells that are predisposed to recognizing host MHC
[14]. The concept of, and evidence for, germline hard
wiring of the TCR to recognize MHC has intrigued the
Current Opinion in Immunology 2013, 25:653–659

654 Immunogenetics and transplantation
field for decades [15]. Both the MHC and TCRs are
thought to have evolved about 500 million years ago in
cartilaginous fish [16], thus potentially allowing for coe-
volution of TCR germline encoded elements to interact
with MHC molecules. An alternative view postulates that
MHC-restricted T cells are selected during positive
selection from a preselected repertoire with unbiased
specificities [17,18�,19]. Here, TCR specificities are
thought of as behaving in an antibody-like fashion where
thymic selection tunes unbiased reactivity towards an
MHC-affinity window that skews their mature reactivity
towards self-MHC-recognition (Figure 1).
Figure 1
Preselectionrepertoire
Thymicselection
Peripheralrepertoire
Bia
sed
Unb
iase
d
non-MHC reactive T cell
T cell reactive with MHC, with high, intermediate,low affinity/avidity
T cells dying by neglect
Negatively selected T cells
Positively selected T cells
Current Opinion in Immunology
Models of selection from an unbiased versus biased (hard wired) TCR
repertoire. The biased and unbiased preselection repertoires consist of T
cells (grey disks) reactive with MHC. The unbiased preselection
repertoire contains in addition T cells that are non-MHC reactive. T cells
reactive with MHC can have high, intermediate or low affinity/avidity for
MHC. During thymic selection T cells interact with MHC presented in the
thymus on APCs (orange irregular shape) if they are MHC-reactive. Only
T cells that interact with intermediate affinity/avidity with MHC get
selected and appear in the peripheral repertoire. In contrast T cells with
low affinity/avidity or high affinity/avidity die by neglect and get
negatively selected, respectively. In addition, non-MHC reactive T cells
present in the unbiased T cell repertoire will not encounter MHC in the
thymus and die by neglect.
Current Opinion in Immunology 2013, 25:653–659
Hard wiring of TCR reactivity for self-MHC suggests the
existence of some ‘hidden logic’ underpinning pMHC
recognition. Conversely, thymic filtration of unbiased
TCRs to retain only those with a narrow window of
affinity for pMHC does not a priori presuppose stereo-
typic patterns of MHC recognition. To put it another way,
how T cells are guided to interact with MHC molecules
remains unclear. Nevertheless, studies investigating viral
immunity; T cell alloreactivity [20]; co-receptor usage
[18�], TCR bias [9], and the TCR-MHC-I-like axis [1],
have provided some insight, albeit often conflicting, into
this issue of the driving force between the TCR-pMHC
interaction. Front and centre stage to many of these
studies are structurally focused investigations
[1,11�21�,22,23�,24,25].
Structural correlates of MHC restrictionDespite the growing list of distinct TCR-pMHC struc-
tures, the ability to define rules that govern classical
pMHC interactions has proven difficult [21�]. Further,
these investigations involve a limited array of MHC
allotypes, which is an issue as the Human Leucocyte
Antigen (HLA) locus is the most polymorphic region in
the human genome. With that in mind, some general
observations can be made. For example, the TCR sits
atop the pMHC, forming an imperfect interface that is
consistent with the weak affinity of this interaction, with
the Va and Vb domains of the TCR docked over the
MHC-I a2-helix and a1-helix respectively [21�]. As this
approximate docking mode is maintained between differ-
ent TCR-pMHC complexes, it suggests some ‘logic’ to
this key interaction. However, this logic appears not to
rest with conserved structural ‘landing spots’ on the
MHC, such as the previously hypothesized ‘restriction
triad’ [26,27]. Moreover, the precise TCR binding modes
can vary markedly for individual TCR-pMHC complexes
and likely reflect the polymorphic pMHC landscape, and
diversity within the vast TCR repertoire.
