The Institute of Global Health Innovation and charge state ... · Results (simulated data) MALDI 0...
Transcript of The Institute of Global Health Innovation and charge state ... · Results (simulated data) MALDI 0...

Results (simulated data)
MAL
DI
0 0.5 10
0.5
11 Protein
seaMS λ = 0.001seaMS λ = 0.01seaMS λ = 0.1Isotope Wavelet + Baseline RemovalNITPICK + Post−processing 0 0.5 1
0
0.5
14−Mix
0 0.5 10
0.5
116−Mix
ESI L
ow N
oise
0 0.5 10
0.5
1
0 0.5 10
0.5
1
0 0.5 10
0.5
1
Prec
isio
n (1
− F
alse
Dis
cove
ry R
ate)
ESI H
igh
Noi
se
0 0.5 10
0.5
1
0 0.5 10
0.5
1
Recall (Sensitivity)0 0.5 10
0.5
1
a New isotope distRibutioN model
a sparse signal restoration framework for simultaneous baseline estimation, deisotoping and charge state deconvolution of complex mass spectra
andrew w dowsey and Guang-Zhong Yang, Hamlyn Centre, Institute of Global Health Innovation, Imperial College London, UK
oveRview
We present an automatic feature detection algorithm for general mass spectrometry which markedly increases accuracy and robustness over existing techniques. It works directly on the raw signal and the novel aspects are:
» A new isotope distribution model that greatly improves on the ubiquitous averagine approach by efficiently:• Encapsulating the full range of isotope distributions at each m/z interval.• Ensuring distributions from other charge states are fully differentiated.
» Concurrent baseline modelling that dynamically separates chemical noise from peptide isotope signals, of particular benefit with MALDI.
» A robust and scalable sparse optimisation framework with Poisson noise model.
The Hamlyn Centre The Institute of Global Health Innovation
iNtRoductioN
backGRouNdRecent feature detection methods take roughly three different approaches [1]:
» Wavelet denoising / matched filtering e.g. Isotope Wavelet [2].• Fast linear solution to a nonlinear problem.• Issues with separating coincident isotope distributions.
» Mixture modelling e.g. Expectation-Maximisation or MCMC.• High separation power, can compute posterior distributions.• Dependent on prior seeding, generally not scalable.
» Sparse regression e.g. NITPICK [3].• High separation power but no posterior distributions (yet).• No prior seeding (templates placed at all discretised m/z intervals).
iteRative shRiNkaGeSparse regression iteratively selects the template with the greatest signal response to add to the candidate set:
» So the algorithm can terminate when further candidates lack statistical support.
» But scalability is compromised as each pass only determines a single feature, and highly correlated features can be missed*.
Conversely, ‘iterative shrinkage’ simultaneously reduces the whole set of template weights until most end up at zero:
» This is highly scalable, which has led to the development of redundant dictionaries (large sets of templates) for the specialised encoding of signal families.
» However, the statistically suitable amount of shrinkage must be set in advance.
In this project, we have designed a set of trained dictionaries for isotope distributions at multiple charge states, and a redundant multiscale dictionary to represent the periodic chemical baseline. In this application we demonstrate results which are robust over a magnitude variation of the shrinkage parameter.
* The recent ‘Elastic-Net’ method adds candidates in groups to attempt to mitigate these issues.
simultaNeous chemical Noise modelliNG
coNclusioN
We present a sparse signal restoration framework that markedly improves feature detection for general mass spectrometry. The technique models raw signal directly and is not reliant on high resolution or a specific instrument configuration, also making it promising for pervasive and hand-held spectrometry. In the short-term:
» We aim to provide validation of quantification reliability and mass accuracy, and objective validation on real data by improved MS/MS protein identification [7].
» We wish to incorporate mass defect into the seaMS isotope distribution model.
The algorithm is also highly scalable. To this end, progress towards processing full LC/MS datasets is described in our other poster.
Results (Real data)
http://www.proteomegrid.org/
ackNowledGemeNts
Project funded by EPSRC UK grant EP/E03988X/1 awarded to AWD.
