Running title: Synthesis of (1 β D-glycans* · 04/11/2010 · terminus of the acceptor glucan...
Transcript of Running title: Synthesis of (1 β D-glycans* · 04/11/2010 · terminus of the acceptor glucan...
-
1
Running title: Synthesis of (1→4)-β-D-glycans*
Title: Update on Mechanisms of Plant Cell Wall Biosynthesis: How plants make
cellulose and other (1→4)-β-D-glycans
Author: Nicholas C. Carpita
Department of Botany & Plant Pathology, and
Bindley Bioscience Center
Purdue University
915 West State Street
West Lafayette, IN 47907-2054
For correspondence:
Tel. +1-765-494-4653
FAX: +1-765-494-0363
E-mail: [email protected]
1This material is based upon work supported as part of the Center for Direct Catalytic
Conversion of Biomass to Biofuels (C3Bio), an Energy Frontier Research Center funded by the
U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, Award Number
DE-SC0000997
*Corresponding author; e-mail [email protected].
The author responsible for distribution of materials integral to the findings presented in this
article in accordance with the policy described in the Instructions for Authors
(www.plantphysiol.org) is: Nicholas C. Carpita ([email protected]).
Plant Physiology Preview. Published on November 4, 2010, as DOI:10.1104/pp.110.163360
Copyright 2010 by the American Society of Plant Biologists
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
2
The discovery of a gene that encodes a cotton (Gossypium hirsutum) cellulose synthase (Pear et
al., 1996) revolutionized and invigorated the plant cell wall community to find the genes that
encode the machinery of cell wall polysaccharide synthesis. The landscape was framed by the
completion of the genome sequence of Arabidopsis thaliana (Arabidopsis Genome Initiative,
2000), which gave a complete gene inventory for a model plant species, but one with many genes
yet to be annotated for function. An estimated 10% of the plant genome, about 2,500 genes, is
devoted to construction, dynamic architecture, sensing functions and metabolism of the plant cell
wall. Based largely on prior discoveries of function in prokaryotic organisms, most of the
tentatively annotated genes are organized into gene families for substrate generation, glycosyl
transfer, targeting and trafficking, cell wall rearrangement and modification by hydrolases,
esterases, and lyases (Yong et al., 2005; Penning et al., 2009). However, the biochemical
activities of most enzymes involved in glycosyl transfer within these families remain to be
verified, and an additional 40% of the genome encodes genes for whose function is not known.
As many of these proteins contain secretory signal peptides (Arabidopsis Genome Initiative,
2000), it is reasonable to infer that some have roles in cell wall construction.
The Arabidopsis cellulose synthase/cellulose-synthase-like (CesA/Csl) gene superfamily,
which includes 10 CesA genes and 29 Csl genes in six distinct groups, was one of the first large
families to be described (Richmond and Somerville, 2000), and comparative analyses of a
reference dicot, Arabidopsis, to a reference grass, rice (Oryza sativa), revealed substantive
differences in family structures, adding two groups not seen in the dicot genome (Hazen et al.,
2002). Extension of these annotations to compare all cell wall-related gene families of the
grasses with those of the dicots reveals some correlation of family structure with the differences
between plants with Type I walls, with those of the grasses with Type II walls (Fig. 1A; Penning
et al., 2009). For CesAs and certain Csl genes, establishment of specific function for the
synthases they encode comes from analysis of mutants lacking a particular function and in some
specific examples, by heterologous expression. However, genetic approaches alone do not
inform us about the biological mechanism of synthesis. The knowledge gained from molecular
genetic approaches now needs to be augmented by biochemical and cell biological approaches to
achieve a greater understanding of proteins and their interactions within a synthase complex,
their organization at membranes, and their dynamics. This update focuses on the biochemical
mechanisms of the synthesis of a single type of linkage, the (1→4)-β-D- linkage, in which one
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
3
sugar is inverted nearly 180° with respect to each neighboring sugar in the chain. This linkage
presents a unique steric problem for processive catalysis that all living organisms have solved,
but we are still struggling to understand.
This update reviews our present state of knowledge of the biochemical mechanisms of
polysaccharide synthesis, including some classic discoveries, and presents an alternative
hypothesis on the biochemical mechanisms and organization of complexes involved in synthase
reactions that yield (1→4)-β-D- linkages.
Cellulose synthesis
In flowering plants, cellulose is a para-crystalline array of about two to three dozen (1→4)-β-D-
glucan chains. Microfibrils of 36 glucan chains have a theoretical diameter of 3.8 nm, but X-ray
scattering and NMR spectroscopy indicate some microfibril diameters could be as small as 2.4
nm, or about two dozen chains (Kennedy et al., 2007). The microfibrils are synthesized at the
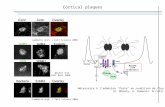
plasma membrane by terminal complexes of six-membered ‘particle rosettes’ that produce a
single microfibril (Giddings et al., 1980; Mueller and Brown, 1980). Thus, each of the six
components of the particle rosette is expected to synthesize four to six of the glucan chains, and
24 to 36 chains are then assembled into a functional microfibril (Doblin et al., 2002). In freeze-
fracture, the particle rosettes, found only on the P-face of the membrane, are about 25 nm in
diameter, but this size represents only the membrane-spanning and short exterior domains (Fig.
2A). Hidden in surface views of rosette structures in the plasma membrane, the much larger
catalytic domains of the cellulose synthases are estimated to be 50 nm wide and extend 35 nm
into the cytoplasm (Bowling and Brown, 2008), a feature that has escaped consideration in many
published models of the rosette structure (Fig. 2B and C).
Cellulose synthase is an ancient enzyme (Nobles et al., 2001), and cellulose synthase genes
in green algae are homologous to those of flowering plants (Roberts et al., 2002). The deduced
amino acid sequences of CesAs share regions of similarity with the bacterial CesA proteins,
namely the four catalytic motifs containing the D, DxD, D, Q/RxxRW that are highly conserved
among those that synthesize several kinds of (1→4)-β-D-glycans (Saxena et al., 1995). The
higher-plant CesA genes are predicted to encode polypeptides of about 110 kDa, each with a
large, cytoplasmic N-terminal region containing zinc-finger (ZnF) domains, and eight membrane
spans sandwiching the four U-motifs of the catalytic domain (Fig. 1B; Delmer, 1999).
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
4
Further evidence for the function of plant CesA genes in cellulose synthesis came from
Arabidopsis mutants of three of the CesA genes involved in primary wall synthesis: a
temperature-sensitive radial swelling mutant rsw1 (AtCesA1; Arioli et al., 1998), a dwarf-
hypocotyl mutant procuste (prc1) (AtCesA6; Fagard et al., 2000), and a stunted root phenotype
with altered jasmonate and ethylene signaling (cev1) and ectopic lignification (eli1) mutant
alleles (AtCesA3; Ellis and Turner, 2002; Caño-Delgado et al., 2003). Despite co-expression in
the same cells and an expectation of redundancy, cellulose synthesis is impaired in each mutant.
The same was observed with the irregular xylem mutants, irx1, irx3, and irx5, which display a
phenotype of collapsed mature xylem cells as a result of lowered cellulose content during
secondary cell wall deposition (Taylor et al., 2001, 2003). A widely accepted hypothesis is that
the AtCesA1, AtCesA3, and AtCesA6 proteins assemble to function in primary wall cellulose
synthesis, while the AtCesA4, AtCesA7 and AtCesA8 proteins assemble to make secondary wall
cellulose (Fig. 1A), with each member of the trio performing a non-redundant function in the
complex (Taylor, 2008). Lack of one CesA prevents incorporation of the other two into the
plasma membrane (Gardiner et al., 2003). However, at least some of the subunits are potentially
interchangeable, as inferred by the dominant negative inhibition of growth and primary wall
thickness caused by constitutive expression of a mutated fra5 (irx3 allele) transgene (Zhong et
al., 2003), and by the semi dominant-negative phenotype observed in the heterozygous AtCesA3
mutant (Daras et al., 2009). AtCesA1 is essential for cellulose synthesis (Beeckman et al., 2002),
whereas knock-outs of AtCesA3 (Ellis and Turner, 2002; Caño-Delgado et al., 2003) and AtCes6
(Fagard et al., 2000) result in partially impaired synthesis but not total inhibition. Desprez et al.
(2007) indicate that the AtCesA2 and AtCesA5 proteins have partially redundant functions with
AtCesA6.
A direct association of three distinct CesA polypeptides was demonstrated in vitro and by co-
localization in vivo by Taylor et al. (2003). Domain swap experiments with wild-type and mutant
AtCesA1 and AtCesA3 proteins in their respective mutants resulted in dominant-positive and
dominant-negative effects, consistent with both catalytic and C-terminus domains being
important for function (Wang et al., 2006). Direct interactions of three distinct CesA
polypeptides in vivo were shown by bimolecular fluorescence complementation (Desprez et al.,
2007). Although some complementary pairs gave stronger fluorescence than others, both homo-
and heterodimers of AtCesA1, AtCesA3 and AtCesA6 are inferred. Wang et al. (2008) used pull-
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
5
down experiments similar to those of Taylor et al. (2003) to show that these three primary wall
CesA proteins interact. Further, they showed that Triton-soluble microsomal preparations
subjected to native PAGE gave an 840 kDa complex, and that null mutants, but not missense
mutations, gave smaller 420 kDa complexes (Wang et al., 2008). Atanassov et al. (2009)
affinity-trapped a ladder of complexes of CesA oligomers to about 700-730 kDa. Consistent with
the observations of Wang et al. (2008), only smaller oligomeric complexes of two of the CesAs
are detected when the third is missing (Atanassov et al., 2009). Such an association of CesAs
was indicated independently in yeast two-hybrid studies (Timmers et al., 2009).
Does synthesis of each (1→4)-β-D-glucan chain require one or two catalytic polypeptides?
After over four decades of study, the biochemical mechanism by which cellulose is made
remains a mystery (Delmer, 1999; Saxena and Brown, 2005; Somerville, 2006; Guerreiro et al.,
2010), with only a few reports of cellulose synthesis in vitro with isolated membranes (e.g.