Some studies consider the TCR’s germline-encoded
regions (the Complementarity Determining Regions
(CDRs) 1 and 2) as the key components that determine
MHC interactions, while the hypervariable CDR3 loops
‘read-out’ the antigenic cargo [23�,28]. Indeed, conserved
interactions (‘codons’) between two tyrosine residues
(Y46 and Y48) within the CDR2b loop of some Vb8.2+
TCRs and some MHC molecules appear maintained,
throughout evolution (Figure 2a,b) [11�,29]. Indeed,
Y48 was determined to be important in T cell positive
selection of CD4+ and CD8+ T cells, while Y46 was only
important for the selection of CD4+ T cells, suggesting
these residues are key for determining MHC-I vs MHC-
II restriction [30]. Nevertheless, even within the Vb8.2
family, varied docking modes on distinct self-MHC and
allo-MHC molecules are observed [31��,32], leading to a
concept of ‘wobble’, whereby that the same V-region
may bind to alternative pMHC-I molecules in a slightly
www.sciencedirect.com

Rules of ab TCR recognition Eckle et al. 655
Figure 2
Y46
Y48 Y46
Y48
Y46 Y48
Y46
Y48
MHC-I
MHC-II
CD1d
Current Opinion in Immunology
Contrasting modes of CDR2b (Vb8.2) recognition on MHC-I, MHC-II and
CD1d. Top-views of MHC helices with isolated CDR2b loops on MHC-I
(H-2Kb, YAe62 TCR: pink, PDB ID: 3RGV; H-2Kb, 2C TCR: violet, PDB
ID: 2CKB), MHC-II (I-Ab, YAe62 TCR: blue, PDB ID: 3C6O; I-AU, 1934
TCR: cyan, PDB ID: 2PXY; I-Ak, D10 TCR: smudge, PDB ID: 1D9K) and
CD1d (NKT TCR: yellow, PDB ID: 3HES). CDR2b Y46 and Y48 are
displayed in stick format.
different mode (Figure 2a) [28]. Moreover, altered Vb8.2
binding modes induced by differential Va-chain pairing
can also affect the role of these residues encoded within
the CDR2b loop [33�,34]. Whilst the degree of differ-
ence/similarity could be considered semantic, it is import-
ant to note that in MHC-restricted immunity, small (less
than 1 A) shifts in MHC helices can dictate T cell
repertoire selection and protective immunity [35], and
have also been linked to taboo mismatches in transplan-
tation [36]. Thus, the thought-provoking ‘codons’ con-
cept needs to be considered a partial and incomplete
structural explanation of TCR-pMHC hard wiring.
In line with this view, a recent analyses of the unique
TCR-pMHC-I structural database [21�] show that the
www.sciencedirect.com
self-perpetuating CDR1/2-MHC and CDR3-peptide
dogma is incorrect, as the germline encoded residues
frequently contact the peptide, whilst the CDR3 regions
often make crucial contact to the MHC [37,38]. Instead,
synthesis of all the available data indicates that TCRs
have a peptide-centric view that drives the MHC-
restricted interaction. Indeed, the protective immune
response is triggered by a foreign peptide within the
MHC; T cell mediated alloreactivity can be attributed
to peptide-dependent molecular mimicry [39��]; the
impact of germline-encoded TCR polymorphisms are
attributable to peptide-centric charge complementarity
[40]; diverse TCR docking modes atop a common pMHC
can reflect peptide-driven contributions [38,41]; TCR
recognition of ‘super-bulged peptides’ can be markedly
peptide centric, or involve atypical MHC-contact sites
[26,42,43]. Thus, we propose that the peptide acts as the
initial ‘lynch-pin’ for the TCR-pMHC interaction while
the MHC interactions subsequently act like a ‘Velcro’ to
stabilize TCR binding. Clearly, many more unique TCR-
pMHC structures are required to shed further insight into
MHC-mediated immunity.
