Protein Structure
-
Upload
ajit-suryawanshi -
Category
Documents
-
view
235 -
download
0
description
Transcript of Protein Structure
-
Protein: Structure and Function
Books:
1. Lehninger Principle of Biochemistry ( by Nelson and Cox)
-
Amino acid
Properties:1. -carbon is bonded to four different groups except glycine2. -carbon is a chiral center3. Two possible stereoisomers (called enantiomers)4. Optically active (except glycine): rotate plan polarized light5. Can act as acids and bases
-
Amino acids in the human body
Essential amino acid
Non-essential amino acid
1. Leucine2. Isoleucine3. Valine4. Histidine5. Lysine
6. Methionine7. Phenylalanine8. Threonine9. Tryptophan
1. Alanine2. Arginine3. Asparagine4. Aspartic acid5. Cysteine6. Glutamic acid7. Glutamine
8. Glycine9. Proline10. Serine11. Tyrosine
Obtained from nutrition
Synthesized by the body
-
Non-polar, aliphatic R groups
Gly/G Ala/A Pro/P Val/V
Leu/L Ile/I Met/M
-
Polar, uncharged R groups
1. Cysteine forms dimercalled cystine
2. Disulfide residues are strongly hydrophobic
Ser/S Thr/T Cys/C
Asn/N Gln/Q
-
Positively charged R groups
Lys/K Arg/R His/H
-
Negatively charged R groups
Asp/D Glu/E
-
Aromatic R groups
1. Non-polar2. Hydrophobic3. Form H-bond4. Absorb UV light (280 nm)
Phe/F Tyr/Y Trp/W
-
pKa values for carboxyl and amino groups
-
Amino acid have characteristic titration curve1. Without ionisable R group
pI = (pK1 + pK2 )/2
pK1 = for acidpK2 = for base
-
2. With ionisable R group
pI = ?
-
2. With ionisable R group
pI = ?
-
1. All protein polymers are constructed from the same set of 20 amino acids.
2. Polymers of proteins are called polypeptides.
3. A protein consists of one or more polypeptides folded and coiled into a specific conformation
4. The physical and chemical characteristics of the R group determine the unique characteristics of a particular amino acid.
Protein
-
Peptide Bond
-
Properties of Peptide bond
1. Planar2. Double bond character (which prevent rotation
about this bond)3. Uncharged (helps to form tightly packed globular
structure4. Two conformations possible (cis and trans)5. All peptide bonds are in proteins are trans6. Other than peptide bonds in protein helps to take
many conformational structure7. Average half-life of peptide bond is 7 years
(intracellular condition)
-
Fully extended polypeptide chain
Both bond can rotate
and are zero
-
trans-Peptide group
-
cis-Peptide group
-
Peptide and Protein
Peptides: 1. Short polymers formed from the linking of (usually less
than or equal to 100) amino acids and comprise 2. Some of the basic components of human biological
processes, including enzymes, hormones, and antibodies.
Protein:1. A functional, polypeptide chain composed of at
least around fifty amino acids put together. 2. They play a critical role in biochemical reactions
within cells.
-
1. Simple protein: contain only amino acid residues2. Conjugated proteins: contain permanently associated chemical
components in addition to amino acids(a) Lipoproteins: contain lipid(b) glycoproteins: contain sugar(c) metalloprotein: contain specific metal
