Structure peculiarities of α - crystallin studied by small angle neutron and X-ray scattering
description
Transcript of Structure peculiarities of α - crystallin studied by small angle neutron and X-ray scattering

STRUCTURE PECULIARITIES OF α-CRYSTALLIN STUDIED BY SMALL ANGLE NEUTRON AND X-RAY SCATTERING
TN Murugova1 OI Ivankov15 AI Kuklin13 KO Muranov2 NB Poliansky2 VM Garamus4 AV Krivandin2
1FLNP JINR Dubna Russia2Institute of Biochemical Physics of RAS Moscow Russia (the Tasking)3Moscow Institute of Physics and Technology Dolgoprudny Russia4Helmholtz-Zentrum Geesthacht Zentrum fuumlr Material- und Koumlstenforschung GmbH Geesthacht Germany5Taras Shevchenko National University of Kyiv Kyiv Ukraine
Function
bullThe main component of the vertebrate eye lens (300 mgml)
bull Provides a proper refractive index of the lens
bullChaperon-like activity (forms soluble complexes with destabilized proteins and prevents their unspecific aggregation and uncontrolled denaturation)
bullSlows down the age-dependent loss of the lens transparency (cataract)
Problem
bullHow does this protein protect the lensbullWhat is the mechanism of its binding of target proteins bullWhat is the role of oligomeric structure and subunit
exchange in this mechanism
Knowledge of the quaternary structure of α-crystallin is a missing link in defining a structure-function relationship and the possible role of the a-crystallin subunits in the lens and other organs in normal and abnormal conditions
The quaternary structure of α-crystallin is not known to date
Structure oligomeric protein
Monomers αA- and αB-crystallin20 kDa
Oligomer20-40 monomers400-1000 kDa
Models of α-crystallin structure
Aquilina J A and Morris A M Evidence for specific subunit distribution and interactions in the quaternary structure of α-crystallin2010httprouoweduauscipapers175
Open micellar configurationB GROTH-VASSELLI ET AL Exp Eye Res (1995) 61 249-253
Single particle reconstructions of native a-crystallin from Electron microscopyDA Haley et al J Mol Biol (2000) 298 261plusmn272
Micellar three-layer modelWalsh et al THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol 266 pp20079-200841991
3-layer modelTardieu et al I Mol Biol (1986) 192711 -724
A model of αB-crystallin with bound α-lactalbumin Electron microscopyScale bar represents 100 Aring J Horwitz Exp Eye Res (2003) 76 145-153
Why Small Angle Scattering
bullPolydisperse system variable size and number of monomers
bullDynamic system monomer exchange between oligomers
bullEfforts to crystallize α-crystallin have failed
bullThe aggregate is too large for high resolution 2D NMR
Small angle scattering allows to study structure of macromolecules (10-1000 Aring) in solution
ShapeSizeMolecular massVolume
Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
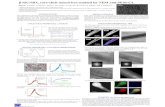
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Function
bullThe main component of the vertebrate eye lens (300 mgml)
bull Provides a proper refractive index of the lens
bullChaperon-like activity (forms soluble complexes with destabilized proteins and prevents their unspecific aggregation and uncontrolled denaturation)
bullSlows down the age-dependent loss of the lens transparency (cataract)
Problem
bullHow does this protein protect the lensbullWhat is the mechanism of its binding of target proteins bullWhat is the role of oligomeric structure and subunit
exchange in this mechanism
Knowledge of the quaternary structure of α-crystallin is a missing link in defining a structure-function relationship and the possible role of the a-crystallin subunits in the lens and other organs in normal and abnormal conditions
The quaternary structure of α-crystallin is not known to date
Structure oligomeric protein
Monomers αA- and αB-crystallin20 kDa
Oligomer20-40 monomers400-1000 kDa
Models of α-crystallin structure
Aquilina J A and Morris A M Evidence for specific subunit distribution and interactions in the quaternary structure of α-crystallin2010httprouoweduauscipapers175
Open micellar configurationB GROTH-VASSELLI ET AL Exp Eye Res (1995) 61 249-253
Single particle reconstructions of native a-crystallin from Electron microscopyDA Haley et al J Mol Biol (2000) 298 261plusmn272
Micellar three-layer modelWalsh et al THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol 266 pp20079-200841991
3-layer modelTardieu et al I Mol Biol (1986) 192711 -724
A model of αB-crystallin with bound α-lactalbumin Electron microscopyScale bar represents 100 Aring J Horwitz Exp Eye Res (2003) 76 145-153
Why Small Angle Scattering
bullPolydisperse system variable size and number of monomers
bullDynamic system monomer exchange between oligomers
bullEfforts to crystallize α-crystallin have failed
bullThe aggregate is too large for high resolution 2D NMR
Small angle scattering allows to study structure of macromolecules (10-1000 Aring) in solution
ShapeSizeMolecular massVolume
Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Problem
bullHow does this protein protect the lensbullWhat is the mechanism of its binding of target proteins bullWhat is the role of oligomeric structure and subunit
exchange in this mechanism
Knowledge of the quaternary structure of α-crystallin is a missing link in defining a structure-function relationship and the possible role of the a-crystallin subunits in the lens and other organs in normal and abnormal conditions
The quaternary structure of α-crystallin is not known to date
Structure oligomeric protein
Monomers αA- and αB-crystallin20 kDa
Oligomer20-40 monomers400-1000 kDa
Models of α-crystallin structure
Aquilina J A and Morris A M Evidence for specific subunit distribution and interactions in the quaternary structure of α-crystallin2010httprouoweduauscipapers175