Kudlicka and Brown, 1997; Lai-Kee-Him et al., 2002). For both cellulose and the related (1→4)-
β-D-glycan, chitin, synthesis proceeds by attachment of glucosyl residues to the non-reducing
terminus of the acceptor glucan chain (Koyama et al., 1997; Imai et al., 2003). The simplest
hypothesis is that each CesA polypeptide synthesizes a single glucan chain. In the Delmer
(1999) model, the eight membrane spans form a channel through which a single glucan chain is
extruded (Fig. 3A). This mode of synthesis comes with a very big steric problem for synthesis.
To make a (1→4)-β-D- linkage means that each glucosyl residue is turned 180° with respect to
each neighbor. Thus, the O-4 position of non-reducing terminal sugar of the acceptor chain is
displaced several Ångstroms upon addition of each successive unit (Fig. 3B). For the next
glycosyl transfer to occur, then the site of catalysis must move several Ångstroms within the
protein, the acceptor chain must swivel 180°, or the catalytic or acid-base amino acids must
toggle between two forms to account for the displacement. To overcome this steric problem,
several models have proposed that two sites or modes of glycosyl transfer reside within the
catalytic complex, so that disaccharide units are added iteratively (Carpita et al., 1996; Koyama
et al., 1997; Carpita and Vergara, 1998; Buckeridge et al., 1999, 2001; Saxena et al., 2001), or
that two polypeptides associate to form two opposing catalytic sites (Vergara and Carpita, 2001;
Buckeridge et al., 2001). In either model, glycobiosyl units, or any even-numbered oligomeric
units, are added to the non-reducing end to ensure that the (1→4)-β-D- linkages are strictly
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
6
preserved without inversion of substrate, active site, or terminus of the growing chain (Fig. 3C).
Despite the rationale for a two-site model of catalysis, biochemical evidence from other
types polysaccharide synthases indicate that a single polypeptide is sufficient. Hyaluronan (HA)
is an unbranched polysaccharide composed of repeating units of (1→3)-β-D-N-
acetylglucosamine and (1→4)-β-D-glucuronic acid (DeAngelis and Weigel, 1994). HA synthases
exist in three distinct classes, with Class I containing integral membrane proteins to transport the
HA across the membrane (Fig. 4). Bacterial and mammalian HA synthases have been shown to
contain both transferase activities in a single polypeptide (DeAngelis and Weigel, 1994; Yoshida
et al., 2000; Williams et al., 2006). Such a finding argues that synthesis of a single HA polymer
requires only a single polypeptide.
The crystal structure of a non-processive family 2 glycosyltransferase (GTs) sharing
sequence similarity with a portion of the catalytic domain of a CesA, the Bacillus subtilis SpsA
synthase, provided the first conformation of the active site and the role of the aspartyl residues in
the positioning of the uridinyl group of a UDP-sugar (Fig. 5A; Charnock and Davies, 1999;
http://www.pdb.org/pdb/explore/explore.do?structureId=1QGS). Charnock and colleagues
(2001) argued that only a single site for a nucleotide-sugar substrate is accommodated within a
single polypeptide of SpsA.
A catalytic dimer hypothesis
From the studies of the Class I HA synthases and the SpsA crystal structure, it is inferred that a
single polypeptide alone has all the features needed for synthesis. However, these features still
do not address mechanistically the fundamental steric problem of synthesis of a repeating
(1→4)-β-D-glucosyl linkage of the cellulose glucan chains. In fact, two recent studies
demonstrate unequivocally that at least some HA synthases and homologs of SpsA synthase
form dimers. An identical structure as the SpsA synthase is the 3BCV polypeptide from
Bacteroides fragilis, which is also predicted to contain a single substrate-binding site. In contrast
to SpsA synthase, the 3BCV protein crystallizes as a dimer, and each monomer possesses a
bound UDP (Fig. 5B; (http://www.pdb.org/pdb/explore/explore.do?structureId=3BCV). The
dimerization occurs through the C-terminal regions, which appear to be flexible and, for this
reason, were deleted from the crystal structure of the SpsA synthase (Charnock and Davies,
1999). This is the first direct evidence by crystal structure of a homodimer formed by GT2
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
7
proteins, but even more complicated structures are also found. For example, an E. coli strain K4
chondroitin polymerase contains two ‘Rossmann fold-like’ domains within a single polypeptide,
each binding a UDP-GlcA or UDP, and it also crystallizes as a dimer giving a total four
nucleotide binding domains (Fig. 5C;
(http://www.pdb.org/pdb/explore/explore.do?structureId=2Z86). Although the fit is not
exceptionally good, the CesA catalytic domain threads through the E. coli chondroitin
polymerase best of all for known crystal structures of glycosyl transferases involving nucleotide
sugars (D. Kihara, Purdue University, personal communication), and it is consistent with the
suggestion by Brown and Saxena (2000) and Saxena et al. (2001) of a conformation within
which the catalytic domain of a single CesA would allow synthesis of cellobiose units of the
chain within a single polypeptide.
The Class II synthases contain two different types of GT2 modules but not the membrane-
spanning domains (Fig. 4). One type of HA synthase possesses two repeats of the UDP-Glc and
acceptor binding domains, so the synthesis of the characteristic disaccharide of HA by a single
synthase is rationalized (Jing and DeAngelis, 2000). However, a direct interaction of two
synthases is inferred for HA synthesis to explain the finding that host cells harboring constructs
in which each site is independently disrupted are still able to make HA (Jing and DeAngelis,
2000; Weigel and DeAngelis, 2007). Because the Class I HA synthases have both activities on a
single peptide does not preclude the possibility that formation of homodimers of single isoforms
of HA synthase is necessary for function, which, like the chrondroitin polymerase, would give
four nucleotide sugar binding sites per dimer.
Solving the steric problem aside, one must also ask if a channel of 8 membrane spans
proposed for the cellulose synthase of sufficient size to extrude a (1→4)-β-D-glucan chain. The
question about sufficient channel size was raised also with respect to the GT family 2 HA
synthases by Weigel and DeAngelis (2007), who suggested that certain phospholipids required
for activity possibly integrate with the membrane spans to widen the channel for extrusion.
However, whether lipid interactions with a small number of domains would be significant is still
in question. Callose synthases are about twice the size of CesAs and contain 16 membrane spans,
i.e. double those of a CesA (Hong et al., 2001). Plasma membrane hexose and maltose
transporters of prokaryotes and eukaryotes are homologous (Maiden et al., 1987), and virtually
all of them contain a minimum of 11 to as many as 18 transmembrane spans per functional unit
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
8
(Pao et al., 1998; Reifenberger et al., 1995; Sherson et al., 2000; Klepek et al., 2009).
The sensitivity of detergent-solubilized CesAs complexes to DTT suggested to Atanassov et
al. (2009) that disulfide bonds are involved in the coupling into the larger complexes. Other
features of the protein outside the region of catalysis, such as the ZnFs, which show high
similarity to RING-finger domains that bind Zn in a ‘cross-brace’ manner (Freemont, 2000),
might function in the organization of these into the larger rosette structure. Kurek et al. (2002)
proposed that CesAs are coupled through the ZnF domain in a redox-dependent manner,
constituting the first step in the clustering of CesAs into rosettes. Moreover, the discovery that a
thioredoxin-like protein associates with the CesA ZnF domain in a yeast two-hybrid screen led to
the suggestion that reduction of the domains by oxidoreductases returns the CesAs to monomeric
forms, which are directed to the ubiquitin-dependent turnover pathway (Kurek et al., 2002). The
experimental herbicide CGA 325’615 blocks crystallization of the β-D-glucan chains into
cellulose microfibrils, phenocopying the rsw1 swollen root tip (Peng et al., 2001). This
phenotype can be abrogated completely in the presence of H2O2, suggesting the inhibitor blocks
rosette assembly by enzymatic oxidation (Kurek et al., 2002).
If ZnF domains of two CesAs couple as part of the recruitment into rosette particles, the
question remaining is how all the others interact to form a complete complex. Although not
discussed specifically, the study by Kurek et al. (2002) presented data that full-length CesA
proteins formed tetramers and even higher ordered pairings, while the ZnF domains were limited
to coupling of a single pair. Timmers et al. (2009) showed that heterodimer interactions indicated
by yeast two-hybrid analysis do not require the ZnF domains. Taken together, these data provide
evidence that domains other than the ZnFs of the CesA participate in coupling reactions if two to
three dozen CesAs or more are aggregated to form a rosette complex.
Comparison of CesA sequences suggests potential heterodimeric interaction domains
within the catalytic domain. The initial scarcity of CesA protein sequences and the apparent
variability within the so-called HVR led to the assumption that this region was probably not
essential in catalysis (Pear et al., 1996). However, it is now understood that these regions are
well conserved across grass and dicot species with distinct sub-clade structure. Potential protein-
protein interactions through sub-domains of this region containing conserved Cys residues,
clusters of consecutive basic Lys and Arg residues, and clusters of acidic Asp and Glu residues
forming the basis of a Class Specific Region (Fig. 1B; Vergara and Carpita, 2001).
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
9
To test the catalytic dimer hypothesis, we expressed fusion proteins containing only the
catalytic domain of Arabidopsis and maize (Zea mays) CesAs with affinity tags and observed
that dimers and higher-order aggregates collapse reversibly to monomeric forms by thiol
reducing agents (C. Rayon, A. Olek, L. Paul, S. Ghosh, unpublished results). Because the ZnF
was absent in these constructs, dimerization must occur through thiol-sensitive sequences in the
catalytic domain. Such an interaction of CesAs to form homo- or hetero-dimers solves the three
basic problems of the single polypeptide-single polymer conundrum: 1) the steric problem is
solved by coordinate synthesis and attachment of cellobiose units instead of monomers,
preserving the integrity of the O-4 site of attachment at the non-reducing terminus of the chain,
2) a channel of 16 membrane spanning domains is consistent with sugar transport and callose
extrusion, and 3) the interaction produces two ZnF domains for recruitment of the catalytic dimer
into rosette particles (Fig. 6). An exciting prospect is that conservation of space would be
maintained if CesAs turn out to have a structure like the chondroitin polymerase dimers and
function like Class II HA synthases, because a CesA catalytic dimer with four nucleotide binding
domains would be capable of generating two (1→4)-β-D-glucan chains instead of just one.