TCR interactions with MHC-I like moleculesThe innate-like NKT and MAIT cells draw their TCR
from the same ab-TCR repertoire as MHC-restricted T
cells and yet they interact with CD1d and MR1 respect-
ively. Given the evolutionary conserved nature of both
MR1 and CD1d, we could expect studies of TCR-CD1d/
MR1 interactions to provide insights into the hard wiring
or TCR recognition of MHC and MHC-I like molecules.
TCRs expressed by Type I NKT cells generally exhibit
invariant a-chain usage (Va24-Ja18 in humans, Va14-
Ja18 in mice), and a restricted Vb repertoire (Vb11 in
humans and Vb8.2, 7 and 2 in mice) [44]. Interestingly,
despite common Vb8.2 (and orthologous Vb11) usage in
MHC-restricted and CD1d-restricted immunity, the
location of the corresponding CDR2b regions atop these
MHC and MHC-like molecules is markedly different
(Figures 2c and 3) [45]. This suggests that there is
inherent flexibility within the TCR gene segments to
accommodate alternate binding modes on distinct MHC-
Ag landscapes. What is more astounding is that despite
the varied Type I NKT TCR gene usage [46–49], and a
variety of chemically distinct lipid-based antigens recog-
nized by Type I NKT cells [50], the mode of Type I
NKT TCR-CD1d-Ag docking is conserved – namely it
adopts a parallel docking mode above the F0-pocket of
CD1d (Figure 3) [51,52]. Thus, distinct components of
the available abTCR repertoire are utilized to recognize
CD1d-Ag in the same way. Indeed, Type I NKT TCRs
act more like a pattern recognition receptor in which the
germline-encoded regions of the TCR solely interact with
the lipid antigen [1,53].
The inherent flexibility within the available abTCR
repertoire to see restricted CD1d-Ag complexes is
Current Opinion in Immunology 2013, 25:653–659

656 Immunogenetics and transplantation
Figure 3
TCR-MHC-peptide
Type I NKT TCR-CD1d-αGalCer
MAIT TCR-MR1-RL-6-Me-7-OH
CDR1α
CDR2α
CDR3α
CDR2β
CDR1β
CDR3βCDR1α
CDR2α
CDR3αCDR2β
CDR1β
CDR3β
CDR1α
CDR2α
CDR3α
CDR2β
CDR1β
CDR3β
β2mβ2mβ2m
CD1d MR1 MHC-I
Vα VβVα Vβ Vα Vβ
Cα Cβ Cα Cβ Cα Cβ
MAIT-TCR NKT TCR TCR
peptide RL-6Me-7-OH
αGalCer
(a)
(b)
Current Opinion in Immunology
TCR recognition of peptides, lipids and vitamin B Ags. LC13 TCR in complex with HLA-B8-FLR (left panels, PDB ID: 1MI5), MAIT TCR in complex with
MR1-RL-6-Me-7-OH (middle panels, PDB ID: VL4V) and NKT TCR in complex with CD1d-aGalCer (left panels, PDB ID: 3HE6). Depicted are the side-
views of the overall complexes (a) and the top-views of the Ag-MHC with isolated CDR loops and centres of mass for the Va and Vb domains (black
spheres) (b). TCR a — chains are coloured in pale green, CDRa — loops in blue; TCR b — chains are coloured in cyan, CDRb — loops in green. MHC
molecules are coloured in grey; peptide, vitamin B and lipid antigens are shown in stick format and coloured in pink, light green and yellow,
respectively.
evidenced by analysis of Type II NKT cells. Here
Type II NKT cells, which exhibit more diverse TCR
usage than Type I NKT cells, can dock in a markedly
different manner onto CD1d [54�,55�]. As observed with
some conventional abTCR-pMHC interactions, the non-
germline encoded regions of sulfatide-reactive TCRs
dominated contacts with the Ag and the Ag-presenting
molecule.
MAIT cells also exhibit invariant TCR a-chain usage
(Va7.2-Ja33 in humans) and express a limited repertoire
of TCR b-chains (Vb2,13 in humans) [56] and recognize
MR1 presenting vitamin B metabolites [5��]. Interest-
ingly, the recently described GEM T cells, which are
restricted to CD1b presenting GMM, also exhibited
biased TRAV1-2 usage, although it remains to be deter-
mined how these GEM TCRs interact with CD1b-Ag [2].