RefeReNces
[1] Dowsey et al., Proteomics 10, pp. 4226-4257, 2010 [2] Hussong et al., Bioinformatics 25, pp. 1937-1943, 2009 [3] Renard et al., BMC Bioinformatics 9, Article 355, 2008 [4] Dolui et al., Proc. 2011 IEEE International Symposium on Biomedical Imaging (ISBI) [5] Sun et al., BMC Bioinformatics 11, Article 490, 2010 [6] Bielow et al., Journal of Proteome Research, DOI: 10.1021/pr200155f, 2011 [7] Falkner et al., Journal of the ASMS 18, pp. 850-855, 2007
motivatioNNearly all current techniques detect peptide isotope distributions using a single template at each mass value. This template is either the averagine [2], fractional averagine [3] or an average distribution computed from a sequence database.
» Actual isotope distributions can deviate significantly from this template.
Rather, we propose learning n templates (‘factors’) for the isotope distributions in each mass interval and fit these to the data simultaneously.
» Any isotope distribution that can be made from a weighted sum of these factors can be represented without error, apart from mass defect (which is not modelled).
» For n = 3, Figure 1 shows a major decrease in encoding error compared to the averagine over a simulated tryptic digest of Swiss-Prot.
methodFor each charge/adduct state, the factors at each small m/z interval (e.g. 1Da) are learnt from the set of all possible distributions around that interval (e.g. ±1Da):
» Non-negative Matrix Factorisation (NMF) is the defacto approach for this task, but the optimisation is non-convex and the factors need not lie near the distributions.
» Adding a soft Minimum Volume Constraint (MVC) solves these problems by requiring the factors to be as similar (i.e. lie as near) as possible to each other and, as a side effect, be dissimilar to factors from other charge/adduct states.
Figure 2 provides a visual representation of the relationship between sets of isotopes distributions and their learnt factors.
motivatioNSince baseline removal is performed prior to feature detection, basic assumptions such as low frequency and/or exponential decay are used to avoid affecting peaks.
» By performing both simultaneously we improve the performance of both...
» ... and can differentiate peptide peaks from chemical noise peaks.
methodWe model chemical noise peaks rather than the baseline - the later is inferred by convolving the chemical noise peaks with the peak spread function.
» We use a dictionary of multiscale B-spline basis functions to model smoothly varying intensity between chemical noise peaks 1Da apart.
» Since the chemical noise offset is unknown and there may be multiple chains, the dictionary is duplicated at each sub-Dalton m/z location (see Figure 3)
We ran seaMS on a Bruker ultraFlex MALDI-ToF 7-Mix spectrum provided by [5]. The peak spread function was estimated using the technique described in our other
poster. Figures 4 & 5 demonstrate the recovered baseline and isotope distributions, including the one barely discernible from noise the methods tested in [5] missed.
spaRse RestoRatioN with poissoN Noise
Most techniques assume the simpler-to-handle case of Gaussian noise. The literature has suggested mainly Poissionian statistics mixed with Multinomial variation from isotope distributions. Poisson counting noise also explains the observed heteroskedasticity as ToF increases. We apply a Biggs-Andrews accelerated form of the recent Sparse Richardson-Lucy method [4] to the following formation model:
figure 1: Root-mean-square error (RMSE), averaged over 1Da intervals, between each trypically digested protein from Swiss-Prot and: (blue) the averagine template; (red) the seaMS regression fit to the seaMS model with 3 learnt factors.
figure 2: Isotope distributions for trypically digested proteins from Swiss-Prot of 1000±1 m/z for charge states 1,3,5, and their derived factors. (top) PCA plot applied only to provide a 2D visualisation of multidimensional isotope distribution variability (green circles) and to illustrate the position of the derived factors and averagine amongst them. Peptides within the blue triangles can be encoded perfectly. (bottom) The templates of the averagine and derived factors.