Further experiments are needed to establish preferred heterodimer interactions, the stoichiometry
of UDP-Glc binding, and the role of the ZnF in recruitment of the catalytic dimers into larger
complexes.
The biological synthesis of cellulose
Cellulose synthase has a half-life of less than thirty minutes, remarkably short for a
membrane protein (Jacob-Wilk et al., 2006). Assembly of rosettes occurs in the Golgi stacks, and
they must be continually secreted to the plasma membrane to maintain cellulose synthesis
(Haigler and Brown, 1986). Additional proteins are suspected to be necessary for the formation
of primers of polymer synthesis, metabolic channeling of substrates, chain crystallization of the
chains, and termination of chains (Doblin et al., 2002; Somerville, 2006; Guerreiro et al., 2010).
Further, a proteomics survey of plasma membrane proteins shows that certain CesAs are
phosphorylated at several locations within the catalytic and N-terminal domains (Nühse et al.,
2004). Modification of some potential phosphorylation sites with amino acids that either prevent
(Ala) or mimick (Glu) phoshorylation have multiple effects that either reduce synthesis rates or
interactions with the microtubule cytoskeleton independently (Chen et al., 2010).
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
10
In affinity labeling experiments with [32P]UDP-Glc, an 84 kD polypeptide was found to be
associated with a plasma membrane fraction containing the highest activity of callose synthase
(Delmer et al., 1991), and subsequently was identified as sucrose synthase (SuSy). Confirmation
of plasma membrane association was made immunocytochemically (Amor et al., 1995), and
Delmer and Amor (1995) proposed that the association of SuSy represented a UDP-Glc delivery
mechanism to cellulose synthase. β-Glucan microfibrils are synthesized from sucrose and UDP
on immobilized tobacco plasma membrane sheets (Hirai et al., 1998). More recently, SuSy was
immunologically associated with CesA proteins in the rosette structures (Fujii et al., 2010),
strengthening the idea of an association of SuSy directly with cellulose synthases for metabolic
channeling of glucose through a localized pool of UDP-Glc. However, a quadruple mutant that
eliminates all detectable SuSy in vegetative tissue does not impair cellulose synthesis (Barratt et
al., 2009). Overexpression of SuSy in developing vascular tissue of transgenic poplar yields
small but significant increases in cellulose content (Coleman et al., 2009). Taken together, SuSy
does not appear to be required for cellulose synthesis but may enhance rates by concentrating
substrate at the site of synthesis.
Peng et al. (2002) provided evidence that sitosterol-cellodextrins synthesized from
sitosterol-β-glycoside serve as primers of glucan chain initiation, with the KORRIGAN
glucanohydrolase trimming the sitosterol from the growing chain. Debolt and colleagues (2009)
question this role, as they find double mutants of two major sterol-β-glucoside synthases result in
severe defects in cuticle formation but not in cellulose synthesis. However, the sterol-glucosides
are substantially reduced in the double mutant but not entirely eliminated, leaving open the
question.
Outside the Golgi stacks, a membrane compartment also containing the KORRIGAN
(Robert et al., 2005) might represent a dynamic factory associated with both the microtubule
network and plasma membrane that aligns and directs the cellulose synthase complex, and
coordinate that function with the deposition of the many polysaccharides directed to it from
packaged Golgi vesicles. Bimolecular fluorescence complementation techniques, as shown for
CesA interactions (Desprez et al. 2007) can give important clues to selected participants in
protein-protein interactions within a complex in vivo. Fluorescence tagging has also allowed
visualization of the movement of cellulose synthase complexes at the plasma membrane (Paredez
et al., 2006; Wightman and Turner, 2006; Gutierrez et al., 2009). These studies also established
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
11
the dynamics of the relationships with the cortical microtubule network in real time. As reviewed
by Baskin (2001) and Szymanski and Cosgrove (2009), these studies bring resolution to ideas on
alignment of cortical microtubules and cellulose microfibrils generated long ago through
observations with inhibitors (Green, 1962) and in the electron microscope (Ledbetter and Porter,
1963). Improvements in imaging tools are still needed that permit visualization of the delivery
of Golgi-derived vesicles to the sites of cellulose synthesis much progress has already been made
along these lines (Held et al., 2008; Konopka and Bednarek, 2008; Crowell et al., 2009).
The synthesis of non-cellulosic polysaccharides with (1→4)-β-D-glycan backbones
Because of the same conserved domains of nucleotide-sugar binding and catalysis as those
encoding CesAs, sub-families of Csl genes were predicted to encode the synthases of non-
cellulosic polymers with (1→4)-β-D-glycan backbones, primarily (gluco)mannans and
galacto(gluco)mannans, xyloglucans, glucuronoarabinoxylans (GAX), and the grass-specific
(1→3),(1→4)-β-D-glucans (Delmer, 1999; Richmond and Somerville, 2000; Hazen et al., 2002).
For the most part this has turned out to be true, but GAXs are a clear exception. There is still an
incomplete knowledge of most of the gene products and interactions among them to make
specific β-D-glycans, even among families where at least one member has a confirmed glycosyl
transferase activity. There is also a growing disconnection between the classic studies on the
synthesis of these polysaccharides in vitro and the discovery of genes encoding the machinery
that warrants a revisit.
CslF and CslH: Mixed-linkage (1→3),(1→4)-β-D-glucan synthase is the topological
equivalent of cellulose synthase
The mixed-linkage (1→3),(1→4)-β-D-glucan is made in the grasses (Poales)(Carpita, 1996;
Buckeridge et al., 2004), certain lichens (Wood et al., 1994), and Equisetum (Fry et al., 2008;
Sørensen et al., 2008), but differences in the distribution of their cellodextrin oligomers indicate
they probably arose by convergent evolution of synthases. For the grasses, this glucan is not a
random mixture of (1→3)-β-D- and (1→4)-β-D-glucosyl linkages, but is composed primarily of
cellotriosyl and cellotetraosyl units in a ratio of about 2.5:1 connected by single (1→3)-β-D-
linkages (Wood et al., 1994). Upon cleavage with a Trichoderma cellulase, smaller amounts of
higher cellodextrin series are observed with the odd-numbered cellodextrin 2-fold higher in
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
12
abundance than the next even-numbered unit in the series. The synthesis of the (1→3),(1→4)-β-
D-glucan has been demonstrated in vitro with isolated, intact Golgi membranes and UDP-Glc
(Gibeaut and Carpita, 1993; Buckeridge et al., 1999, 2001; Urbanowicz et al., 2004). Whereas
μM concentrations of UDP-[14C]-glucose result in much shorter oligomers and polymers
enriched in cellotetraosyl units rather than cellotriosyl units (Buckeridge et al. 1999), much
larger polysaccharides enriched in cellotriosyl units are observed with higher concentrations of
substrate (Buckeridge et al., 1999, 2001). The ratios of cellotriosyl : cellotetraosyl and
cellopentosyl : cellohexaosyl are increased proportionally with substrate concentrations higher
than 250 μM (Buckeridge et al., 1999), indicating that the mechanism of synthesis of the odd-
numbered cellodextrin unit is fundamentally different than synthesis of the even-numbered units.
Proteolysis protection assays show further that the active site of catalysis is on the outward
facing Golgi membrane (Urbanowicz et al., 2004). Golgi membranes treated with proteinase K
specifically lost their ability to make the odd-numbered cellodextrin units, whereas the synthesis
of the cellotetraosyl and higher order even-numbered units was unaffected. Again, loss of the
ability to make cellotriosyl units is correlated with significant loss in size of the (1→3),(1→4)-β-
D-glucan product (Urbanowicz et al., 2004). We proposed a similar catalytic dimer model
wherein even-numbered units are synthesized by core cellulose synthase-like proteins and the
odd-numbered units arise by an additional glycosyl transferase (GT) that has yet to be identified
(Buckeridge et al., 2001, 2004).
Limited proteolysis and detergent reconstitution experiments with the mixed-linkage
(1→3),(1→4)-β-D-glucan of grass species provides kinetic evidence for three sites of glycosyl
transfer within the catalytic domain—two from the cellulose synthase-like core domain and a
third, separable activity (Urbanowicz et al., 2004). (1→3),(1→4)-β-D-Glucan is the topological
equivalent of cellulose synthase at the Golgi membrane. Limited proteolysis or detergent
treatment causes loss of the ability to make the diagnostic odd-numbered cellotriose units for
synthesis, without affecting the ability to generate the even-numbered cellotetraosyl unit
(Urbanowicz et al., 2004).
As the topologic equivalent of cellulose synthase at the Golgi membrane, the
(1→3),(1→4)-β-D-glucan synthase shares another feature with cellulose synthase. When its
resident membranes are damaged, cellulose synthase (Delmer, 1977), and the (1→3),(1→4)-β-D-
glucan synthase (Buckeridge et al., 2001) ‘default’ to synthesis of the (1→3)-β-D-glucan,
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
13
callose, possibly by disruption of the complete active site to a single glycosyl transferase activity
(Buckeridge et al., 2001; Urbanowicz et al., 2004). Loss of the cellobiosyl generating system to a
single site of glycosyl transfer would thereafter make only callose (Buckeridge et al., 2001),
whose synthesis does not require turning the catalytic site or acceptor 180° (Fig. 3B). While it
can be argued that membrane disruption activates callose synthase in vitro in plasma membrane
preparations of all angiosperms (Nishamura et al., 2003), only Golgi membranes from grasses–
the only angiosperms that make the (1→3),(1→4)-β-D-glucan, make callose when damaged
(Gibeaut and Carpita, 1993; Buckeridge et al., 1999). The most direct evidence for the default
synthesis of callose from a damaged cellulose synthase comes from the experiments of Blanton
et al. (2000), who showed that isolated membranes of a cellulose synthase mutant of the cellular
slime mold Dictyostelium discoideum also lost the ability to make callose in vitro. Whereas
some cellulose is made in vitro with wild-type membrane preparations, callose linkages
predominate. Because cellulose synthase is a single gene in Dictyostelium, loss of the ability to
make (1→3)-β-D-glucan in vitro as well as (1→4)-β-D-glucan in membranes from the mutant
strongly suggests that the single polypeptide is responsible for both activities.