Current Opinion in Immunology 2013, 25:653–659
The structure of the human MAIT TCR complexed to
MR1 bound to a non-activating and activating ligand
provided the first definitive insight into vitamin B metab-
olite mediated recognition (Figure 3) [57�]. The MAIT
TCR docked approximately centrally and orthogonally to
the main axis of the MR1-Ag-binding cleft, consistent
with a xenoreactive MAIT TCR-MR1 complex in which
ligands were hypothetically modelled into the MR1 cleft
[58]. Alongside mutagenesis studies of the human MAIT
TCR-MR1 system [59], these findings revealed that the
Va7.2-Ja33 chain formed many contacts with MR1, while
the germline-encoded regions of the Vb chain played a
lesser role, although the CDR3b loop sat above the Ag,
and interacted extensively with MR1 [57�]. Several of the
residues from the invariant a-chain that contact MR1 are
either conserved or synonymously substituted across all
known eutherian and marsupial a-chain orthologues,
www.sciencedirect.com

Rules of ab TCR recognition Eckle et al. 657
suggesting longstanding co-evolution of this TCR-MR1
interaction, consistent with the high sequence conserva-
tion of MR1 across species. Interestingly, the docking
mode of the MAIT TCR-MR1-Ag complex resembled a
Va7.2+ TCR-pMHC-I complex [60], indicating that
MAIT TCR recognition shared more ‘adaptive features’
than the innate-like characteristics of the Type I NKT
TCR. Nevertheless, it is likely that a common mode of
MAIT TCR-MR1 docking will underpin recognition of
any other (yet to be identified) MR1-restricted Ags.
ConclusionsPresently, the field of MHC-mediated immunity is con-
founded by the various docking strategies employed by
the T cell repertoire. Ironically, whilst the field of CD1
and MR1-restricted biology is less well established than
MHC-recognition, the ‘rules of engagement’ are much
clearer in these systems in which invariant TCR usage
seems to be a predominant driving force in recognizing a
monomorphic Ag presenting molecule. It is likely not to
be coincidental that the TRAV1-2 gene of the TCR has
been used for recognizing peptides, vitamin B metab-
olites and lipids bound to their respective Ag-presenting
molecules [2,57�,60]. Here, the TCR hard wiring seems
geared to interacting with the monomorphic MHC-I like
molecule as well as antigen, independent of the chemistry
of its antigenic cargo. Perhaps the field of MHC biology
could learn much from examining TCR recognition of
MHC-I like molecules.
AcknowledgementsThis research was supported by the National Health and Medical ResearchCouncil of Australia (NHMRC). S.J.T. was supported by an AustralianResearch Council Future Fellowship, J.R. was supported by an NHMRCAustralia Fellowship.
References and recommended readingPapers of particular interest, published within the period of review,have been highlighted as:
� of special interest
�� of outstanding interest
1. Rossjohn J, Pellicci DG, Patel O, Gapin L, Godfrey DI: Recognitionof CD1d-restricted antigens by natural killer T cells. Nat RevImmunol 2012, 12:845-857.
2. Van Rhijn I, Kasmar A, de Jong A, Gras S, Bhati M,Doorenspleet ME, de Vries N, Godfrey DI, Altman JD, de Jager Wet al.: A conserved human T cell population targetsmycobacterial antigens presented by CD1b. Nat Immunol 2013,14:706-713.
3. Gapin L: Where do MAIT cells fit in the family of unconventionalT cells? PLoS Biol 2009, 7:e1000070.
4. Adams EJ, Luoma AM: The adaptable major histocompatibilitycomplex (MHC) fold: structure and function of nonclassicaland MHC class I-like molecules. Annu Rev Immunol 2013,31:529-561.
