A new selective phage cloning vector, λ2001, with sites for XbaI, BamHI, HindIII, EcoRI, SstI and...
Transcript of A new selective phage cloning vector, λ2001, with sites for XbaI, BamHI, HindIII, EcoRI, SstI and...

217 Gene, 32 (1984) 217-224
Elsevier
GENE 1158
A new selective phage cloning vector, L2001, with sites for XbaI, BarnHI, HindIII, EcoRI, SstI and XhoI
(Recombinant DNA; E. coli I host; synthetic oligonucleotide polylinker; Spi selection; bacteriophage M13mp12)
Jonathan Karn, Hans W.D. Matthes*, Michael J. Gait and Sydney Brenner**
MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH (U.K.) Tel. 0223-248011
(Received March 3rd, 1984)
(Accepted April 17th, 1984)
SUMMARY
An improved bacteriophage 1 cloning vector, 12001, has been constructed. The phage includes a 34-bp polylinker oligonucleotide which introduces cleavage sites for XbaI, SstI, XhoI, EcoRI, Hind111 and BumHI, and can accommodate lo-kb to 23-kb fragments. Inserts that destroy the BumHI or XhoI cloning sites may be recovered by excision at flanking sites in the polylinker sequence. Insertion of foreign DNA into A2001 generates phage with a Spi- phenotype. The recombinant phage are able to grow on P2 lysogens but the parental vector phages are not. In the course of this work, the polylinker sequence was also introduced into M13mp8. This produced a new vector, M13mp12, with cloning sites for EcoRI, SmaI, XbaI, SstI, XhoI, BarnHI, and HindIII.
INTRODUCTION
Bacteriophage 1 cloning vectors are the vehicles of choice for cloning DNA fragments 5 kb to 23 kb in
* Present address: CNRS, Laboratoire de Gdnetique Mole-
culaire des Eucaryotes, Fact&C de Mtdecine, 11 Rue Humann,
67085 Strasbourg Cedex (France).
** To whom all correspondence should be directed.
Abbreviations: am, amber mutations; att, phage Iz attachment
site; BCIG, 5-bromo-4-chloro-indolyl-p-D-galactopyranoside;
bp, base pairs; A, deletion; h, host range; HPLC, high-perform-
ance liquid chromatography; .& insertion; imm, bacteriophage
immunity; IPTG, isopropyl-@-thiogalactoside; kb, kilobase
pairs; s, restriction site; Spi- , PZ-interference-less (1 mutants
that are red-, gum- and are able to grow on E. coli strains
lysogenic for coli phage P2); su’, suppressorless; su + , suppres-
sor-generating mutations; vir, virulence; ( ) , I att sites generated
by lifting crosses; ‘, as in IucZ’, indicates that the gene is
truncated at its 3’ end.
0378-l 119/84/$03.00 0 1984 Elsevier Science Publishers
length (Williams and Blattner, 1980; Brammar, 1982; Murray, 1983; Karn, 1983). Recombinants of I may be efficiently recovered by in vitro packaging (Hohn and Murray, 1977; Becker and Gold, 1975; Stemberg et al., 1977), are stable during multiple cycles of growth, easily stored and screened (Benton and Davis, 1977), and amenable to genetic manip- ulation. We developed a phage ,J BamHI vector, L1059, with a positive genetic selection for large DNA inserts (Kam et al., 1980; 1983), which sim- plitied the construction of genome libraries. In this communication, we describe the construction of an improved derivative of A 1059, 12001. A related series of phages have been constructed by Frischauf et al. (1983).

Syn
thet
ic
olig
on
ucl
eoti
des
5’ G
GG
TT
CT
AG
AG
CT
CG
AG
GA
TC
CA
AG
CT
3’
3’C
TC
CT
AG
GT
TC
GA
AC
TT
AA
GA
TC
TG
GG
5
mp
8 /
41-m
er
seq
uen
ce
(lac
- ph
age)
4 1 -
mer
d
up
lex
AM
V p
oly
mer
ase
5’G
GG
TT
CT
AG
AG
CT
CG
AG
GA
TC
CA
AG
CT
TG
AA
TT
CT
AG
AC
CC
3’
3’C
CC
AA
GA
TC
TC
GA
GC
TC
CT
AG
GT
TC
GA
AC
TT
AA
GA
TC
TG
GG
5’
I Inse
rt i
nto
Sm
a I s
ite
of
mp
8
CA
G
GA
A
AC
A
GC
T
AT
G
AC
C
AT
G
AT
T
AC
G
AA
TT
CC
C
GG
G
TT
C
TA
GA
G
CT
C
GA
G
GA
T
CC
A
AG
C
TT
G
AA
T
TC
T
AG
A
CC
C
GG
G
AT
C
CG
T
CG
AC
CT
GC
A
GC
C
AA
G
CT
T
GG
C
AC
T
GG
C
CG
T
CG
T
TT
T
AC
--
1--
Eco
R
I sm
a I X
ba
I X
ho&
+y
Hin
d E
CO
xXba
, -S
t -S
ri-
.-S
E-
pst-
H
ind
II!