Two groups of Csl genes, CslF and CslH, which are found only in grasses (Hazen et al.,
2002), have been shown to catalyze (1→3),(1→4)-β-D-glucan biosynthesis. Heterologous
expression of a rice CslF in Arabidopsis, a species that does not make (1→3),(1→4)-β-D-glucan,
results in small amounts of the β-D-glucan in the cell walls (Burton et al., 2006). However,
considerably greater amounts of the (1→3),(1→4)-β-D-glucan result when a CslH is co-
expressed with CslF, suggesting a synergistic role for both CslH and CslF in the synthesis of the
polysaccharide, and that a catalytic heterodimer enhances the activity (Doblin et al., 2009). If an
accessory glycosyl transferase is necessary to make the odd-numbered cellodextrin unit, then
Arabidopsis must produce a related isoform. This finding of concerted action by two distinct
group members highlights the possibility that synthases of other cross-linking glycans might be
encoded by Csls of different groups.
CslA: Mannan and Glucomannan Biosynthesis
One of the first cell wall polysaccharides to be synthesized in vitro was glucomannan. An early
conclusion that GDP-Glc is the substrate for (1→4)-β-D- linkages of cellulose, and UDP-Glc is
the substrate for (1→3)-β-D-glucans (Chambers and Elbein, 1970), had already been disproven.
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
14
Still today GDP-Glc is still listed erroneously as the substrate for cellulose synthesis on most
wall charts of biochemical pathways, even though kinetic evidence obtained with cotton fiber
cells cultured in vitro showed unequivocally that UDP-Glc is the substrate for cellulose synthesis
(Carpita and Delmer, 1981). Addition of both GDP-Glc and GDP-Man to membrane
preparations resulted in marked stimulation of incorporation into a glucomannan product (Elbein
and Hassid, 1966; Piro et al., 1993).
The knowledge of GDP-nucleotide sugars as substrates were instrumental in the discovery
that a CslA gene encodes a mannan synthase by expression profiling of guar seed development, a
species that accumulates large amounts of galactomannan as a cell wall storage carbohydrate
(Dhugga et al., 2004). Liepman et al. (2005, 2007) confirmed that at least four members of the
CslA group function in mannan and/or glucomannan synthesis. However, mixed substrates of
GDP-Glc and GDP-Man in the heterologous expression system (Liepman et al., 2007) do not
give the marked enhancement of glucomannan synthesis long ago observed in vitro (Elbein and
Hassid, 1966). While the CesA genes appear orthologous across several species, the Csl genes
are not (Penning et al., 2009). In fact, the CslA group is resolved into three sub-groups that
either are Arabidopsis-dominated, grass-dominated, or mixed. The CslA members defined as
mannan synthase genes (Liepman et al., 2007), fall into both the Arabidopsis-dominated and the
mixed sub-group (Penning et al., 2009). The functions of these other sub-group members of the
grasses need to be defined.
CslC: Xyloglucan Biosynthesis
Xyloglucans were among the first complex cell wall polysaccharides whose synthesis was
demonstrated in vitro. Early studies showed that labeled sugars from UDP-Glc and UDP-Xyl are
incorporated into several polysaccharides using microsomal membranes, and were later refined
by isolation of Golgi membranes (Ray et al., 1969, 1980). Small amounts of xyloglucan-like
oligomers with the characteristic α-D-Xyl-(1→6)-D-glucosyl unit, isoprimeverose, are made with
small amounts of UDP-Glc and UDP-Xyl, but Gordon and Maclachlan (1989) found that when
concentrations of each nucleotide-sugar are increased to millimolar, large polymers containing
the characteristic heptasaccharide XXXG (for nomenclature, see Table I) unit structure are
synthesized. The tetraglucosyl unit of the xyloglucan backbone and the precisely repeated three
xylosyl units added to make the XXXG structure is consistent with an even-number cellobiose
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
15
unit synthase reaction for the glucan backbone. Even in the structural variant of Solanaceous
xyloglucan, where just two xylosyl units are added, a tetraglucosyl unit backbone is preserved by
the replacement of the third xylosyl group with an acetate (Sims et al., 1996). An apparent
exception is the ability of certain tree legumes, such as jatobá (Hymenaea courbaril), to make
XXXXG units in addition to XXXG (Buckeridge et al., 2000). Curiously, partial digestion of this
polymer with a Trichoderma cellulase, which cleaves only at unbranched positions, yields
octomer, nonamer, and decamer backbone oligomers whose ratios predict that the polymer
consists of 4-5-5-4 frameworks separated by variable amounts of XXXG, rather than a random
distribution of XXXXG and XXXG units (Tiné et al., 2006). This is an intriguing result for two
reasons: 1) the 4-5-5-4 framework of these types of xyloglucans preserves the even-numbered
unit symmetry of the backbone, and 2) to make such a framework, as many as 18 glucosyl
residues might be contained within the complex in order to be ‘read’ properly to preserve the unit
structure, making the complex much larger than expected.
The topology of xyloglucan synthesis at the Golgi membrane is still uncertain. Several
lines of evidence suggest that synthesis relies on transporters for UDP-Glc for backbone
synthesis of (1→4)-β-D-glucans in vitro (Orellana, 2005), and UDP-GlcA is transported to
provide UDP-Xyl for transfer within the lumen (Hayashi et al., 1988). Lerouxel et al. (2006)
propose that synthesis of the backbone is the topological equivalent of cellulose synthase, but the
addition of all subtending sugars of Xyl, Gal, and Fuc are within the lumen of the Golgi.
Based on heterologous expression in Pichia, Cocuron et al. (2006) provide evidence that
CslC genes encode the synthases of the xyloglucan backbone. Although Pichia is unable to
make UDP-Xyl, co-expression of the xylosyl transferase with CslC is sufficient to induce
extension of (1→4)-β-D-glucan chains, indicating that a close interaction of these proteins might
stabilize the synthase to allow extension of the backbone. A complication to unequivocal
annotation of function of this sub-family is the finding of a CslC at the plasma membrane instead
of the Golgi membrane (Dwivany et al., 2009). Three members of the GT34 family are
established as the xylosyl transferases involved in xyloglucan synthesis, and these Golgi resident
proteins are predicted to face the lumen. (Cavalier et al. 2008; Zabotina et al., 2009)
Xyloglucan is decorated in various ways in a species-specific manner (Hoffman et al.,
2005; Peña et al., 2008). Most are primarily galactosylated, with a characteristic α-L-Fuc-
(1→2)-β-D-Gal-(1→2)-α-D-Xyl- trisaccharide extension (Bauer et al., 1973), but others, such as
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
16
those of Solanaceous species, have a truncated unit structure substituted with α-L-Ara-(1→2)-α-
D-Xyl extensions instead of Gal (Sims et al., 1996), and the Asteridae and Oleales species have
mixtures of these two forms of xyloglucan substitution (Hoffman et al., 2005). Further, the
xyloglucans synthesized in the Golgi are modified in ways that make them structurally different
than those that are assembled onto the cellulose microfibrils in the wall (Obel et al., 2009).
The purification of a xyloglucan-specific fucosyl transferase that led to the discovery of a
glycosyl transferase family 37 (GT37) gene encoding it (Perrin et al., 1999). While the synthase
complex must involve a close interaction between the glucan synthases and the xylosyl
transferases that decorate it (Cavalier et al., 2008), the association of the galactosyl and fucosyl
transferases might be more transient. Transferases extracted from the membrane are able to add
Gal from UDP-Gal (Madson et al., 2003) and Fuc from GDP-Fuc (Perrin et al., 1999; Vanzin et
al., 2002) to exogenous xyloglucan in vitro.
Xylan Biosynthesis
The synthesis of grass (1→4)-β-D-xylans from UDP-Xyl with microsomal membranes was
first demonstrated by Bailey and Hassid (1966). Cooperative action of two nucleotide-sugar
substrates, in this instance UDP-Xyl and UDP-GlcA, resulted in the synthesis of (1→4)-β-D-
xylans with subtending GlcA units (Waldron and Brett, 1983; Baydoun et al., 1989). Similar
studies have shown that membrane preparations from grasses and mixtures of nucleotide sugars
made GAXs, in part employing a nascent C-4 epimerase that interconverts UDP-Xyl and UDP-
Ara (Porchia and Scheller, 2000; Porchia et al., 2002; Kuroyama and Tsumuraya, 2001; Zeng et
al., 2008).
Apart from a hint that the AtCslD5 may be involved (Bernal et al., 2007) in the synthesis of
(1→4)-β-D-xylans, it has yet to be directly demonstrated that any Csl gene plays a role. In fact,
informatics approaches yield non-Csl genes as more likely candidates for encoding the
machinery for xylan synthesis (Mitchell et al., 2007), and several mutants with deficiencies in
normal xylan synthesis, such as parvus (Lao et al., 2003; Lee et al., 2007), irx8 (Brown et al.,
2005), irx7/fra8 (Zhong et al., 2005; Brown et al., 2007), irx9 and irx14 (Brown et al., 2007), and
irx10 and irx10-L (Brown et al., 2009), do not include a member of the Csl gene family.
