Analysis, gene cloning and expression of two α-amylases from Bacillus cereus
Transcript of Analysis, gene cloning and expression of two α-amylases from Bacillus cereus

Chinese Journal of Agricultural Biotechnologyhttp://journals.cambridge.org/CJA
Additional services for Chinese Journal of Agricultural Biotechnology:
Email alerts: Click hereSubscriptions: Click hereCommercial reprints: Click hereTerms of use : Click here
Analysis, gene cloning and expression of two αamylases from Bacillus cereus
Zhang Linlin, Wang Qi, Li Rongxi, Wang Yongjun and Mei Ruhong
Chinese Journal of Agricultural Biotechnology / Volume 6 / Issue 02 / August 2009, pp 135 140DOI: 10.1017/S1479236209002538, Published online: 30 October 2009
Link to this article: http://journals.cambridge.org/abstract_S1479236209002538
How to cite this article:Zhang Linlin, Wang Qi, Li Rongxi, Wang Yongjun and Mei Ruhong (2009). Analysis, gene cloning and expression of two αamylases from Bacillus cereus. Chinese Journal of Agricultural Biotechnology,6, pp 135140 doi:10.1017/S1479236209002538
Request Permissions : Click here
Downloaded from http://journals.cambridge.org/CJA, IP address: 128.148.252.35 on 08 Sep 2012

Analysis, gene cloning and expression of twoa-amylases from Bacillus cereus
Zhang Lin-lin1,2, Wang Qi1,*, Li Rong-xi2, Wang Yong-jun1 and Mei Ru-hong2
1Department of Plant Pathology, China Agricultural University, Beijing 100094, China; 2Department
of Agronomy, Inner Mongolia Agricultural University, Huhehaote 010018, China
Received 25 January 2008; Accepted 7 May 2008
First published in Journal of Agricultural Biotechnology 2008, 16(4): 676–680
AbstractBacillus cereus can produce a-amylase, which has important industrial production value and
can endure high temperatures. The enzymatic characteristics of two a-amylases from B. cereus
(B905 and B904) were studied. The results showed that both preserved their activity at
90–100 �C. Their thermal stability and enzymatic activity did not depend on Ca2+. However,
their protein molecular weights were obviously different. The structural genes of amy905 and
amy904 were cloned successfully using the polymerase chain reaction (PCR) and expressed in
Escherichia coli. The amy904 and amy905 genes were 1362 bp and 1761 bp long and their
protein molecular weights were about 55 kDa and 68 kDa, respectively.
Keywords: Bacillus cereus; a-amylase; gene clone; gene expression
Introduction
Amylase, the enzyme that hydrolyses starch, is found
widely in animals, plants and microbes (Li and Cai, 2004).
Nowadays, amylase is the most important, industrially
produced and, so far, the most frequently used enzyme in
starch processing (Kong and Wang, 1989; MacGregor and
Janecek, 2001). Starch liquefaction is usually conducted
under high temperatures (Du and Hao, 2006), therefore
high-temperature resistance and thermal stability are the
main features of interest regarding the use of amylase
(Guo and Li, 2006). The a-amylase from B. cereus
presents these characteristics as well as a high activity
(Kang and Lee, 2004).
Bacillus cereus B905 and B904 are beneficial endo-
phytic bacteria isolated from plants. The results from
greenhouse and field experiments have shown that both
strains promote plant growth and have a prophylactic
effect on plants. Before colonizing the plant host, these
two strains use available nutrients to intensify their meta-
bolism and increase their population (Kang and Lee,
2004). Extracellular amylase activities have been found
on both strains.
In this experiment, the characteristics of extracellular
amylase of B. cereus B905 and B904 strains were analysed
and the amylase genes were cloned. Results from this
study could help in revealing the molecular mechanism
of plant colonization by Bacillus cereus, and provide new
genetic resources for the industrial production of highly
thermoresistant amylase.