5.��
Kjer-Nielsen L, Patel O, Corbett AJ, Le Nours J, Meehan B, Liu L,Bhati M, Chen Z, Kostenko L, Reantragoon R et al.: MR1 presentsmicrobial vitamin B metabolites to MAIT cells. Nature 2012,491:717-723.
This paper describes the elucidation of the long-sought after ligand forMAIT cells.
www.sciencedirect.com
6.��
Illing PT, Vivian JP, Dudek NL, Kostenko L, Chen Z, Bharadwaj M,Miles JJ, Kjer-Nielsen L, Gras S, Williamson NA et al.: Immuneself-reactivity triggered by drug-modified HLA-peptiderepertoire. Nature 2012, 486:554-558.
This is the first report to describe how a small molecule drug canmodulate the HLA peptide repertoire.
7. Illing PT, Vivian JP, Purcell AW, Rossjohn J, McCluskey J: Humanleukocyte antigen-associated drug hypersensitivity. Curr OpinImmunol 2013, 25:81-89.
8. Gras S, Kjer-Nielsen L, Burrows SR, McCluskey J, Rossjohn J: T-cell receptor bias and immunity. Curr Opin Immunol 2008,20:119-125.
9. Turner SJ, Doherty PC, McCluskey J, Rossjohn J: Structuraldeterminants of T-cell receptor bias in immunity. Nat RevImmunol 2006, 6:883-894.
10. Broughton SE, Petersen J, Theodossis A, Scally SW, Loh KL,Thompson A, van Bergen J, Kooy-Winkelaar Y, Henderson KN,Beddoe T et al.: Biased T cell receptor usage directed againsthuman leukocyte antigen DQ8-restricted gliadin peptides isassociated with celiac disease. Immunity 2012, 37:611-621.
11.�
Yin L, Scott-Browne J, Kappler JW, Gapin L, Marrack P: T cellsand their eons-old obsession with MHC. Immunol Rev 2012,250:49-60.
A focused review on TCR hardwiring within the Vb8.2 axis.
12. Gapin L, Godfrey DI, Rossjohn J: Natural killer T cell obsessionwith self-antigens. Curr Opin Immunol 2013, 25:168-173.
13. Treiner E, Duban L, Bahram S, Radosavljevic M, Wanner V, Tilloy F,Affaticati P, Gilfillan S, Lantz O: Selection of evolutionarilyconserved mucosal-associated invariant T cells by MR1.Nature 2003, 422:164-169.
14. Zinkernagel RM, Doherty PC: The discovery of MHC restriction.Immunol Today 1997, 18:14-17.
15. Jerne NK: The somatic generation of immune recognition. Eur JImmunol 1971, 1:1-9.
16. Flajnik MF, Kasahara M: Origin and evolution of the adaptiveimmune system: genetic events and selective pressures. NatRev Genet 2010, 11:47-59.
17. Tikhonova AN, Van Laethem F, Hanada K-i, Lu J, Pobezinsky LA,Hong C, Guinter TI, Jeurling SK, Bernhardt GN, Park JH et al.: ab Tcell receptors that do not undergo major histocompatibilitycomplex-specific thymic selection possess antibody-likerecognition specificities. Immunity 2012, 36:79-91.
18.�
Van Laethem F, Tikhonova AN, Singer A: MHC restriction isimposed on a diverse T cell receptor repertoire by CD4 andCD8 co-receptors during thymic selection. Trends Immunol2012, 33:437-441.
A thoughoutful and opposing view on TCR hardwiring and MHC restric-tion.
19. Van Laethem F, Sarafova SD, Park JH, Tai X, Pobezinsky L,Guinter TI, Adoro S, Adams A, Sharrow SO, Feigenbaum L et al.:Deletion of CD4 and CD8 coreceptors permits generation ofalphabetaT cells that recognize antigens independently of theMHC. Immunity 2007, 27:735-750.
20. Ely LK, Burrows SR, Purcell AW, Rossjohn J, McCluskey J: T-cellsbehaving badly: structural insights into alloreactivity andautoimmunity. Curr Opin Immunol 2008, 20:575-580.