(1→4)-β-D-Glucuronoxylan (GX) synthesis appears to involve a complex initiation
sequence. Peña et al. (2007) discovered that collapsed xylem mutants deficient in xylan, irx8
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
17
and fra8, were essentially devoid of a complex tetrasaccharide, β-D-Xyl-(1→3)-α-L-Rha-(1→2)-
α-D-GalA-(1→4)-D-Xyl, located at the reducing end of the xylan polymer, whereas an irx9
mutant also severely deficient in xylan contained an overabundance of the tetrasaccharide. The
presence or absence of this tetrasaccharide greatly affected the size distribution of the xylans. In
irx8 and fra8 a broader distribution is observed, with some polymers longer than observed in
wild type. Xylans of irx9 have short chains, with nearly all of them containing the
tetrasaccharide (Peña et al., 2007). These results suggested a model whereby short chains of
(1→4)-β-D-xylan are primed by the tetrasaccharide, and these are spliced, cleaving the primer,
to make the long polysaccharides (York and O’Neill, 2008).
The IRX8 (GAUT12) and PARVUS (GATL1) encode members of the GT8 of Group C, and
IRX7/FRA8 of the GT47 Group E, that synthesize the primer tetrasaccharide, whereas IRX9 and
IRX14 encode members of GT43 that are likely to encode the synthases of the (1→4)-β-D-xylan
oligomeric backbones that are stitched together by a yet unidentified glycosyl transferase (Brown
et al., 2007; Peña et al., 2007). GT8 family members encode retaining-type transferases and the
GT47 members encode inverting-type transferases with respect to the anomeric linkages formed
compared to the anomeric linkage in the nucleotide sugar, so GT8 members make α-D-linkages,
whereas the GT47 members make β-D- or α-L- linkages. While an α-D-GalA is found in the
tetrasaccharide, (1→2)-α-D-GlcA (4-O-Me-GlcA) side groups are also attached at precise
intervals along the xylan chain (Nishitani and Nevins, 1991), and a general blockwise synthesis
of six consecutive branched xylosyl residues in grass xylans is observed (Carpita and Whittern,
1986). IRX10 and IRX10-L appear to encode xylosyl transferases also from GT47, but double
mutants of these genes have greatly reduced GlcA substitutions along the shorter chains (Brown
et al., 2009). It will be interesting to determine if addition of these GlcA side-groups play a role
as an attachment or recognition point where short xylan chains are grafted together to make a
long chain–a suggestion foretold by the original work on the cooperative action of UDP-GlcA
and UDP-Xyl in the in vitro synthesis of glucuronoxylan (Waldron and Brett, 1983; Baydoun et
al., 1989).
York and O’Neill (2008) take a broader perspective of xylan synthesis and have suggested
that reducing end addition should not been ruled out. In their model a type of alternating
‘pendulum’ mechanism is proposed for the introduction of the (1→4)-β-D-xylan linkages, a
mechanism analogous to the dimer synthesis described here for (1→4)-β-D- linkages in general.
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
18
We are just learning the identity of the GlcA and Araf transferases of the GX and GAX polymers
and the distinctions between the GX devoid of arabinose that is abundant in the secondary xylem
and the GAX with its rich Ara substitution that is the major polymer of the primary cell walls of
grasses (Scheller and Ulvskov, 2010). Recently, proteomic approaches of isolated GAX synthase
complexes from wheat membranes demonstrated a close interaction of GT43 and GT47 family
members with a GT75 UDP-Ara mutase, an enzyme that interconverts the UDP-arabinopyranose
and UDP-arabinofuranose conformations required for incorporation of the latter into the
polysaccharide (Zeng et al., 2010). These kinds of studies represent the path forward to identify
all the components of synthase complexes whose activities are preserved in vitro.
Concluding remarks
While the plant cell wall community is making steady progress on defining biochemical
functions of backbone synthases and the glycosyl transferases that decorate them, the
biochemical details of these large, coordinated complexes are still unknown. Fading from
memory are a great many foundation studies of polysaccharide synthesis in vitro that give
insights to the differences between an isolated activity and the function of a complex as a whole.
The major differences between the behaviors of protein complexes in vitro and in vivo are
compounded by the topology of the synthases and associated glycosyl transferases at the plasma
membrane and Golgi membranes, across which both pH gradients and electrical potentials are
generated. Early studies with excised cotton fibers cultured in vitro showed that resealing of the
plasma membrane was essential to reconstitute cellulose synthesis, and evidence provided that
regeneration of the electrical potential rather than the pH gradient led to that was critical to
preservation of synthesis (Carpita and Delmer, 1980). This work was extended to show that
artificial potentials stimulate (1→3)-β-D-glucan synthesis in vitro (Bacic and Delmer, 1981). In
Gluconacetobacter, rates of cellulose synthesis could be modulated directly through adjustment
of the electrical potential in living cells under conditions that did not impair metabolism (Delmer
et al., 1982). Such gradients exist across other compartments, and, in contrast to cellulose
synthesis at the plasma membrane, it is maintenance of a pH gradient that prolongs the synthesis
of the mixed-linkage (1→3),(1→4)-β-D-glucan in vitro in isolated maize Golgi membranes
(Gibeaut and Carpita, 1993). The physiological and biochemical bases for the effect of these
potentials and gradients on polysaccharide synthesis are still not understood. They do serve to
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
19
illustrate that, beyond the biochemical mechanism of synthesis, technologies that preserve not
only the protein complexes but also their cellular context need to be developed to truly
understand synthesis of macromolecules across membrane surfaces.
Acknowledgments
I thank Maureen McCann (Purdue University) and Peter Ulvskov (University of Copenhagen)
for their review of the manuscript and their many helpful suggestions. I also thank Daisuke
Kihara (Purdue University) for his contributions to the discussion on protein structure and
modeling. The artwork in Figure 6 is by Pamela Burroff-Murr (Purdue University).
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
20
References
Amor Y, Haigler CH, Johnson S, Wainscott M, Delmer DP (1995) A membrane-associated
form of sucrose synthase and its potential role in synthesis of cellulose and callose in
plants. Proc Natl Acad Sci USA 92: 9353-9357
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant
Arabidopsis thaliana. Nature 408: 796-815
Arioli T, Peng LC, Betzner AS, Burn J, Wittke W, Herth W, Camilleri C, Höfte H,
Plazinski J, Birch R, Cork A, Glover J, Redmond J, Williamson RE (1998) Molecular
analysis of cellulose biosynthesis in Arabidopsis. Science 279: 717-720
Atanassov II, Pittman JK, Turner SR (2009) Elucidating the mechanisms of assembly and
subunit interaction of the cellulose synthase complex of Arabidopsis secondary cell walls. J
Biol Chem 284: 3833-3841
Bailey RW, Hassid WZ (1966) Xylan synthesis from uridine-diphosphate-D-xylose by
particulate preparations from immature corncobs. Biochemistry 56: 1586–1593
Bacic A, Delmer DP (1981) Stimulation of membrane-associated polysaccharide synthetases by
a membrane potential in developing cotton fibers. Planta 152: 346-351
Barratt DHP. Derbyshire P, Findlay K, Pike M, Wellner N, Lunn J, Feil R, Simpson C,
Maule AJ, Smith AM (2009) Normal growth of Arabidopsis requires cytosolic invertase
but not sucrose synthase. Proc Natl Acad Sci USA 106: 13124-13129
Baskin TI (2001) On the alignment of cellulose microfibrils by cortical microtubules: a review
and a model. Protoplasma 215: 150-171
Bauer WD, Talmadge KW, Keegstra K, Albersheim P (1973) The structure of plant cell
walls. II. The hemicellulose of the walls of suspension-cultured sycamore cells. Plant
Physiol 51: 174-187
Baydoun EA-H, Waldron KW, Brett CT (1989) The interaction of xylosyltransferase and
glucuronyltransferase involved in glucuronoxylan synthesis in pea (Pisum sativum)
epicotyls. Biochem J 257: 853-858
Beeckman T, Przemeck G K, Stamatiou G, Lau R, Terryn N, De Rycke R, Inze D, Berleth
T (2002) Genetic complexity of cellulose synthase A gene function in Arabidopsis
embryogenesis. Plant Physiol 130: 1883-1893
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
21
Bernal AJ, Jensen JK, Harholt J, Sørensen S, Moller I, Blaukopf C, Johansen B, de Lotto
R, Pauly M, Scheller HV, Willats WG (2007) Disruption of ATCSLD5 results in reduced
growth, reduced xylan and homogalacturonan synthase activity and altered xylan
occurrence in Arabidopsis. Plant J 52: 791-802
Blanton RL, Fuller D, Iranfar N, Grimson MJ, Loomis WF (2000) The cellulose synthase
gene of Dictyostelium. Proc Natl Acad Sci USA 97: 2391-2396
Bowling AJ, Brown RM (2008) Cytoplasmic domain of cellulose synthesizing complexes in
plants. Protoplasma 233: 115-127
Brown DM, Zeef LAH, Ellis J, Goodacre R, Turner SR (2005) Identification of novel genes
in Arabidopsis involved in secondary cell wall formation using expression profiling and
reverse genetics. Plant Cell 17: 2281-2295
Brown DM, Goubet F, Vicky WWA, Goodacre R, Stephens E, Dupree P, Turner SR (2007)
Comparison of five xylan synthesis mutants reveals new insight into the mechanisms of
xylan synthesis. Plant J 52: 1154-1168
Brown DM, Zhang Z, Stephens E, Dupree P, Turner SR (2009) Characterization of IRX10
and IRX10-like reveals an essential role in glucuronoxylan biosynthesis in Arabidopsis.