Materials and methods
Bacterial strains and plasmids
The bacterial strains and plasmids used in this experiment
are shown in Table 1. Strains were cultured on plates of
liquid or agarose Luria–Bertani (LB) media. The selective
medium for a-amylase was LBS (LB medium with 1%
soluble starch), supplemented, when necessary, with
50mg/ml ampicillin (Amp) to ensure plasmid stability in
the bacteria.* Corresponding author. E-mail: [email protected]
g China Agricultural University 2009 Chinese Journal of Agricultural Biotechnology 6(2); 135–140ISSN 1479-2362 doi:10.1017/S1479236209002538

Enzymes and biochemical reagents
Tris-saturated phenol and Amp were produced by Beijing
Xinjingke Biotechnology (Beijing, China). Restriction
endonucleases XbaI, HindIII and BamHI were from
Promega (Wisconsin, USA). Agarose was from Spain,
packed by TanWei Biotechnology (Beijing, China). dNTP
came from TaKaRa (Dalian, China) and polymerase Taq
DNA and ligase T4 DNA were purchased from TanWei
Biotechnology. Other analytically pure reagents were all
of domestic origin.
Enzymological studies of a-amylase
Plaque detection and determination of enzymeactivity of amylaseB. cereus 905 and 904 were inoculated in solid medium
LBS (Zhou and Ma, 2006). After cultivation at 32 �C for
72 h, 3 mmol/l iodine and 10 mmol/l KI were added for
staining (Zhang and Zhang, 2004). Enzyme activity was
determined following QB/T1803 (Niu and Xu, 2006).
(Enzyme activity: when 1 g enzyme powder or 1 ml
enzyme liquid dissolves 1 mg soluble starch at 60 �C in
1 min, it is one enzyme activity unit expressed in U/g or
U/ml.)
Electrophoretic analysis of amylasesAfter 48 h the culture was centrifuged and the supernatant
was taken as a crude extract of amylase. An aliquot of 5ml
SDS-PAGE upper sample buffer was added to 20ml crude
extract. After electrophoresis by native-PAGE, the gel,
containing 0.5% soluble starch, was treated overnight with
Tris–HCl (pH 6.8) for protein renaturation, and also for
iodine staining activity.
Detection of heat resistance of amylaseEnzyme solutions were treated at different temperatures
(40–100 �C) for 10, 30, 60, 120 and 130 min, respectively.
Then, they were rapidly cooled on ice to room
temperature. The relative activity of the residual enzymes
was measured by comparison with the enzyme activity of
the unheated enzyme solution (100%).
Detection of the acid and alkali resistance of amylaseEnzyme solutions were diluted in buffer solutions at dif-
ferent pH (Cl–HCl pH 2.0–2.2, Gly–HCl pH 3.0, HAc-NaAc
pH 4.0–5.0, Tris–HCl pH 6.0–8.0, Gly–NaOH pH 9.0–10.0,
Na2HPO4–NaOH pH 11.0, KCl–NaOH pH 12.0–13.0)
(Deutch, 2002). After incubating at 40 �C for 1 h, the resi-
dual enzyme activity was measured (the highest enzyme
activity was considered as 100%).
Detection of the impact of metal ions and proteaseinhibitor on enzyme activityDifferent metal ions and the protease inhibitor sodium
dodecyl sulphate (SDS) and ethylenediaminetetraacetic
acid (EDTA) were added into the reaction systems at a
final concentration of 1.0 mmol/l. The enzyme solution
without metal ions was used as a control.
Cloning and expression of genes encoding a-amylase
Total DNA extraction from B. cereusOne Bacillus cereus strain was selected and put in 5 ml LB
liquid at 37 �C, before shaking at 200 rpm until OD600 =0.8. The bacteria were collected from 1 ml culture by
centrifugation and were extracted in 600ml Tris–EDTA
(TE) buffer pH 8.0, containing 2 mg/ml lysozyme, 10% SDS
and 20 mg/ml protease K. After 1 h in a water bath at 37 �C,
solutions of 5 mol/l NaCl and cationic surfactant hexa-
decyltrimethylammonium bromide (CTAB)/NaCl (10%/
0.7 mol/l) were added and well mixed. After incubation in
a water bath at 65 �C for 10 min, the same volume of
phenol/chloroform (1 : 1,v/v) was added and the super-
natant was precipitated with 0.6 volumes of isopropanol.