21.�
Gras S, Burrows SR, Turner SJ, Sewell AK, McCluskey J,Rossjohn J: A structural voyage toward an understanding ofthe MHC-I-restricted immune response: lessons learned andmuch to be learned. Immunol Rev 2012, 250:61-81.
A comprehensive review on TCR-pMHC-I interactions.
22. Gras S, Kjer-Nielsen L, Chen Z, Rossjohn J, McCluskey J: Thestructural bases of direct T-cell allorecognition: implicationsfor T-cell-mediated transplant rejection. Immunol Cell Biol2011, 89:388-395.
23.�
Garcia KC, Adams JJ, Feng D, Ely LK: The molecular basis ofTCR germline bias for MHC is surprisingly simple. Nat Immunol2009, 10:143-147.
A clear explanation of the ‘interaction codon’ concept.
Current Opinion in Immunology 2013, 25:653–659

658 Immunogenetics and transplantation
24. Godfrey DI, Rossjohn J, McCluskey J: The fidelity, occasionalpromiscuity, and versatility of T cell receptor recognition.Immunity 2008, 28:304-314.
25. Feng D, Bond CJ, Ely LK, Maynard J, Garcia KC: Structuralevidence for a germline-encoded T cell receptor-majorhistocompatibility complex interaction ‘codon’. Nat Immunol2007, 8:975-983.
26. Tynan FE, Burrows SR, Buckle AM, Clements CS, Borg NA,Miles JJ, Beddoe T, Whisstock JC, Wilce MC, Silins SL et al.: T cellreceptor recognition of a ‘super-bulged’ majorhistocompatibility complex class I-bound peptide. NatImmunol 2005, 6:1114-1122.
27. Burrows SR, Chen Z, Archbold JK, Tynan FE, Beddoe T, Kjer-Nielsen L, Miles JJ, Khanna R, Moss DJ, Liu YC et al.: Hard wiringof T cell receptor specificity for the major histocompatibilitycomplex is underpinned by TCR adaptability. Proc Natl AcadSci U S A 2010, 107:10608-10613.
28. Garcia KC: Reconciling views on T cell receptor germline biasfor MHC. Trends Immunol 2012, 33:429-436.
29. Scott-Browne JP, Crawford F, Young MH, Kappler JW, Marrack P,Gapin L: Evolutionarily conserved features contribute to ab Tcell receptor specificity. Immunity 2011, 35:526-535.
30. Scott-Browne JP, White J, Kappler JW, Gapin L, Marrack P:Germline-encoded amino acids in the alphabeta T-cellreceptor control thymic selection. Nature 2009, 458:1043-1046.
31.��
Colf LA, Bankovich AJ, Hanick NA, Bowerman NA, Jones LL,Kranz DM, Garcia KC: How a single T cell receptor recognizesboth self and foreign MHC. Cell 2007, 129:135-146.
This paper provides first insight into mouse and human TCR alloreactivity,respectively.
32. Adams JJ, Narayanan S, Liu B, Birnbaum ME, Kruse AC,Bowerman NA, Chen W, Levin AM, Connolly JM, Zhu C et al.: T cellreceptor signaling is limited by docking geometry to peptide-major histocompatibility complex. Immunity 2011, 35:681-693.
33.�
Stadinski BD, Trenh P, Smith RL, Bautista B, Huseby PG, Li G,Stern LJ, Huseby ES: A role for differential variable gene pairingin creating T cell receptors specific for unique majorhistocompatibility ligands. Immunity 2011, 35:694-704.
This finding shows that the TCR b-chain ‘interaction codon’ can bemodulated by the TCR a-chain.
34. Turner S, Rossjohn J: abT cell receptors come out swinging.Immunity 2011, 35:660-662.
35. Miles JJ, Elhassen D, Borg NA, Silins SL, Tynan FE, Burrows JM,Purcell AW, Kjer-Nielsen L, Rossjohn J, Burrows SR et al.: CTLrecognition of a bulged viral peptide involves biased TCRselection. J Immunol 2005, 175:3826-3834.