Plant J 57: 732-746
Brown RM Jr, Saxena IM (2000) Cellulose biosynthesis: a model for understanding the
assembly of biopolymers. Plant Physiol Biochem 38: 57-67
Buckeridge MS, Vergara CE, Carpita NC (1999) Mechanism of synthesis of a cereal mixed-
linkage (1→3)(1→4)-β-D-glucan: evidence for multiple sites of glucosyl transfer in the
synthase complex. Plant Physiol 120: 1105-1116
Buckeridge MS, dos Santos HP, Tiné MAS (2000) Mobilisation of storage cell wall
polysaccharides in seeds. Plant Physiol Biochem 38: 141-152
Buckeridge MS, Vergara CE, Carpita NC (2001) Insight into multi-site mechanisms of
glycosyl transfer in (1→4)-β-D-glycan synthases provided by the cereal mixed-linkage
(1→3)(1→4)-β-D-glucan synthase. Phytochemistry 57: 1045-1053
Buckeridge MS, Rayon C, Vergara CE, Carpita NC (2004) Mixed linkage (1→3)(1→4)-β-D-
glucans of grasses. Cereal Chem 81: 115-127
Burton RA, Wilson SM, Hrmova M, Harvey AJ, Shirley NJ, Medhurst A, Stone BA,
Newbigin EJ, Bacic A, Fincher GB (2006) Cellulose synthase-like CSLF genes mediate
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
22
the synthesis of cell wall (1,3;1,4)-β-D-glucans. Science 311: 1940–1942
Caño-Delgado A, Penfield S, Smith C, Catley M, Bevan M (2003) Reduced cellulose
synthesis evokes lignification and defense responses in Arabidopsis thaliana. Plant J 34:
351-362
Carpita NC (1996) Structure and biogenesis of the cell walls of grasses. Annu Rev Plant
Physiol Plant Molec Biol 47: 445-476
Carpita NC, Delmer DP (1980) Protection of cellulose synthesis in detached cotton fibers by
polyethylene glycol. Plant Physiol 66: 911-916
Carpita NC, Delmer DP (1981) Concentration and metabolic turnover of UDP-glucose in
developing cotton fibers. J Biol Chem 256: 308-315
Carpita NC, Vergara CE (1998) A recipe for cellulose. Science 279: 672-673
Carpita NC, Whittern D (1986) A highly substituted glucuronoarabinoxylan from developing
maize coleoptiles. Carbohydr Res 146: 129-140
Carpita N, McCann M, Griffing LR (1996) The plant extracellular matrix: News from the
cell’s frontier. Plant Cell 8: 1451-1463
Cavalier DM, Lerouxel O, Neumetzler L, Yamauchi K, Reinecke A, Freshour G, Zabotina
OA, Hahn MG, Burgert I, Pauly M, Raikhel NV, Keegstra K (2008) Disrupting two
Arabidopsis thaliana xylosyltransferase genes results in plants deficient in xyloglucan, a
major primary cell wall component. Plant Cell 20: 1519-1537
Chambers J, Elbein AD (1970) Biosynthesis of glucans in mung bean seedlings – Formation of
β-(1→4)-glucans from GDP-glucose and β-(1→3)-glucans from UDP-glucose. Arch
Biochem Biophys 138: 620-631
Charnock S J Davies G J (1999) Structure of the nucleotidediphospho-sugar transferase SpsA
from Bacillus subtilis in native and nucleotide-complexed forms. Biochemistry 38: 6380-
6385
Charnock S J Henrissat B Davies G J (2001) Three-dimensional structures of UDP-sugar
glycosyltransferases illuminate the biosynthesis of plant polysaccharides. Plant Physiol
125: 527-531
Chen S, Ehrhardt DW, Somerville CR (2010) Mutations of cellulose synthase (CESA1)
phosphorylation sites modulate anisotropic cell expansion and bidirectional mobility of
cellulose synthases. Proc Natl Acad Sci USA 107: 17188-17193
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
23
Cocuron JC, Lerouxel O, Drakakaki G, Alonso AP, Liepman AH, Keegstra K, Raikhel N,
Wilkerson CG (2007) A gene from the cellulose synthase-like C family encodes a β-1,4
glucan synthase. Proc Natl Acad Sci USA 104: 8550-8555
Coleman HD, Yan J, Mansfield SD (2009) Sucrose synthase affects carbon partitioning to
increase cellulose production and altered cell wall ultrastructure. Proc Natl Acad Sci USA
106: 13118-13123
Crowell EF, Bischoff V, Desprez T, Rolland A, Stierhof Y-D, Schumacher K, Gonneau M,
Vernhettes S (2009) Pausing of Golgi bodies on microtubules regulates secretion of
cellulose synthase complexes in Arabidopsis. Plant Cell 21: 1141-1154
Daras G, Rigas S, Penning B, Milioni D, McCann MC, Carpita NC, Fasseas C, Hatzopoulos
P (2009) The thanatos mutation in Arabidopsis thaliana cellulose synthase3 (AtCesA3) has
a dominant-negative effect on cellulose synthesis and plant growth. New Phytol 184: 114-
126
DeAngelis PL, Weigel PH (1994) Immunochemical confirmation of the primary structure of
streptococcal hyaluronan synthase and synthesis of high molecular weight product by
recombinant enzyme. Biochemistry 33: 9033-9039
Debolt S, Scheible W-R, Schrick K, Auer M, Beisson F, Bischoff V, Bouvier-Navé P,
Carroll A, Hematy K, Li Y, Milne J, Nair M, Schaller H, Zemla M, Somerville C
(2009) Mutations in UDP-glucose:sterol glucosyltransferase in Arabidopsis cause
transparent testa phenotype and suberization defect in seeds. Plant Physiol 151: 78-87
Delmer DP (1977) Biosynthesis of cellulose and other plant cell wall polysaccharides. Rec Adv
Phytochem 11: 105-153
Delmer DP (1999) Cellulose biosynthesis: exciting times for a difficult field of study. Annu Rev
Plant Physiol Plant Mol Biol 50: 245-276
Delmer DP, Amor Y (1995) Cellulose biosynthesis. Plant Cell 7: 987-1000
Delmer DP, Benziman M, Padan E (1982) Requirement for a membrane potential for cellulose
synthesis in intact cells of Acetobacter xylinum. Proc Natl Acad Sci USA 79: 5282-5286
Delmer DP, Solomon M, Read SM (1991) Direct labeling with [32P]UDP-glucose for
identification of a subunit of cotton fiber callose synthase. Plant Physiol 95: 556-563
Desprez T, Juraniec M, Crowell EF, Jouy H, Pochylova Z, Parcy F, Höfte H, Gonneau M,
Vernhettes S (2007) Organization of cellulose synthase complexes involved in primary
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
24
cell wall synthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 104: 15572-15577
Dhugga KS, Barreiro R, Whitten B, Stecca K, Hazebroek J, Randhawa GS, Dolan M,
Kinney AJ, Tomes D, Nichols S, Anderson P (2004) Guar seed β-mannan synthase is a
member of the cellulose synthase super gene family. Science 303: 363-366
Doblin MS, Kurek I, Jacob-Wilk D, Delmer DP (2002) Cellulose biosynthesis in plants: from
genes to rosettes. Plant Cell Physiol 43: 1407-1420
Doblin MS, Pettolino FA, Wilson SM, Campbell R, Burton RA, Fincher GB, Newbigin E,
Bacic A (2009) A barley cellulose synthase-like CSLH gene mediates (1,3;1,4)-β-D-glucan
synthesis in transgenic Arabidopsis. Proc Natl Acad Sci USA 106: 5996-6001
Dwivany FM, Yulia D, Burton RA, Shirley NJ, Wilson SM, Fincher GB, Bacic A, Newbigin
E, Doblin MS (2009) The CELLULOSE SYNTHASE-LIKE C (CSLC) family of barley
includes members that are integral membrane proteins targeted to the plasma membrane.
Molecular Plant 2: 1025-1039
Elbein AD, Hassid WZ (1966) The enzymatic synthesis of a glucomannan. Biochem Biophys
Res Comm 23: 311-318
Ellis C, Turner JG (2001) The Arabidopsis mutant cev1 has constitutively active jasmonate and
ethylene signal pathways and enhanced resistance to pathogens. Plant Cell 13: 1025-1033
Fagard M, Desnos T, Desprez T, Goubet F, Refregier G, Mouille G, McCann M, Rayon C,
Vernhettes S, Höfte H (2000) PROCUSTE1 encodes a cellulose synthase required for
normal cell elongation specifically in roots and dark-grown hypocotyls of Arabidopsis.