Prim
er
I
Bar
n H
I
%&
C
ut
wit
h H
ind
lll, l
igat
e w
ith
T4
DN
A l
igas
e
mp
12
seq
uen
ce
(lac
+ p
hag
e)
ME
T
TH
R
ME
T
ILE
T
HR
A
SN
S
Ef?
A
RG
V
AL
LEU
G
LU
LEU
G
LU
AS
P
PR
O
SE
R
LEU
A
LA
LE”
ALA
“A
L V
AL
LEU
. .
GA
G
GA
A
AC
A
GC
T
AT
G
AC
C
AT
G
AT
T
AC
G
AA
T
TC
C
CG
G
GT
T
CT
A
GA
G
CT
C
GA
G
GA
T
CG
A
AG
C
TT
G
GC
A
CT
G
GC
C
GT
C
GT
T
TT
A
C...
L1
E
co
RI
I X
ba
I .M
X
ho
I -
Hin
d III
S
ma
I B
arn
HI
Pr,
mer
ssi
I
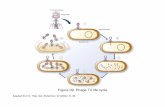
Fig.
1.
Sc
hem
e fo
r th
e co
nstr
uctio
n of
M
13m
p12
by
mol
ecul
ar
clon
ing
of a
syn
thet
ic
olig
onuc
leot
ide
poly
linke
r.

219
EXPERIMENTAL
(a) Synthesis of a 4Lnucleotide polylinker segment
Oligonucleotide linkers were synthesized by phos- photriester chemistry on polyamide/kieselguhr solid supports (Gait et al., 1982). The chains were purified by HPLC using Partisil 10 SAX resin. Fig. 1 outlines the method employed to synthesize a 41-nucleotide polylinker sequence with restriction endonuclease cleavage sites for X&I, BarnHI, SstI, XhoI, EcoRI and HindIII. Two partially complementary 27nucleotide chains, d(GGG- TTCTAGAGCTCGAGGATCCAAGCT) and d (GGGTCTAGAATTCAAGC’GGATCCTC) ,
were synthesized. The two chains overlap with 13 complementary base pairs at their 3’ ends. AMV polymerase (reverse transcriptase) was used to fill in the single-stranded regions to yield a duplex 41-mer. AMV polymerase was more effective at completing the duplex DNA (Bahl et al., 1977) than was the Klenow subfragment of Escherichiu coli DNA poly- merase (Kieny et al., 1983), which accumulated sub- stantial amounts of 39- and 40-nucleotide chains.
(h) Insertion of the polylinker sequence into bac- teriophage Ml3
The 41-mer duplex was initially cloned into the SmaI site of phage M13mp8 (Messing and Vieira, 1982) to purify the completed duplex and verify its sequence. The insertion of the 41-bp polylinker sequence produces a frame shift in the coding se- quence for the cr-peptide of the 1acZ’ gene product of M 13mp8. Phages harbouring the 41-mer are unable to complement lacZAM15 and do not pro- duce active /3-galactosidase and appear as white plaques. After transfection of E. coli strain JMlOl, and plating in the presence of BCIG indicator dye and IPTG inducer (Messing et al., 1977) a single white plaque was recovered from a pool of approx. 800 blue plaques. Sequencing of the inserted DNA by the dideoxy method (Sanger et al., 1980) showed that the 41-mer had been incorporated in the orien- tation depicted in Fig. 1. Replicative form DNA was prepared by primed synthesis on the Ml3 mp 8/41-mer phage DNA template. The DNA was then cleaved with Hind111 and recircularized with T4- DNA ligase. After transfection of JM 10 1, numerous
blue plaques were obtained. This lac’ phage has been named M 13mp 12, and may be used as a cloning vector in the mp series. It should be noted that M13mp12 does not contain any amber mutations. The M 13mp12 phage has unique cloning sites for EcoRI, SmaI,XbaI, SstI,XhoI, BumHI and Hind111 (Fig. 1).