Types of Protein
-
Structure of Protein
I. The primary structure determines the folding of the polypeptide to give a functional protein
II. Polar amino acids (acidic, basic and neutral) are hydrophilic and tend to be placed on the outside of the protein.
III. Non-polar (hydrophobic) amino acids tend to be placed on the inside of the protein
IV. The possible conformations are very large
V. Most are useless, natural selection picks out the best
Primary Structure
-
1. Elongated2. Dynamic, heterogeneous3. Very rich in H-bond4. Every protein has unique amino acid sequence5. Sequence decide the mechanism of action6. Sequence determine the 3-D structure7. Sequence reveals about its evolutionary history
Primary structure of protein
1. Starting of the polypeptide chain : N-terminal (amino group)
2. End terminal of the polypeptide chain: C-terminal (carboxylic group)
-
Human insulin
-
1. Strong interactions(a) Covalent bonding(b) Ionic bonding(c) Resonance bonding
2. Weak Interaction(a) Van der Waals interaction
(i) Polar-polar (ii) Polar-non polar(iii) Non polar- non polar
(b) Hydrogen bonding
3. Effect of medium(a) Screening of field(b) Surface interaction(c) Hydrophobic environment(d) Hydrophilic environment
Molecular interactions
-
The polypeptide chain can fold into regular structures like:
1. Alpha helix2. Beta-sheet3. Turns and loops
These are called secondary structure which helps to form final 3-D structure
Secondary structure of proteinThe folding of the N-C terminals of the chain using many different interactions
-
Alpha-helix1. It is coiled structure stabilized by intra-chain H-bonds
between NH and CO groups (situated 4 residue ahead in the sequence) except end terminal groups
2. Pitch of alpha-helix 5.4 A
3. Both right handed and left handed helix are allowed , however right handed alpha-helices are energetically more favourable because there is less steric clash between the side chain and backbone
4. Content in the protein: alpha-helices may be 100%
5. Glycine, serine and threonine make amino terminal residue (N-cap) in alpha-helix
6. Glycine and asparagine makes carboxyl terminal (C-cap) of alpha-helix
-
Helical Wheel: Each residue can be plotted every 360/3.6=100 around a circle or spiral
Representation of -helices in protein
-
-helicesH bond between residues i, i+4
Rise per residue, d = 1.5
# of residues per turn, n = 3.6
Pitch of helix= n x d = 5.4
-
The -helix has a dipole moment
The dipole of a peptide unit. Numbers in boxes give the approximate fractional charges of the atoms of the peptide unit
The Dipoles of peptide units are aligned along the helical axis
-
Beta- sheet/strand1. It is stabilized by H-bonding between chains
2. This secondary structure may associated through side chain interactions and form super-secondary structure called motif
3. Beta-sheet is almost fully extended
4. The distance between adjacent amino acids along a beta-strand is roughly 3.5 A (in contrast to 1.5 A in alpha-helix)
5. A beta- sheet is formed by linking two pr more beta strands by H-bonds
6. Beta strand represented by broad arrows pointing in the direction of C-terminal
7. Beta sheet formation is important in fatty acid-binding proteins and lipid metabolism
-
-Strand
The side chain (green) are alternatively above and below the plane of the strand
-
NN
C
C
Anti-parallel -sheet
Adjacent -strand run in opposite direction. H-bond between NH and CO groups connect each amino acid on an adjacent strand and stabilize the structure
-
Parallel -sheet
Adjacent -strand run in same direction. H-bond between NH and CO groups connect each amino acid on one strand with two different amino acids on the adjacent strand
N
N
C
C
-
Mixed -sheet
-
Turns and Loops
1. Polypeptide chain can change direction with the help of turn and loops
2. CO group of residue i is H-bonded with NH group of residue i+3
3. This particular H-bond interaction stabilizes abrupt changes in the direction of polypeptide chain
4. Turn and loops connect alpha-helices and beta strand and allow a peptide chain to fold back on itself to make a compact structure
5. Loops often contain hydrophilic residues and are found on the protein surface
6. Turn or loops contain 5 residues or less
7. Beta turn connects different anti-parallel beta strands
-
Ramachandran plot
Phi ()
Psi ()
Phi () and Psi () rotate, allowing the polypeptide to assume its various conformations
some conformations of the polypeptide backbone result in steric hindrance and are disallowed
glycine has no side chain and is therefore conformationally highly flexible (it is often found in turns)
no stericclashes
permittedif atoms aremore closelyspaced
-
Tertiary structure of protein