Plant Cell 12: 2409-2423
Freemont PS (2000) Ubiquitination: RING for destruction? Curr Biol 10: R84-R87
Fry SC, York WS, Albersheim P, Darvill A, Hayashi T, Joseleau JP, Kato Y, Lorences EP,
Maclachlan GA, McNeil M, Mort AJ, Reid JSG, Seitz HU, Selvendran RR, Voragen
AGJ, White AR (1993) An unambiguous nomenclature for xyloglucan-derived
oligosaccharides. Physiol Plant 89: 1-3
Fry SC, Nesselrode BHWA, Miller JG, Mewburn BR (2008) Mixed-linkage (1 3),(1 4)-β-D-
glucan is a major hemicellulose of Equisetum (horsetail) cell walls. New Phytol 179: 104-
115
Fujii S, Hayashi T, Mizuno K (2010) Sucrose synthase is an integral component of the
cellulose synthase machinery. Plant Cell Physiol 51: 294-301
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
25
Gardiner JC, Taylor NG, Turner SR (2003) Control of cellulose synthase complex
localization in developing xylem. Plant Cell 15: 1740-1748
Gibeaut DM, Carpita NC (1993) Synthesis of (1→3),(1→4)-β-D-glucan in the Golgi apparatus
of maize coleoptiles. Proc Natl Acad Sci USA 90: 3850-3854
Giddings TH, Brower DL, Staehelin LA (1980) Visualization of particle complexes in the
plasma membrane of Micrasterias denticulata associated with the formation of cellulose
fibrils in primary and secondary cell walls. J Cell Biol 84: 327-339
Gordon R, Maclachlan G (1986) Incorporation of UDP-[14C]Glucose into xyloglucan by pea
membranes. Plant Physiol 91: 373-378
Green PB (1962) Mechanism for plant cellular morphogenesis. Science 138: 1404-1405
Guerreiro G, Fugelstad J, Bulone V (2010) What do we really know about cellulose
biosynthesis in higher plants? J Integr Plant Biol 52: 161-175
Gutierrez R, Lindeboom JJ, Paredez AR, Emons AMC, Ehrhardt DW (2009) Arabidopsis
cortical microtubules position cellulose synthase delivery to the plasma membrane and
interact with cellulose synthase trafficking compartments. Nature Cell Biol 11: 797-808
Haigler CH, Brown RM Jr (1986) Transport of rosettes from the Golgi apparatus to the plasma
membrane in isolated mesophyll cells of Zinnia elegans during differentiation to tracheary
elements in suspension culture. Protoplasma 134: 111-120
Hayashi T, Koyama T, Matsuda K (1988) Formation of UDP-xylose and xyloglucan in
soybean Golgi membranes. Plant Physiol 87: 341-345
Hazen SP, Scott-Craig JS, Walton JD (2002) Cellulose synthase-like genes of rice. Plant
Physiol 128: 336-340
Held MA, Boulaflous A, Brandizzi F (2008) Advances in fluorescent protein-based imaging for
the analysis of plant endomembranes. Plant Physiol 147: 1469-1481
Herth W (1985) Plasma-membrane rosettes involved in localized wall thickening during xylem
vessel formation of Lepidium sativum L. Planta 164: 12-21
Hirai N, Sonobe S, Hayashi T (1998) In situ synthesis of β-glucan microfibrils on tobacco
plasma membrane sheets. Proc Natl Acad Sci USA 95: 15102-15106
Hoffman M, Jia ZH, Peña MJ, Cash M, Harper A, Blackburn AR, Darvill A, York WS
(2005) Structural analysis of xyloglucans in the primary cell walls of plants in the subclass
Asteridae. Carbohydr Res 340: 1826-1840
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
26
Hong Z, Delauney AJ, Verma DPS (2001) A cell plate-specific callose synthase and its
interaction with phragmoplastin. Plant Cell 13: 755-768
Imai T, Watanabe T, Yui T, Sugiyama J (2003) The directionality of chitin biosynthesis: a
revisit. Biochem J 374: 755-760
Jacob-Wilk D, Kurek I, Hogan P, Delmer DP (2006) The cotton fiber zinc-binding domain of
cellulose synthase A1 from Gossypium hirsutum displays rapid turnover in vitro and in
vivo. Proc Nat Acad Sci USA 103: 12191-12196
Jing W, DeAngelis PL (2000) Dissection of the two transferase activities of the Pasteurella
multocida hyaluronan synthase: two active sites exist in one polypeptide. Glycobiology 10:
883-889
Kennedy CJ, Cameron GJ, ˇSturcová A, Apperley DC, Altaner C, Wess TJ, Jarvis MC
(2007) Microfibril diameter in celery collenchyma cellulose: X-ray scattering and NMR
evidence. Cellulose 14: 235-246
Klepek Y-S, Volke M, Konrad KR, Wippel K, Hoth S, Hedrich R, Sauer N (2009)
Arabidopsis thaliana POLYOL/MONOSACCHARIDE TRANSPORTERS 1 and 2:
fructose and xylitol/H+ symporters in pollen and young xylem cells. J Exp Bot 61: 537-550
Konopka CA, Bednarek SY (2008) Variable-angle epifluorescence microscopy: a new way to
look at protein dynamics in the plant cell cortex. Plant J 53: 186-196
Koyama M, Helbert W, Imai T, Sugiyama J, Henrissat B (1997) Parallel-up structure
evidences the molecular directionality during biosynthesis of bacterial cellulose. Proc Natl
Acad Sci USA 94: 9091-9095
Kudlicka K, Brown RM (1997) Cellulose and callose biosynthesis in higher plants. 1.
Solubilization and separation of (1→3)- and (1→4)-β-glucan synthase activities from mung
bean. Plant Physiol 115: 643-656
Kurek I, Kawagoe Y, Jacob-Wilk D, Doblin M, Delmer D (2002) Dimerization of cotton fiber
cellulose synthase catalytic subunits occurs via oxidation of the zinc-binding domains. Proc
Natl Acad Sci USA 99: 11109-11114
Kuroyama H, Tsumuraya Y (2001) A xylosyltransferase that synthesizes β-(1→4)-xylans in
wheat (Triticum aestivum L.) seedlings. Planta 213: 231-240
Lai-Kee-Him J, Chanzy H, Muller M, Putaux JT, Imai T, Bulone V (2002) In vitro versus in
vivo cellulose microfibrils from plant primary wall synthases: Structural differences. J Biol
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
27
Chem 277: 36931-36939
Lao NT, Long D, Kiang S, Coupland G, Shoue DA, Carpita NC, Kavanagh TA (2003)
Mutation of a family 8 glycosyl transferase gene alters cell wall carbohydrate composition
and cause a humidity-sensitive semi-sterile dwarf phenotype in Arabidopsis. Plant Mol
Biol 53: 647-661
Ledbetter MC, Porter KR (1963) A “microtubule” in plant cell fine structure. J Cell Biol 19:
239-250
Lee CH, Zhong RQ, Richardson EA, Himmelsbach DS, McPhail BT, Ye ZH (2007) The
PARVUS gene is expressed in cells undergoing secondary wall thickening and is essential
for glucuronoxylan biosynthesis. Plant Cell Physiol 48: 1659-1672
Lerouxel O, Cavalier DM, Liepman AH, Keegstra K (2006) Biosynthesis of plant cell wall
polysaccharides. Curr Opin Plant Biol 9: 621-630
Liepman AH, Wilkerson CG, Keegstra K (2005) Expression of cellulose synthase-like (Csl)
genes in insect cells reveals that CslA family members encode mannan synthases. Proc
Natl Acad Sci USA 102: 2221-2226
Liepman AH, Nairn CJ, Willats WGT, Sørensen I, Roberts AW, Keegstra K (2007)
Functional genomic analysis conservation of function among cellulose synthase-like A
gene family members and suggests diverse roles of mannans in plants. Plant Physiol 143:
1881-1893
Madson M, Dunand C, Verma R, Vanzin GF, Caplan J, Li X, Shoue DA, Carpita NC,
Reiter W-D (2003) The MUR3 gene of Arabidopsis encodes a xyloglucan
galactosyltransferase that is evolutionarily related to animal exostosins. Plant Cell 15:
1662-1670
Maiden MCJ, Davis EO, Baldwin SA, Moore DCM, Henderson PJF (1987) Mammalian and
bacterial sugar transport proteins are homologous. Nature 325: 641-643
Mitchell RAC, Dupree P, Shewry PR (2007) A novel bioinformatics approach identifies
candidate genes for the synthesis and feruloylation of arabinoxylan. Plant Physiol 144: 43-
53
Mueller SC, Brown Jr RM (1980) Evidence for an intramembrane component associated with a
cellulose microfibril synthesizing complex in higher plants. J Cell Biol 84: 315-326
Nishimura MT, Stein M, Hou B-H, Vogel JP, Edwards H, Somerville SC (2003) Loss of a
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
28
callose synthase results in salicylic acid-dependent disease resistance. Science 301: 969-
972
Nishitani K, Nevins DJ (1991) Glucuronoxylan xylanohydrolase. A unique xylanase with a
requirement for appendant glucuronosyl units. J Biol Chem 266: 6539-6543
Nobles DR, Romanovicz DK, Brown Jr RM (2001) Cellulose in Cyanobacter: Origin of
vascular plant cellulose synthase? Plant Physiol 127: 529-542
Nühse TS, Stensballe A, Jensen ON, Peck SC (2004) Phosphoproteomics of the Arabidopsis
plasma membrane and a new phosphorylation site database. Plant Cell 16: 2394-2405
Obel N, Erben V, Schwartz T, Kuhnel S, Fodor A, Pauly M (2009) Microanalysis of plant
cell wall polysaccharides. Molecular Plant 2: 922-932
Orellana A (2005) Biosynthesis of non-cellulosic polysaccharides in the Golgi apparatus.
Topological considerations. Plant Biosystems 139: 42-45
Pao SS, Paulsen IT, Saier Jr MH (1998) Major facilitator superfamily. Microbiol Mol Biol
Rev 62: 1-34
Paredez AR, Somerville CR, Ehrhardt DW (2006) Visualization of cellulose synthase
demonstrates functional association with microtubules. Science 312: 1491-1495
Pear JR, Kawagoe Y, Schreckengost WE, Delmer DP, Stalker D M (1996) Higher plants
contain homologs of the bacterial celA genes encoding the catalytic subunit of cellulose
synthase. Proc Natl Acad Sci USA 93: 12637-12642
Peña MJ, Zhong R, Zhou G-K, Richardson EA, O’Neill MA, Darvill AG, York WS, Ye Z-
H (2007) Arabidopsis irregular xylem8 and irregular xylem9: Implications for the
complexity of glucuronoxylan biosynthesis. Plant Cell 19: 549-563
Peña MJ, Darvill AG, Eberhard S, York WS, O’Neill MA (2008) Moss and liverwort
xyloglucans contain galacturonic acid and are structurally distinct from the xyloglucans
synthesized by hornworts and vascular plants. Glycobiology 18: 891-904
Peng LC, Xiang F, Roberts E, Kawagoe Y, Greve LC, Kreuz K, Delmer DP (2001) The
experimental herbicide CGA 325'615 inhibits synthesis of crystalline cellulose and causes
accumulation of non-crystalline β-14-glucan associated with CesA protein. Plant Physiol
126: 981-992
Peng L, Kawagoe Y, Hogan P, Delmer D (2002) Sitosterol-β-glucoside as primer for cellulose
synthesis in plants. Science 295: 147-150
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
29
Penning B, Hunter CT, Tayengwa R, Eveland AR, Dugard CK, Olek A, Vermerris W,
Davis M, Thomas SR, Koch KE, McCarty DR, McCann MC, Carpita NC (2009)
Genetic resources for maize cell wall biology. Plant Physiol 151: 1703-1728
Perrin RM, DeRocher AE, Bar-Peled M, Zeng W, Norambuena L, Orellana A, Raikhel
NV, Keegstra K (1999) Xyloglucan fucosyltransferase, an enzyme involved in plant cell
wall biosynthesis. Science 284: 1976-1979
Piro G, Zuppa A, Dalessandro G, Northcote DH (1993) Glucomannan synthesis in pea
epicotyls: The mannose and glucose transferases. Planta 190: 206–220
Porchia AC, Scheller HV (2000) Arabinoxylan biosynthesis: identification and partial
characterization of β-1,4-xylosyltransferase from wheat. Physiol Plant 110: 350-356
Porchia AC, Sørensen SO, Scheller HV (2002) Arabinoxylan biosynthesis in wheat.