(c) Construction of A2001
12001 is a derivative of i11059 (Karn et al., 1980; 1983) with an extended range of cloning sites. The phage is composed of three fragments, separated by two inverted polylinker sequences (Fig. 2). The left arm of 19 985 bp carries the NU 1 to J genes of I and specifies all the functions required for phage encap- sidation. The central 12785 bp fragment carries the red and gam genes under the control of p,_. This fragment is replaced by foreign DNA during cloning experiments. Deletion of the red and gum genes permits growth of recombinant 1 phage on E. coli
strains lysogenic for bacteriophage P2. The right 9 129-bp arm carries the A replication and lysis functions. Although both the left and right arms of the vector contain all the genes required for 2 growth and maturation, DNA composed of the two arms alone is too short to be viably packaged.
Fig. 3 is a genetic and restriction map of AcI857Sarn7 based on the DNA sequencing studies of Sanger et al. (1982). The map shows the positions of the deletions used in the construction of 12001. Endpoints for the deletions were obtained from the annotated A DNA sequence of Daniels et al. (1983). The restriction map of the lbio256 regions was reported by deWet et al. (1980) and the genetic physical map by Szybalski and Szybalski (1982). Restriction maps of 21059 are given in Karn et al. (1980; 1983).
Construction of 12001 involved a series of phage crosses and molecular cloning steps. In the first step, the target sites for EcoRI were removed by genetic crosses. The pACL29 plasmid and one of the dupli- cated 2 att sites were then replaced with an E. coli DNA fragment containing part of the bio operon. A synthetic oligonucleotide adapter was used to replace the BamHI sites with XbaI sites, into which the polylinker sequences were cloned. Finally, a short right arm devoid of EcoRI, HindIII, BamHI and XhoI sites was introduced by phage crosses. This

gamma
cos L cos R
10 - 23 kb inserts Sst 19989 RI 32746 Xho 19992 Hind 32752
Restriction Barn 19997 map
Barn 32758 HI;:’ ~~~o~3 Sst 32766
Xho 32763 Xba 19985, 20013 Xba 32742, 32770
Kpn 18556 Kpn 17053 j
I
!
Bgl 415
tl
Sal 25290 Sal 24791
1 BglZ9023 12gl56Bgl 34302
Bgl 26157 gl33987 Bgl 34953 k Bgl 35013
Fig. 2. Genetic and restriction maps of 12001. The coding sequences for 1 protein are drawn as boxes. The Z genes in boxes above the line are translated from right to left, genes in boxes below the line area translated from left to right. Genes that are essential for 1 growth and regulation are shaded, dispensable genes are shown as open boxes. The bio substitution is shown as a striped box. The 1 aft site is shown by a solid circle. The restriction map was calculated from the nucleotide sequence of lcI857Sam7 of Sanger et al. (1982) using the deletion endpoints of Daniels et al. (1983). The bio region in 12001 has not been sequenced, but a detailed restriction map was obtained from deWet et al. (1980) (see also Szybalski and Szybalski, 1982). Restriction cut sites for SsrI (Sst),XhoI (Xho), Bum HI (Barn), Hind111 (Hin), EcoRI (RI), XbaI (Xba), KpnI (Kpn), Sal1 (Sal), and Bg/II (Bgl) are shown. Numbers refer to the position of the tirst nucleotide in the recognition sequence taken from the nucleotide sequence of r’.. The total length of 12001 is 41.9 kb. Fragments between 10 and 23 kb may be cloned between the polylinker sequences at residues 19985 to 20013 and residues 32742 and 32770. Note that after insertion of new fragments, the gum and exe (red) genes are deleted, giving the recombinants a Spi _ phenotype. Recombinants retain a single d art site and remove the duplication of the N gene. For genotype of 12001 see Table I and Fig. 3.