0 1000 2000 3000 4000 5000 6000 70000
0.01
0.02
0.03
0.04
0.05
0.06
mass/Daltons
Roo
t Mea
n Sq
uare
d Er
ror
Averagine3 Factors
0.25 0.3 0.35 0.4 0.45 0.5−0.1
−0.08
−0.06
−0.04
−0.02
0
0.02
0.04
Factor 1
Factor 2
Factor 3
1st Principal Component (72.3% of variance)2nd
Prin
cipa
l Com
pone
nt (2
7.6%
of v
aria
nce) 1000±1Da mass and H+ charge
Averagine
Averagine Factor 1 Factor 2 Factor 30
50
100
% In
tens
ity
0.02 0.04 0.06 0.08 0.1 0.12 0.14 0.16 0.18 0.2 0.220.15
0.16
0.17
0.18
0.19
0.2
0.21
0.22
Factor 1
Factor 2
Factor 3
1st Principal Component (94.0% of variance)
2nd
Prin
cipa
l Com
pone
nt (5
.7%
of v
aria
nce) 5000±5Da mass and 5H+ charge
Averagine
Averagine Factor 1 Factor 2 Factor 30
20
40
% In
tens
ity
0.05 0.1 0.15 0.2 0.25 0.3−0.2
−0.19
−0.18
−0.17
−0.16
−0.15
−0.14
−0.13
−0.12
−0.11
Factor 1
Factor 2
Factor 3
1st Principal Component (88.9% of variance)2nd
Prin
cipa
l Com
pone
nt (1
0.9%
of v
aria
nce) 3000±3Da mass and 3H+ charge
Averagine
Averagine Factor 1 Factor 2 Factor 30
20
40
% In
tens
ity
figure 3: Three representative L2-normalised B-spline basis function templates of increasing scale. Each basis function is sampled at 1Da intervals but offset by 0.33Da. A chemical noise chain is constructed from a weighted combination of templates.
0 10 20 30 40 50 60 70 80 90 1000
5
10
15
20
25
30
m/z
% In
tens
ity
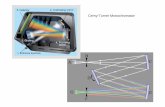
figure 7: Precision-Recall
plots compare Sensitivity
against the False Discovery Rate as an intensity
threshold on the set of detected
features is varied.
Nine datasets of simulated ToF data were generated using MSSimulator from the OpenMS TOPP pipeline [6]. 64 proteins were randomly selected from the Aurum dataset FASTA file [7], tryptically digested and analysed on:
» Separate spectra.
» A mixture of 4 proteins per spectra.
» A mixture of 16 proteins per spectra.
Noise in MSSimulator is simulated as a mixture of Poisson and Gaussian components. 3 different instrument configurations were tested, as derived from [5]:
» MALDI-ToF with a simple baseline* and a medium noise level.
» ESI with low noise, to test coincident peptide separation power.
» ESI with high noise, to test algorithm robustness.
Figure 6 illustrates the simulated data and Figure 7 presents Precision-Recall plots comparing seaMS with shrinkage λ to the Isotope Wavelet [2] and NITPICK [3].
* MSSimulator does not currently generate chemical noise peaks.
The results show:
» seaMS significantly outperforms the Isotope Wavelet and NITPICK in all tests.
» Too much shrinkage (λ = 0.1) can be harmful, particularly when estimating the baseline. The algorithm is robust to too little shrinkage, mainly affecting speed.
» NITPICK* is much improved by its post-processing technique to remove spurious peaks. seaMS does not employ post-processing but would actually also benefit.
» The Isotope Wavelet is less robust to noise. Even with prior OpenMS baseline subtraction, the technique struggled on the simulated MALDI data.
*NITPICK could not run all the tests as it is ‘proof of concept’ code which appears to be memory-limited.
figure 6: Representative simulated spectra and ground truth features for (top) MALDI 4-mix (middle) ESI Low Noise 16-mix and (bottom) ESI High Noise 1 protein.
400 1400 2400 3400 440002468
H+2H+
400 600 800 1000 1200 140002468
Inte
nsity
3H+4H+
400 600 800 1000 1200 140002468
m/z
5H+
g = P H BcB + AqcAq∈Q∑⎡
⎣⎢
⎤
⎦⎥
⎛
⎝⎜
⎞
⎠⎟
g is the observed spectrum.h is the varying peak spread function of the spectrometer (estimated in our other poster). a are the isotope distribution dictionaries over charge/adduct states Q.b is the multiscle baseline dictionary.c = {ca, cb} are the unknowns to estimate.
figure 4: seaMS results showing (green) original spectrum, (red) reconstructed baseline, (blue) detected isotope distributions and (cyan/magenta/yellow) factor responses.
2500 2600 2700 2800 2900 3000 3100 3200−500
0
500
m/z
Inte
nsity
930 935 940 945 950 955 960 965 970 975−1000
−500
0
500
1000
m/z
Inte
nsity