Characterization of arabinosyltransferase activity in Golgi membranes. Plant Physiol 130:
432-441
Ray PM, Shininger TL, Ray MM (1969) Isolation of β-glucan synthetase particles from plant
cells and identification with Golgi membranes. Proc Natl Acad Sci USA 64: 605-612
Ray PM (1980) Cooperative action of β-glucan synthetase and UDP-xylose xylosyl transferase
of Golgi membranes in the synthesis of xyloglucan-like polysaccharide. Biochim Biophys
Acta 629: 431-444
Reifenberger E, Freidel K, Ciriacy M (1995) Identification of novel HXT genes in
Saccharomyces cerevisiae reveals the impact of individual hexose transporters on
glycolytic flux. Mol Microbiol 16: 157-167
Richmond TA, Somerville CR (2000) The cellulose synthase superfamily. Plant Physiol 124:
495-498
Robert S, Bichet A, Grandjean O, Kierzkowski D, Satiat-Jeunemaître B, Pelletier S,
Hauser M-T, Höfte H, Vernhettes S (2005) An Arabidopsis endo-1,4-β-D-glucanase
involved in cellulose synthesis undergoes regulated intracellular cycling. Plant Cell 17:
3378-3389
Roberts AW, Roberts EM, Delmer DP (2002) Cellulose synthase (CesA) genes in the green
alga Mesotaenium caldariorum. Eukaryotic Cell 1: 847-855
Saxena IM, Brown Jr RM (2005) Cellulose biosynthesis: Current views and evolving concepts.
Ann Bot 96: 9-21
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
30
Saxena IM, Brown Jr RM, Fevre M, Geremia RA, Henrissat B (1995) Multidomain
architecture of β-glucosyl transferases: implications for mechanism of action. J Bacteriol
177: 1419-1424
Saxena IM, Brown Jr RM, Dandekar T (2001) Structure-function characterization of cellulose
synthase: relationship to other glycosyltransferases. Phytochemistry 57: 1135-1148
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61: 263-289
Sherson SM, Hemmann G, Wallace G, Forbes S, Germain V, Stadler R, Bechtold N, Sauer
N, Smith SM (2000) Monosaccharide/proton symporter AtSTP1 plays a major role in
uptake and response of Arabidopsis seeds and seedlings to sugars. Plant J 24: 849-857
Sims IM, Munro SLA, Currie G, Craik D, Bacic A (1996) Structural characterization of
xyloglucan secreted by suspension cultured cells of Nicotiana plumbaginifolia. Carbohydr
Res 293: 147-172
Somerville C (2006) Cellulose synthesis in higher plants. Annu Rev Cell Devel Biol 22: 53-78
Sørensen I, Pettolino FA, Wilson SM, Doblin MS, Johansen B, Bacic A, Willats WGT
(2008) Mixed-linkage (1→3),(1→4)-β-D-glucan is not unique to the Poales and is an
abundant component of Equisetum arvense cell walls. Plant J 54: 510-521
Szymanski DB, Cosgrove DJ (2009) Dynamic coordination of cytoskeletal and cell wall
systems during plant cell morphogenesis. Curr Biol 19: R800-R811
Taylor NG (2008) Cellulose biosynthesis and deposition in higher plants. New Phytol 178: 239-
252
Taylor NG, Laurie S, Turner SR (2001) Multiple cellulose synthase catalytic subunits are
required for cellulose synthesis in Arabidopsis. Plant Cell 12: 2529-2539
Taylor NG, Howells RM, Huttly AK, Vickers K, Turner SR (2003) Interactions among three
distinct CesA proteins essential for cellulose synthesis. Proc Natl Acad Sci USA 100:
1450-1455
Timmers J, Vernhettes S, Desprez T, Vincken J-P, Visser RGF, Trindade LM (2009)
Interactions between membrane-bound cellulose synthases involved in the synthesis of the
secondary wall. FEBS Lett 583: 978-982
Tiné MAS, Silva CO, de Lima DU, Carpita NC, Buckeridge MS (2006) Fine structure of a
mixed-oligomer storage xyloglucan from seeds of Hymenaea courbaril. Carbohydr Polym
66: 444-454
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
31
Urbanowicz BR, Rayon C, Carpita NC (2004) Topology of the maize mixed-linkage
(1→3)(1→4)-β-D-glucan synthase at the Golgi membrane. Plant Physiol 134: 758-768
Vanzin GF, Madson M, Carpita NC, Raikhel NV, Keegstra K, Reiter WD (2002) The mur2
mutant of Arabidopsis thaliana lacks fucosylated xyloglucan because of a lesion in
fucosyltransferase AtFUT1. Proc Natl Acad Sci USA 99: 3340-3345
Vergara CE, Carpita NC (2001) β-D-Glucan synthases and the CesA gene family: lessons to
learned from the mixed-linkage (1→3)(1→4)-β-D-glucan synthase. Plant Mol Biol 47:
145-160
Waldron KW, Brett CT (1983) A glucuronosyltransferase involved in glucuronoxylan
synthesis in pea (Pisum sativum) epicotyls. Biochem J 213: 115-122
Wang J, Howles PA, Cork AH, Birch R, Williamson RE (2006) Chimeric proteins suggest
that the catalytic and/or C-terminal domains give CesA1 and CesA3 access to their specific
sites in the cellulose synthase of primary walls. Plant Physiol 142: 685-695
Wang J, Elliott JE, Williamson RE (2008) Features of the primary wall CESA complex in wild
type and cellulose deficient mutants of Arabidopsis thaliana. J Exp Bot 59: 2627-2637
Weigel PH, DeAngelis PL (2007) Hyaluronan synthases: a decade-plus of novel
glycosyltransferases. J Biol Chem 282: 36777-36781
Wightman R, Turner SR (2008) The roles of the cytoskeleton during cellulose deposition at the
secondary wall. Plant J 54: 794-805
Williams KJ, Halkes KM, Kamerling JP, DeAngelis PL (2006) Critical elements of
oligosaccharide acceptor substrates for the Pasteurella multocida hyaluronan synthase. J
Biol Chem 281: 5391-5397
Wood PJ, Weisz J, Blackwell BA (1994) Structural studies of (1→3)(1→4)-β-D-glucan by 13C-
nuclear magnetic resonance spectroscopy and by rapid analysis of cellulose-like regions
using high-performance anion exchange chromatography of oligosaccharides released by
lichenase. Cereal Chem 71: 301-307
Yong W, O’Malley R, Link B, Binder B, Bleecker A, Koch KE, McCann MC, McCarty
DR, Patterson S, Reiter W-D, Staiger CJ, Thomas S, Vermerris W, Carpita NC (2005)
Plant cell wall genomics. Planta 221: 747-751
York WS, O’Neill MA (2008) Biochemical control of xylan biosynthesis – which end is up?
Curr Opin Plant Biol 11: 258-265
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
32
Yoshida M, Itano N, Yamada Y, Kimata K (2000) In vitro synthesis of hyaluronan by a single
protein derived from mouse HAS1 gene and characterization of amino acid residues
essential for activity. J Biol Chem 275: 497-506
Zabotina OA, van de Ven WTG, Freshour G, Drakakaki G., Cavalier D, Mouille G, Hahn
MG, Keegstra K, Raikhel NV (2008) Arabidopsis XXT5 gene encodes a putative α-1,6-
xylosyltransferase that is involved in xyloglucan biosynthesis. Plant J 56: 101-115
Zeng W, Chatterjee M, Faik A (2008) UDP-xylose-stimulated glucuronyltransferase activity in
wheat microsomal membranes: Characterization and role in glucurono(arabino)xylan
biosynthesis. Plant Physiol 147: 78-91
Zeng W, Jiang N, Nadella R, Killen TL, Nadella V, Faik A (2010) A glucurono(arabino)
xylan synthase complex from wheat contains members of the GT43, GT47, and GT75
families and functions cooperatively. Plant Physiol 154: 78-97
Zhong R, Morrison III WH, Freshour G, Hahn MG, Ye Z-H (2003) Expression of a mutant
form of cellulose synthase AtCesA7 causes dominant negative effect on cellulose
biosynthesis. Plant Physiol 132: 786-795
Zhong R, Peña MJ, Zhou G-K, Nairn CJ, Wood-Jones A, Richardson EA, Morrison III
WH, Darvill AG, York WS, Ye Z-H (2005) Arabidopsis Fragile Fiber8, which encodes a
putative glucuronyltransferase, is essential for normal secondary wall synthesis. Plant Cell
17: 3390-3408
https://plantphysiol.orgDownloaded on April 2, 2021. - Published by Copyright (c) 2020 American Society of Plant Biologists. All rights reserved.
https://plantphysiol.org
-
33
Table I. Six possible xyloglucan oligomers are produced by digestion with a Trichoderma
endoglucanase. Because of characteristic side-group construction of these
complex glucans, a single-letter code was devised to denote the non-reducing
terminal sugar of the side chain, with other constituents understood (Fry et al.,
1993). Thus, XXXG signifies a tetraglucosyl backbone with three residues
bearing a xylosyl side-groups. For many angiosperms, this heptasaccharide is
decorated further with variable amounts of Gal at the two Xyl residues closer to
the reducing end, and if Gal is added to the first Xyl residue, a Fuc residue is
usually added. In addition to the glucan backbone synthase, at least three types of
glycosyl transferases, and as many as six, are needed to construct all the side-
gro