The precipitate was washed in 70% ethanol, air-dried and
dissolved in 20ml TE buffer (pH 8.0) and preserved
according to Xia (2002).
Table 1. Bacterial strains and plasmids used
Bacterial strains orplasmids Relevant characteristics Source
Escherichia coliDH5a
f80 d lacZDM15 recA1 endA1 gyrA96 thi-1 hsdR17 supE44 relA1 deoRD(lacZYA-argF )U169 F -
This lab
E. coli BL21 hsdS gel(lcIts857 ind1 Sam7 nin5 lac UV-5-T7) This labBacillus cereus 905 Wild type This labB. cereus 904 Wild type This labpET-22b (+) Expression vector; T7 promoter; KanR This labpET-amy904 pET-22b(+) harbouring the entire open reading frame of amy904 from
B. cereus B904Thiswork
pET-amy905 pET-22b(+) harbouring the entire open reading frame of amy905 fromB. cereus B905
This work
136 Zhang Lin-lin et al.

Cloning of amylase genesBased on B. cereus a-amylase DNA sequences in
GenBank, the following primers were designed:
AmyS-5: (50-ACAGGATCCATGCGTGTGGGGAAAATAC-30);
AmyS-3: (50-ACACTCGAGTTTCCGTCTCTTTTTAACC-30).
AmyL-5: (50-ACAGGATCCGATGCTTAAAGAAGC-30);
AmyL-3: (50-ACACTCGAGCCATTTTAATATGGAG-30).
The underlined regions indicate restriction sites, for
BamHI in AmyS-5 and AmyL-5 and for XhoI in AmyS-3
and AmyL-3.
PCR amplifications with genome DNA encoding
B. cereus 905 and B. cereus 904 were carried out, using
the following parameters: denaturation at 94 �C for 5 min;
35 cycles of denaturation at 94 �C for 30 s, annealing at
50 �C for 30 s, and elongation at 72 �C for 40 s; extension
at 72 �C for 10 min. The amplified fragment was recovered
by electrophoresis after double digestion. The recovery
product was inserted into pET-22b(+) after a double
digestion with BamHI and XhoI. Escherichia coli BL21
(DE3) was transformed with the resulting plasmid for
protein expression and sequence analysis (Wang and
Peng, 2007).
Expression of amylase genes in E. coliRecombinant E. coli was cultured overnight at 37 �C in
5 ml LB containing 100mg/ml AMP. It was then inoculated
in 50 ml LB [1% (v/v)] and cultured at 37 �C until OD600 =0.6–0.8. The inducer isopropyl b-D-thiogalactopyranoside
(IPTG) was added at a final concentration of 1 mmol/l.
The culture was incubated at 37 �C for 5 h to induce the
expression of target genes.
Induced bacteria (1 ml) were centrifuged at 12 000 rpm
for 30 s. The pellet was washed with 50 mmol/l
phosphate buffer solution (pH 6.0) and received the
same volume of SDS upper sample buffer (40 mmol/l
Tris–HCl, 10% glycerol, 2% SDS, 5% mercaptoethanol and
0.1% bromophenol blue). The tube was bathed in boiling
water for 5 min, rapidly cooled on ice for 2 min and cen-
trifuged at 5000 rpm for 10 min. A 12% SDS-PAGE of 5ml
supernatant was carried out (uninduced bacteria were
used as a control).
Results
Amylase test of the two strains
After B905 and B904 were cultured in LBS medium for
48 h, they were stained with iodine. The results showed a
bright circle of starch degradation around the growing
colonies (Fig. 1).
The native PAGE, protein renaturation staining and
iodine activity showed differences in the form of the active
protein between these two bacteria (Fig. 2), suggesting
that they (AMY905 and AMY904) are different amylases.
Effect of pH on enzyme activity and stability
The crude amylase enzymes from the two B. cereus
strains were extracted and tested for acid and alkali
resistance at 40 �C. The results showed that their levels of
sensitivity to acidity and alkalinity were similar. The
optimum pH ranged between 6.0 and 10.0. In this range,
both amylases retained more than 50% enzyme activity
in 48 h (Fig. 3), which meant that the two enzymes could
have relatively high activity over a wide pH range.