Table I
Phage lambda cloning vectors with positive selection for inserts
Strain Genotype”
Al059
11672
12004
A1259
11274 12053 12149
l2OOl
ial sBam1’ 6189 (int29 ninL44 ~I857 pACL29) d[Bam] KH54 sRI4” nin5
h/l sBam1” b189 (i&29 sRI3” ninL44 ~I857 pACL29) d[Bam] KH54 sRI4” nin5 sRI5” sHind7”
ha sBam1” b189 a&. int29 sRI3” ninL44 EsHind3[bio] d[Bam] ~1857 sRI4” ninJ sRI5”
hd sBam1” Earn2001 Earn424 b189 (int29 ninL44 pACL29)
&Barn] KH54 sRI4” nin5 X/z01 linker in 1059 BamHI sites XbuI linker in 2004 BamHI sites hA sBam1” b189 ad. C sBam3 [XbaI] ninL44 I: sHind3[bio]
X[XbaI] d[Bam] ~1857 hl sBam1’ b189 attlZ I: sBam3[polylinker] ninL44 ZsHind3[bio]
Z[polylinker] d[Bam] KH54 sR14” nin5 sRI5” sHind7”
1 chi Cloning sites Capacity (kb)
D BumHI IO-23
c BumHI lo-23
C Bum HI 8-21
D BamHI IO-23
D BumHI, XhoI IO-23 C XbaI 8-21 C XbaI 5-18
C XbuI, SstI, XhoI, lo-23 BamHI, EcoRI, Hind111
a See Figs. 2 and 3.

short arm includes the chic mutation, to ensure
efficient growth of the Spi- recombinants. Table I lists the derivatives of 11059 constructed
during the course of this work. Original sources for 2 mutations and bacterial strains are given in Brenner et al. (1982) and Kam et al. (1983). In the sections that follow we describe the structures of these phages in detail.
(i) 11672 A 1059 was constructed by cloning the BglII frag-
ment of the A imm region (residues 35711 to 38103) into the BamHI sites of a small ColEl plasmid, pACL29, “lifting” the plasmid into phage arms, and crossing in various markers (Karn et al., 1980; Bren- ner et al., 1982). One of the intermediates in the construction was 1983, $80d[att80-cIIIlc1857 pACL29 attl b1319 $8lcIts chiD. This phage was crossed with the phasrnid, 11321, sBam1” b189 (pACL29) d[att80-cIII] KH54 chic sRI4” nin5 sRI5” sHind7” and recombinants were selected on strains resistant to I and immune to $81. The result- ing phage was 11341, $80d[att80-cIIIlcI857 pACL29 attl d[ Bam] KH54 chiC sRI4” nin5 sRI5’
221
sHind7”. A1321 was crossed with 11543, sBam1” b 189 int29 sRI3” ninL44 ~1857 chic, and recom-
binants were selected on a grolv strain resistant to $80 yielding 11672, sBam1” b189 (int29 sRI3” ninL44 ~1857 pACL29) d[Bam] KH54 c&C sRI4” nin5 sRI5” sHind7”. This phage is similar to 11059, but has no EcoRI sites on the central frag- ment and the right arm and no Hind111 sites on the right arm. The chiD mutation used in 11059 was replaced by chiC because it proved difficult to com- bine the sRI4” and the sHind7” mutations with chiD.
(ii) 12004
The pACL29 region of 21672 was replaced by a fragment of the bio operon of E. co& as follows. Charon 4a, which contains the bio256 substitution, was digested with EcoRI (cuts to the right of bio operon; Szybalski and Szybalski, 1982) and the cohesive ends tilled in by treatment with AMV poly- merase and each of the deoxynucleoside triphos- phates. The phage DNA was then digested with BamHI (cuts in gene bioA; Szybalski and Szybalski, 1982) and the EcoRI-BamHI fragment from the
b189 ’
ABarn- KH54 nin5
I ninL44H k---i -
Lambda gamma
int exe \ clll N cl
Nu 1 W E H J CII 0 P QSRRZ Barn RI
bio256 Restriction map
RI 21226 RI 26104 RI 31747 RI 39168 RI 44972 ! IIL I
Barn 5509 Barn 22346 Barn 27972 Barn 34499 Barn 41732 t I
Hind 2313: ‘T” 25’57 w
H,ind 27479 Hind36895 Hind 37459 Hind 44141
i I I Kpn 18556
Kpn 17053 I
Sal 33244 Sal 327451
I II i
+- Xba 24508
I 4 Sst 25877
Sst 247721
Xho 33498
Fig. 3. Restriction and gene map of kI857Sam7. The positions of the deletions and substitutions used in the construction of 12001 are
shown above and below the gene map. (For gene designations and symbols see Szybalski and Szybalski, 1979; Daniels et al., 1983, and
Fig. 2.)