36. Archbold JK, Ely LK, Kjer-Nielsen L, Burrows SR, Rossjohn J,McCluskey J, Macdonald WA: T cell allorecognition and MHCrestriction — a case of Jekyll and Hyde? Mol Immunol 2008,45:583-598.
37. Borg NA, Ely LK, Beddoe T, Macdonald WA, Reid HH,Clements CS, Purcell AW, Kjer-Nielsen L, Miles JJ, Burrows SRet al.: The CDR3 regions of an immunodominant T cell receptordictate the ‘energetic landscape’ of peptide-MHC recognition.Nat Immunol 2005, 6:171-180.
38. Gras S, Wilmann PG, Chen Z, Halim H, Liu YC, Kjer-Nielsen L,Purcell AW, Burrows SR, McCluskey J, Rossjohn J: A structuralbasis for varied abTCR usage against an immunodominantEBV antigen restricted to a HLA-B8 molecule. J Immunol 2012,188:311-321.
39.��
Macdonald WA, Chen Z, Gras S, Archbold JK, Tynan FE,Clements CS, Bharadwaj M, Kjer-Nielsen L, Saunders PM,Wilce MCJ et al.: T cell allorecognition via molecular mimicry.Immunity 2009, 31:897-908.
See annotation to Ref. [39��].
40. Gras S, Chen Z, Miles JJ, Liu YC, Bell MJ, Sullivan LC, Kjer-Nielsen L, Brennan RM, Burrows JM, Neller MA et al.: Allelicpolymorphism in the T cell receptor and its impact on immuneresponses. J Exp Med 2010, 207:1555-1567.
Current Opinion in Immunology 2013, 25:653–659
41. Gras S, Burrows SR, Kjer-Nielsen L, Clements CS, Liu YC,Sullivan LC, Bell MJ, Brooks AG, Purcell AW, McCluskey J et al.:The shaping of T cell receptor recognition by self-tolerance.Immunity 2009, 30:193-203.
42. Liu YC, Chen Z, Burrows SR, Purcell AW, McCluskey J,Rossjohn J, Gras S: The energetic basis underpinning T-cellreceptor recognition of a super-bulged peptide bound to amajor histocompatibility complex class I molecule. J BiolChem 2012, 287:12267-12276.
43. Liu YC, Miles JJ, Neller MA, Gostick E, Price DA, Purcell AW,McCluskey J, Burrows SR, Rossjohn J, Gras S: Highly divergentT-cell receptor binding modes underlie specific recognition ofa bulged viral peptide bound to a human leukocyte antigenclass I molecule. J Biol Chem 2013, 288:15442-15454.
44. Godfrey DI, MacDonald HR, Kronenberg M, Smyth MJ, Van Kaer L:NKT cells: what’s in a name? Nat Rev Immunol 2004, 4:231-237.
45. Borg NA, Wun KS, Kjer-Nielsen L, Wilce MC, Pellicci DG, Koh R,Besra GS, Bharadwaj M, Godfrey DI, McCluskey J et al.: CD1d-lipid-antigen recognition by the semi-invariant NKT T-cellreceptor. Nature 2007, 448:44-49.
46. Uldrich AP, Patel O, Cameron G, Pellicci DG, Day EB, Sullivan LC,Kyparissoudis K, Kjer-Nielsen L, Vivian JP, Cao B et al.: A semi-invariant V[alpha]10+ T cell antigen receptor defines apopulation of natural killer T cells with distinct glycolipidantigen-recognition properties. Nat Immunol 2011, 12:616-623.
47. Pellicci DG, Patel O, Kjer-Nielsen L, Pang SS, Sullivan LC,Kyparissoudis K, Brooks AG, Reid HH, Gras S, Lucet IS et al.:Differential recognition of CD1d-alpha-galactosyl ceramide bythe V beta 8.2 and V beta 7 semi-invariant NKT T cell receptors.Immunity 2009, 31:47-59.