1 2
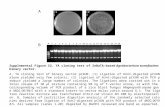
Fig. 1. Activity of Bacillus cereus on the 1% starch of LBS substrate. 1, B. cereus 905; 2, B. cereus 904. (See online for acolour version of this figure.)
Analysis, gene cloning and expression of a-amylases from B. cereus 137

Impact of temperature on enzyme activityand stability
The heat resistance test revealed that the a-amylase
generated from both strains maintained high activity in
the temperature range of 40–100 �C (Fig. 4). Their
activities were more than 70 U/ml for both strains,
showing that they were both high-temperature resistant
a-amylases. Below 100 �C, the enzyme activity of strain
B904 was lower than that of strain B905. Their a-amylases
exhibited high heat stability without Ca2+ dependence
(see below). After treatment at 100 �C for 2 h, the enzyme
activity of the amylases were 80% and 60% for B904 and
B905, respectively.
Impact of metal ions and protease inhibitor onenzyme activity
At room temperature and pH 6.0, the crude enzyme was
treated with 1.0 mmol/l different metal ions and protease
inhibitor for 30 min. The enzyme activity was measured
and compared. The results showed that Ca2+, Na+, Li+,
NH4+, Fe2+ and Mg2+ had a slight promoting effect on
the amylase secreted from B905. Whereas Ca2+, Na+, Li+,
NH4+ and Fe2+ had almost no effect on the amylase
from B904, some inhibitory effect was noticed with Mg2+.
Cu2+ strongly inhibited amylase activity for the two
strains (Fig. 5). The protease inhibitors SDS and EDTA
both had a promoting effect on the amylase from the two
strains.
Cloning of genes encoding a-amylase
The gene sequences of a-amylase from B. cereus strains
in GenBank were compared. There were two completely
different a-amylase genes of B. cereus. According to gene
sequences, two primers AmyL-5/AmyL-3 and AmyS-5/
AmyS-3 were designed. The results of PCR amplification
showed that only a 1700 bp band was amplified by AmyL-
5/AmyL-3 from B905 and a 1000 band by AmyS-5/AmyS-3
from B904 (Fig. 6). The sequence analysis revealed that
B905 and B904 contained 1761 and 1362 bp DNA frag-
ments, respectively. From BLAST sequence analysis we
know that both fragments belonged to a-amylase genes,
namely amy905 and amy904, respectively.
Induced expression of a-amylase genes inEscherichia coli
We inserted amy905 and amy904 into plasmids, produ-
cing pET-amy905 and pET-amy904. E. coli BL21 was
transformed with these. After inducing expression with
ITPG, fragmented cells contained a-amylase activity. The
SDS-PAGE results showed protein bands of 55 kDa
(amy904) and 68 kDa (amy905) (Fig. 7).
1 2
Fig. 2. Identification of B. cereus amylases by nativePAGE. 1, B. cereus 905; 2, B. cereus 904.
%
Fig. 3. Effect of pH value on activity of enzymes producedby B. cereus.
Enz
yme
activ
ity (
U/m
l)
Temperature (°C)
Fig. 4. Effect of temperature on enzyme activity producedby B. cereus.
138 Zhang Lin-lin et al.

Discussion
The enzymological analysis showed that a-amylases from
B. cereus 905 and 904 (B905 and B904) conserved their
activity in a relatively wide pH range. The stability tem-
perature of the enzyme activity reached 90–100 �C, and
the thermal stability as well as the enzymatic activity did
not depend on Ca2+. Therefore, these new proteins,
which are high-temperature resistant and extracellular
Ca2+ independent, could solve ongoing problems re-
lated to amylase production. This finding will help to
improve the heat resistance of the enzyme and reduce the
200
150
100
50
0
Rel
ativ
e ac
tivi
ty
CaCl2 Na2SO4 MgSO4 FeSO4 NH4HPO4 CuSO4 EDTA SDSLiCl
905904
Fig. 5. Effects of various salts, SDS and EDTA on enzyme activity produced by B. cereus.
M 1 2bp
514842693530
2027190415871375
948831
M 3 4bp
514842693530
2027190415871375
948831
Fig. 6. PCR analysis of the a-amylase from B. cereus. Lanes: M, marker; 1 and 4, B. cereus 905 PCR product; 2 and 3,B. cereus 904 PCR product.