Barn Ii I f I30 RI 1 Hind Ill
-Left arm
-Central fragment
-Right arm
Fig. 4. Analysis ofrestriction fragments from 1 vectors. 11059,A12?4,21672,R2QO4,12053 and 12001 DNA were digested~th BuwzHI, EcoRI or NindIfl and the DNA fragments separated on 0.40/, agarose gels. The positions of the left arm, central Fragment and right arm of 12001 are indicated on the right of the figure. Note that substitution of the pACL29 region (present in 11059, A1274 and A1672) with the bio fragment (present in 12004, X%53 and d2001) reduces the central fragment length by approximately 500 bp. In 12053 the right arm carries an intact 2 immunity region and migrates more slowly than the right arms of the other pbage, which are reduced in size by the KH54 deletion. The minor fragments seen migrating more slowly than the left arms of the digested phage result from the annealing of the left and right arms of the phage at their cos sites.
b&256 region purified by ele~trophoresis on low melting temperature agarose gels. This fra~ent contained one blunt end and one BarnHI sticky end. The bio fragment was inserted into the two phage arms by blunt-end ligation on one side and BarnHI- cohesive-end ligation on the other. A left phage arm was obtained by N&zdIII cleavage of A1672 DNA. This arm contained the intact hJ, b189 arm and a portion of the central fragment containing attA int29 sRI3’ ninL44, terminating at the first Hind111 site in the c1 gene (residue 36895 in the wild-type sequence).
The ~j~dII1 site was tilled in with AMV polymerase and purified by agarose gel electrophoresis. The right arm of the phage was derived from the phasmid 21129 by BarnHI digestion, and carried d[ Barn]] ~1857 chic sRI4” nin.5 sRI5”. This arm was also purified by gel electrophoresis. After ligation with T4 DNA ligase and in vitro packaging, a large number of phage plaques were obtained. These phages were shown by restriction mapping to contain the desired three fragments and an isolate was named ~2~~4 (Table I, Fig. 4).

223
(iii) 12053 In 12004, two BamHI sites flank the central frag-
ment. In order to replace the BamHI sites with XbaI sites, the decamer oligonucleotide adapter d(GATCTCTAGA) was synthesized. Although this oligonucleotide is capable of self-annealing to form concatemers, the major species at 20°C (as seen on a native polyacrylamide gel) is a dimer with a double- stranded hexanucleotide sequence and 5’ GATC cohesive ends. After enzymatic 5’ phosphorylation, the adapter was ligated to BamHI-digested 12004 DNA. Reconstituted phage were examined by re- striction enzyme mapping. Most phage had single adapters cloned in one of the two BamHI sites. 12053 is a derivative of 12004 in which both BamHI sites were destroyed by insertion of the XbaI adapters.
(iv) 12522 The synthetic polylinker was recovered from the
M13mp8/41-mer clone by digestion with XbuI. This was cloned into XbaI digested 12053. Phages containing the polylinker were detected by plaque hybridization to probes prepared from the original synthetic 27-mers by 32P-labelling with T4 poly- nucleotide kinase and [Y-~~P]ATP. A clones harbouring polylinkers were checked by restriction mapping. 22522 had polylinkers inserted at both XbuI sites. The linker sequences are inverted with respect to one another, with the EcoRI sites of each linker in the inner positions.
(v) 12001 The right arm of A2522 carrying ~1857 was replac-
ed by a shorter right arm containing the KH54 deletion and devoid of Hind111 sites. In the first step, the il immunity region was replaced by Avir
(Avlv2v3), 12522 was crossed with 12188, $80 a~80 to& trpD’ d[Bam] Avir sRI4” Pam1001 nin5
sHind7”, and recombinants selected on ;1 lysogens resistant to $80. The resultant phage, 12537, was sBam1” b189 uttA Z[polylinker] sRI3” ninL44 Z[bio] Z[polylinker] d[Bam] Ivir Pam1001 nin5
sHind7”. Next, the Avir Pam1001 arm was replaced by crossing with 21277, $80 act80 sRI480” tonB
trpD’ d[Bam] KH54 chic sRI4” nin5 sRI5” sHind7” and selecting for recombinants on su’ strains resistant to $80. The resulting phage was 12001, sBam1’ b 189 Z~Bam3[polylinker] sRI3’
ninL44 Z,sHind3[bio]C[polylinker] d[Bam] KH54 chic sRI4” nin5 sRI5” sHind7”.