48. Patel O, Pellicci DG, Uldrich AP, Sullivan LC, Bhati M, McKnight M,Richardson SK, Howell AR, Mallevaey T, Zhang J et al.: Vb2natural killer T cell antigen receptor-mediated recognition ofCD1d-glycolipid antigen. Proc Natl Acad Sci U S A 2011,108:19007-19012.
49. Lopez-Sagaseta J, Kung JE, Savage PB, Gumperz J, Adams EJ:The molecular basis for recognition of CD1d/a-galactosylceramide by a human non-Va24 T cell receptor.PLoS Biol 2012, 10:e1001412.
50. Godfrey DI, Rossjohn J: New ways to turn on NKT cells. J ExpMed 2011, 208:1121-1125.
51. Godfrey DI, Pellicci DG, Patel O, Kjer-Nielsen L, McCluskey J,Rossjohn J: Antigen recognition by CD1d-restricted NKT T cellreceptors. Semin Immunol 2010, 22:61-67.
52. Girardi E, Zajonc DM: Molecular basis of lipid antigenpresentation by CD1d and recognition by natural killer T cells.Immunol Rev 2012, 250:167-179.
53. Scott-Browne JP, Matsuda JL, Mallevaey T, White J, Borg NA,McCluskey J, Rossjohn J, Kappler J, Marrack P, Gapin L:Germline-encoded recognition of diverse glycolipids bynatural killer T cells. Nat Immunol 2007, 8:1105-1113.
54.�
Patel O, Pellicci DG, Gras S, Sandoval-Romero ML, Uldrich AP,Mallevaey T, Clarke AJ, Le Nours J, Theodossis A, Cardell SL et al.:Recognition of CD1d-sulfatide mediated by a type II naturalkiller T cell antigen receptor. Nat Immunol 2012, 13:857-863.
See annotation to Ref. [55�].
55.�
Girardi E, Maricic I, Wang J, Mac T-T, Iyer P, Kumar V, Zajonc DM:Type II natural killer T cells use features of both innate-like andconventional T cells to recognize sulfatide self antigens. NatImmunol 2012, 13:851-856.
This paper provides first insight into Type II NKT TCR recognition.
56. Gold MC, Lewinsohn DM: Co-dependents: MR1-restrictedMAIT cells and their antimicrobial function. Nat Rev Microbiol2013, 11:14-19.
57.�
Patel O, Kjer-Nielsen L, Le Nours J, Eckle SBG, Birkinshaw R,Beddoe T, Corbett AJ, Liu L, Miles JJ, Meehan B et al.:Recognition of vitamin B metabolites by mucosal-associatedinvariant T cells. Nat Commun 2013:4.
This paper provides first formal insight into how the MAIT TCR binds tovitamin B metabolites bound to MR1.
www.sciencedirect.com

Rules of ab TCR recognition Eckle et al. 659
58. Lopez-Sagaseta J, Dulberger CL, Crooks JE, Parks CD, Luoma AM,McFedries A, Van Rhijn I, Saghatelian A, Adams EJ: The molecularbasis for mucosal-associated invariant T cell recognition ofMR1 proteins. Proc Natl Acad Sci U S A 2013, 110:E1771-E1778.
59. Reantragoon R, Kjer-Nielsen L, Patel O, Chen Z, Illing PT, Bhati M,Kostenko L, Bharadwaj M, Meehan B, Hansen TH et al.:Structural insight into MR1-mediated recognition of the
www.sciencedirect.com
mucosal associated invariant T cell receptor. J Exp Med 2012,209:761-774.
60. Tynan FE, Reid HH, Kjer-Nielsen L, Miles JJ, Wilce MC,Kostenko L, Borg NA, Williamson NA, Beddoe T, Purcell AW et al.:A T cell receptor flattens a bulged antigenic peptide presentedby a major histocompatibility complex class I molecule. NatImmunol 2007, 8:268-276.
Current Opinion in Immunology 2013, 25:653–659