1 2 3 4 5 kDa
Fig. 7. SDS-PAGE of the purified Amy905/Amy904 from E. coli BL21. Lanes: 1, total protein of E. coli BL21 (DE3) withpET-amy905 induced by IPTG; 2 and 3, total protein of E. coli BL21 (DE3) with pET-amy905/904 before inductionwith IPTG; 4, total protein of E. coli BL21 (DE3) with pET-amy904 induced by IPTG; 5, standard protein marker.
Analysis, gene cloning and expression of a-amylases from B. cereus 139

dependence of its thermal stability on Ca2+. Application
of these gene resources in industrial processes involving
amylase could, to some extent, improve the efficiency
and quality of the production of high-temperature
resistant amylase and reduce production costs.
The amylase genes have been cloned in many bacteria.
Two completely different amylases exist in B. cereus. This
experiment has validated this phenomenon at the DNA
level. The differences of gene length and sequence
showed that their proteins have evolved in two entirely
different ways.
Bacterial strains B905 and B904 were both separated
from plant hosts. The amylase produced extracellularly
could very well help bacteria degrade sugar and ease its
absorption, so that bacterial nutrition could be ensured
when the bacteria colonize plant hosts. The two different
amylase genes provide potential for bacterial genetic
modification and improvement of colonization ability in
plant hosts.
Acknowledgements
This work was supported by National High Technology
Research and Development Plan ‘863 plan’ (No.
2006AA10A211) and Special Agricultural Commonwealth
Research Programs: Physaclospora piricola resistance
resources information database establishment (No.
nyhyzx07055).
References
Deutch CE (2002) Characterization of a salt-tolerant extracellulara-amylase from Bacillus dipsosauri. Letters in AppliedMicrobiology 35: 78–84.
Du BB and Hao S (2006) Expression of a thermostablea-amylase mutant into Escherichia coli and Pichia pastoris.Acta Microbiologica Sinica 46(5): 827–830 (in Chinese withEnglish abstract).
Guo JQ and Li YM (2006) Molecular cloning and ex-pression of extremely thermostable and acid-stable amylasegene in Pichia pastoris. Chinese Journal of Biotechnology22(2): 237–242 (in Chinese with English abstract).
Kang HK and Lee JH (2004) Cloning and expression ofLipomyces starkeyi alpha-amylase in Escherichia coli anddetermination of some of its properties. Microbiology Letters233(1): 53–56.
Kong XL and Wang JY (1989) Research and application ofa-amylase. Microbiology 16(5): 282–287 (in Chinese withEnglish abstract).
Li JX and Cai H (2004) Cloning expression of heat-stablea-amylase gene from Bacillus licheniformis in E. coli. Foodand Fermentation Industries 1(30): 70–73 (in Chinese withEnglish abstract).
MacGregor EA and Janecek S (2001) Relationship of sequenceand structure to specificity in the a-amylase family ofenzymes. Biochimica et Biophysica Acta 1548: 1–20.
Niu DD and Xu M (2006) Cloning of the gene encoding athermostable a-amylase from Bacillus licheniformis CICIMB0204 and functional identification of its promoter. ActaMicrobiologica Sinica 46(4): 576–580 (in Chinese withEnglish abstract).
Wang LP and Peng DK (2007) Construction of a highly effectiveexpression system of Bacillus subtilis recombined xylanasegene. Journal of Agricultural Biotechnology 15(1): 133–137(in Chinese with English abstract).
Xia QCH (2002) The progress and development of proteomicresearch. Bulletin of the Chinese Academy of Sciences 17(3):166–169 (in Chinese).
Zhang YW and Zhang ZY (2004) Cloning and expression inE. coli of cDNA encoding sweet potato b-amylase. HighTechnology Letters 14(9): 29–33 (in Chinese with Englishabstract).
Zhou JC and Ma XD (2006) A new method to identify a-amylaseactivity and its producing bacteria. Journal of HuazhongAgricultural University 6: 58–61 (in Chinese with Englishabstract).
140 Zhang Lin-lin et al.