Open micellar configurationB GROTH-VASSELLI ET AL Exp Eye Res (1995) 61 249-253
Single particle reconstructions of native a-crystallin from Electron microscopyDA Haley et al J Mol Biol (2000) 298 261plusmn272
Micellar three-layer modelWalsh et al THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol 266 pp20079-200841991
3-layer modelTardieu et al I Mol Biol (1986) 192711 -724
A model of αB-crystallin with bound α-lactalbumin Electron microscopyScale bar represents 100 Aring J Horwitz Exp Eye Res (2003) 76 145-153
Why Small Angle Scattering
bullPolydisperse system variable size and number of monomers
bullDynamic system monomer exchange between oligomers
bullEfforts to crystallize α-crystallin have failed
bullThe aggregate is too large for high resolution 2D NMR
Small angle scattering allows to study structure of macromolecules (10-1000 Aring) in solution
ShapeSizeMolecular massVolume
Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Structure oligomeric protein
Monomers αA- and αB-crystallin20 kDa
Oligomer20-40 monomers400-1000 kDa
Models of α-crystallin structure
Aquilina J A and Morris A M Evidence for specific subunit distribution and interactions in the quaternary structure of α-crystallin2010httprouoweduauscipapers175
Open micellar configurationB GROTH-VASSELLI ET AL Exp Eye Res (1995) 61 249-253
Single particle reconstructions of native a-crystallin from Electron microscopyDA Haley et al J Mol Biol (2000) 298 261plusmn272
Micellar three-layer modelWalsh et al THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol 266 pp20079-200841991
3-layer modelTardieu et al I Mol Biol (1986) 192711 -724
A model of αB-crystallin with bound α-lactalbumin Electron microscopyScale bar represents 100 Aring J Horwitz Exp Eye Res (2003) 76 145-153
Why Small Angle Scattering
bullPolydisperse system variable size and number of monomers
bullDynamic system monomer exchange between oligomers
bullEfforts to crystallize α-crystallin have failed
bullThe aggregate is too large for high resolution 2D NMR
Small angle scattering allows to study structure of macromolecules (10-1000 Aring) in solution
ShapeSizeMolecular massVolume
Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Models of α-crystallin structure
Aquilina J A and Morris A M Evidence for specific subunit distribution and interactions in the quaternary structure of α-crystallin2010httprouoweduauscipapers175
Open micellar configurationB GROTH-VASSELLI ET AL Exp Eye Res (1995) 61 249-253
Single particle reconstructions of native a-crystallin from Electron microscopyDA Haley et al J Mol Biol (2000) 298 261plusmn272
Micellar three-layer modelWalsh et al THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol 266 pp20079-200841991
3-layer modelTardieu et al I Mol Biol (1986) 192711 -724
A model of αB-crystallin with bound α-lactalbumin Electron microscopyScale bar represents 100 Aring J Horwitz Exp Eye Res (2003) 76 145-153
Why Small Angle Scattering
bullPolydisperse system variable size and number of monomers
bullDynamic system monomer exchange between oligomers
bullEfforts to crystallize α-crystallin have failed
bullThe aggregate is too large for high resolution 2D NMR
Small angle scattering allows to study structure of macromolecules (10-1000 Aring) in solution
ShapeSizeMolecular massVolume
Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Why Small Angle Scattering
bullPolydisperse system variable size and number of monomers
bullDynamic system monomer exchange between oligomers
bullEfforts to crystallize α-crystallin have failed
bullThe aggregate is too large for high resolution 2D NMR
Small angle scattering allows to study structure of macromolecules (10-1000 Aring) in solution
ShapeSizeMolecular massVolume
Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Distance distribution function p(r)(indirect Fourier transform)
001 01
1E-3
001
01
1
10
SANS SAXSIn
tens
ity a
u
q A-1
Curves of SANS and SAXS for the same sample of α-crystallin (concentration 9 mgml)
0 50 100 150 20000
02
04
06
08
10
p(r)
r A
SANS SAXS
Dmax
Distance distribution functions for α-crystallin molecule calculated from SANS and SAXS curves by indirect transform
max max
2 2
0 0
( ) ( ) D D
gR p r r dr p r dr max
0
(0) 4 ( )D
I p r dr
Program GNOM httpwwwembl-hamburgdebiosaxssoftwarehtml
Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Molecular mass M volume V and hydration value
M = (600plusmn17) kDa rarr Vah = Mν = (740plusmn20)103 Aring3 (anhydrous volume)
000 002 004 006 008 010 012 014 016 018 02000000
00002
00004
00006
00008
00010
Iq2
q A-1
Hydrous volume Vh
(Porod invariant) (Fig 3)
22 0 hV I Q2
0
( )where Q q I q dq
Hydration value for α-crystallin026 g of water per g of protein
Iq2 vs q plot for estimation of Porod volume The Porod volume was calculated by program PRIMUS [20]
2 2
(0)A
S
I NM
c
where I(0)=297 cm-2 (from indirect transformation)NA = 60221023 (Avogadro constant)c=9 mgml (concentration of protein in the sample) (average scattering density of protein) [8]ρs = -05481010cm-2 (scattering density of the solvent) (specific volume) [13]
10 219 10 cm
3(0745 0014) cm mg
Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Shape of α-crystallin
001 011E-4
1E-3
001
01
1
SANS experiment
model of 3-axial ellipsoid (programm FITTER)
3-axial ellipsoidal shell
(programm SASFIT)
Inte
nsi
ty
cm-1
q A-101
01
1
10
SAXS experiment
3-exial ellipsoid (program FITTER)
3-exial ellipsoidal shell (programm SASFIT)
Inte
nsi
ty
au
q A-1
Approximation of SANS and SAXS data for α-crystallin by model of 3-axial ellipsoid and 3-axial ellipsoidal shell
Model ellipsoidal shell for α-crystallin
3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