(vi) Other derivatives of AI059
Several other derivatives of L 1059 were construct- ed during this work, and these are listed in Table I. 12 149 has a longer right arm which includes the nin5
region and an XbaI linker. A1259 introduces Eam and Kam mutations into 11059.
(d) Applications and special features of A2001
22001 closely resembles 21059 in its structure, and may be used in the same way. Selection of recombinants on P2 lysogens reduces the numbers of parental phage in genomic libraries by 4 to 5 orders of magnitude. This allows these vectors to be used efficiently without biochemical separation of the vec- tor arms.
22001 contains a pair of polylinker sequences and extends the range of fragments that may be directly cloned into A vectors. The linker has unique sites for XbuI, SstI, XhoI, EcoRI and BumHI, and the phage accommodates DNA fragments from 10 to 23 kb in size. Fragments generated with Sal1 (G/TCGAC) may be inserted into the XhoI sites since both enzymes produce TCGA ends. Similarly fragments prepared with BclI (T/GATCA), XhoII (Pu/GATCPy), Bg/II (A/GATCT) and Suu3A (/GATC) may be cloned directly into the BumHI (G/GATCC) sites. Thus, a total of eleven different restriction enzyme specificities may be used to clone fragments into 1200 1.
Partial digestion of DNA with Suu3A proved an
effective method for generating nearly random 15- to 23-kb fragments of genomic DNA, suitable for gene library construction (Karn et al., 1980; Kam, 1983; Frischauf et al., 1983). When Suu3A fragments are cloned into BumHI sites, on average only 25 y0 of the BumHI sites are retained. The loss of BumHI sites complicates the analysis of Suu3A fragments insert- ed into A1059. In 12001, the flanking XbuI, SstI and XhoI sites may be used to separate inserted frag- ments from the vector arms. Similarly, although cloning of Sal1 fragments into the XhoI sites creates a new sequence resistant to cleavage by both en- zymes, these fragments can be recovered by SstI or XbuI cleavage of the vector arms.
The polylinker sequence in 12001 also allows a

224
method of biochemical selection for inserts involving multiple restriction enzyme digestions of the vector DNA to limit recovery of the original vector se- quence. For example, in a Sau3A cloning experiment using 22001, the vector may be cleaved with both EcoRI and BamHI and the short linker oligo- nucleotide removed by gel filtration or selective pre- cipitation of the vector arms with polyethylene glycol. Frischauf et al. (1983) reported a two- to five-fold increase in recombinants on selective strains when the EMB03 derivative of A1059 was treated in this manner.
Finally, we note that the replacement of the pACL29 sequence by a fragment of the E. coli bio
operon allows genomic libraries in A2001 to be screened directly with probes prepared in homo- logous plasmid vectors such as pBR322.
ACKNOWLEDGEMENTS
We thank Mohinder Singh for help with oligo- nucleotide synthesis, and Leslie Barnett for assis- tance with ,? genetics. Phage strains may be obtained from Leslie Barnett. Joachim Messing has kindly agreed to number our new Ml3 vector (M13mp12) in accordance with his nomenclature. Hans Matthes was supported by a postdoctoral fellowship from EMBO.
REFERENCES
Bahl, C.P., Wu, R., Stawinsky, J. and Narang, S.A.: Minimal
length of the lactose operator sequence for specific recog-
nition by the lactose repressor. Proc. Natl. Acad. Sci. USA
74 (1977) 966-990.
Becker, A. and Gold, M.: Isolation of the bacteriophage I A gene
protein. Proc. Natl. Acad. Sci. USA 72 (1975) 581-585.
Benton, W.D. and Davis, R.W.: Screening Agt recombinant
clones by hybridization to single plaques in situ. Science 196
(1977) 180-182.
Brammar, W.J.: Vectors based on bacteriophage I, in William-
son, R. (Ed. Genetic Engineering), Academic Press, New
York, 1982, pp. 53-79.
Brenner, S., Cesareni, G. and Karn, J.: Phasmids: hybrids
between ColEl plasmids and E. coli bacteriophage lambda.
Gene 17 (1982) 27-44.