3D dumm-models
x y z( )
x y z( )
x y z( )ADUMM-model from SANS
BDUMM-model from SAXS
C3-exis ellipsoid from SANS
Program DAMMIF httpwwwembl-hamburgdebiosaxssoftwarehtml
001 01
1E-3
001
01
1
SANSchi2=142
Inte
nsi
ty c
m-1
q A-1
001 003 005 007 00901
1
10
SAXSchi2=127
Inte
nsi
ty r
elu
n
q A-1
Approximation of experimental data by curves for the 3D-damm models
The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

The summary table of structural parameters of α-crystallin obtained from small angle scattering
Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell

Conclusions
bullThe average values of structural parameters of alpha-cryslallin have been obtained
(Volume size hydration value)
bullEllipsoid-like shape
bullThe presence of internal cavity is available
bullIn case of model ellipsoidal shell the monomers form an unilayer shell










![Characterization of an antibody that recognizes peptides ... · in αA-crystallin (Asp 58 and Asp 151) [3], αB-crystallin (Asp 36 and Asp 62) [4], and βB2-crsytallin (Asp 4) [5]](https://static.fdocument.org/doc/165x107/5ff1e68e89243b57b64135f8/characterization-of-an-antibody-that-recognizes-peptides-in-a-crystallin-asp.jpg)