Daniels, D., Schroeder, J.L., Szybalski, W., Sanger, F., Coulson,
A.R., Hong, G.F., Hill, D.F., Petersen, G.B. and Blattner,
F.R.: Complete annotated I sequence, in Hendrix, R.W.,
Roberts, J.W., Stahl, F.W. and Weisberg, R.A. (Eds.),
Lambda II. Cold Spring Harbor Laboratories, Cold Spring
Harbor, NY, 1983, pp. 519-676.
deWet, J.R., Daniels, D.L., Schroeder, J.L., Williams, B.G.,
Denniston-Thompson, K., Moore, D.D. and Blattner, F.R.:
Restriction maps of twenty-one Charon vector phages. J.
Virol. 33 (1980) 401-410.
Frischauf, A.-M., Lehrach, H., Poustka, A. and Murray, N.:
Lambda replacement vectors carrying polylinker sequences.
J. Mol. Biol. 170 (1983) 827-842.
Gait, M.J., Matthes, H.W.D., Singh, M., Sproat, B.S. and
Titmas, R.C.: Rapid synthesis of oligodeoxyribonucleotides,
VII. Solid phase synthesis of oligodeoxyribonucleotide by a
continuous flow phosphotriester method on kieselguhr-
polyamide support. Nucl. Acids Res. 10 (1982) 6243-6254.
Hohn, B. and Murray, K.: Packaging recombinant DNA mole-
cules into bacteriophage particles in vitro. Proc. Natl. Acad.
Sci. USA 77 (1977) 3259-3263.
Karn, J.: Cloning with bacteriophage, in Flavell, R.A. (Ed.),
Techniques in the Life Sciences, Nucleic Acid Biochemistry
B501. Elsevier, Shannon, 1983, pp. 1-61.
Karn, J., Brenner, S., Barnett, L. and Cesareni, G.: Novel
bacteriophage 1 cloning vector. Proc. Natl. Acad. Sci. USA
77 (1980) 5172-5176.
Karn, J., Brenner, S. and Barnett, L.: New bacteriophage 1
vectors with positive selection for cloned inserts. Methods
Enzymol. 101 (1983) 3-20.
Kieny, M.P., Lathe, R. and Lecocq, J.P.: New versatile cloning
and sequencing vectors based on bacteriophage M 13. Gene
26 (1983) 91-99.
Messing, J. and Vieira, J.: A new pair of M 13 vectors for selecting
either DNA strand of double-digest restriction fragments.
Gene 19 (1982) 269-276.
Messing, J., Gronenborn, B. Muller-Hill, B., and Hofschneider,
P.H.: Filamentous coliphage M 13 as a cloning vehicle: inser-
tion of a Hind111 fragment of the lac regulatory region in the
Ml3 replicative form in vitro. Proc. Natl. Acad. Sci. USA 74
(1977) 3642-3646.
Murray, N.E.: Phage lambda and molecular cloning, in Hendrix,
R.W., Roberts, J.W., Stahl, F.W. and Weisberg, R.A. (Eds.),
Lambda II. Cold Spring Harbor Laboratory, Cold Spring
Harbor, NY, 1983, pp. 395-432.
Sanger, F., Coulson, A.R., Barrell, B.G., Smith, A.J.H. and Roe,
B.A.: Cloning in single-stranded bacteriophage as an aid to
rapid DNA sequencing. J. Mol. Biol. 162 (1980) 161-178.
Sanger, F., Coulson, A.R., Hong, G.F., Hill, D.F. and Petersen,
G.B.: Nucleotide sequence of bacteriophage Iz DNA. J. Mol.
Biol. 162 (1982) 729-773.
Steinberg, N., Tiemeier, D. and Enquist, L.: In vitro packaging
of a I Dam vector containing EcoRI DNA fragments of
Escherichia coli and phage Pl. Gene 1 (1977) 255-280.
Szybalski, E.H. and Szybalski, W.: A comprehensive molecular
map of bacteriophage lambda. Gene 7 (1979) 217-270.
Szybalski, E.H. and Szybalski, W.: A physical map of the
Escherichiu coli bio operon. Gene 19 (1982) 93-103.
Williams, B.G. and Blattner, F.R.: Bacteriophage I vectors for
DNA cloning, in Setlow, J.K. and Hollaender, A. (Eds.),
Genetic Engineering Vol. 2. Plenum, New York, 1980,
pp. 201-235.
Communicated by K. Chater.