Spatial arrangement of amino acid residues that are far apart in the sequence
1. This folding is sometimes held together by strong covalent bonds (e.g. cysteine-cysteine disulphide bridge)
2. Bending of the chain takes place at certain amino acids (e.g. proline)
3. Hydrophobic amino acids tend to arrange themselves insidethe molecule
4. Hydrophilic amino acids arrange themselves on the outside
-
Quaternary structure of protein
1. Polypeptide chains can assemble into multi sub-units
2. Sub units are spatially arranged
3. Helix-loop-helix: two helices connected by a turn
4. Coiled-coil: two alpha helices interact in parallel through their hydrophobic edge
5. Helix-bundle: several alpha-helices that associate in an anti-parallel manner
6. Beta-alpha-beta unit: two parallel beta strand linked to an intervening alpha helix by two loops
-
Levels of structure in proteins
-
Protein structure: overview
Structural element Description
primary structure amino acid sequence of protein
secondary structure helices, sheets, turns/loops
super-secondary structure (motif) association of secondary structures
domain self-contained structural unit
tertiary structure folded structure of whole protein includes disulfide bonds
quaternary structure assembled complex (oligomer) homo-oligomeric (1 protein type) hetero-oligomeric (>1 type)
-
Physical parameters of protein
Size of the protein roughly between 1nm to 10nm
Persistence length of protein ranges between 0.3 nm to 0.8 nm
Elastic modulus of protein ranges between 1200 to 2000 pN/nm2
-
The protein structure must obey
1. The bond lengths and bond angles should be distorted as little as possible
2. No two atoms should approach one another more closely than is allowed by there van der Waals radii
3. The amide group must remain planar and in the trans configuration. This allows only rotation about the two bonds adjacent to the alpha-carbon
4. Some kind of non-covalent binding is necessary to stabilized a regular folding
-
Domain structures core is exclusively built from helices
Domain structures core comprises of antiparallel sheets, usually two sheets packed against each other
/ Domain structures made from combinations of -- motifs that form a predominantly parallel sheets surrounded by helices
Structure of proteins
-
Human plasma retinol binding protein. Retinol molecule (vitamin A) bound inside the barrel
Triosephosphateisomerase
Structure of proteins
-
Structure Function
Structure Mechanism
Structure Origins/Evolution
Structure-based Drug Design
Solving the Protein Folding Problem
Solving Protein Structures
Atomic resolution pictures of macromolecules X-ray Crystallography (first applied in 1961 - Kendrew & Perutz) NMR Spectroscopy (first applied in 1983 - Ernst & Wuthrich)
QHTAWCLTSEQHTAAVIWDCETPGKQNGAYQEDCAHHHHHHCCEEEEEEEEEEECCHHHHHHHCCCCCCC
-
Crystallographic structure of Myoglobin
10
(1958, Sir John Kendrew)
-
Protein Structure solved by X-ray crystallography
-
PDB contains 75000 structures mostly determined by X-ray crystallography and NMR. About 3-5 new structures per day
Total Yearly
-
Hemoglobin A: Val-His-Leu-Thr-Pro-Glu-Glu-Lys-Hemoglobin S: Val-His-Leu-Thr-Pro-Val-Glu-Lys-
sticky patch causes hemoglobin S to agglutinate (stick together) and form fibers which deform the red blood cell
Importance of Protein StructureUsing electrophoresis, Pauling showed that individuals with sickle cell disease had a modified form of Hb
-
101 residues. each residue can assume three different conformations
the total number of structures would be 3100, which is equal to 5 1047
If it takes 10-13 s to convert one structure into another
the total search time would be 5 1047 10-13 s
which is equal to 5 1034 s, or 1.6 1027 years.
Protein folding: Levinthals paradox
The enormous difference between calculated and actual folding times is called Levinthal's paradox.
There should be some pathways for folding
-
Factors affecting protein folding
1. Space packingproteins are like liquid and gases instead of crystalline solidit helps in forming structure but space packing is not enough
2. Internal residue: Folding is directed mainly by internal residues not by surface residues. (Hydrophobic force-driven folding)
3. Protein structures are hierarchically organized
4. Protein structures are highly adaptable
5. Secondary structure can be context dependent and can be predicted by algorithms
6. Changing the fold of a protein
-
Speed limit of protein folding
For a single domain protein The approximate folding time (folding) is given by N/100 s
-protein fold faster than the protein or protein
folding = k exp(G/kBT)
-
Hydrophobic effect
Conformational entropy
Electrostatics
Hydrogen bonding
van der Waals interaction
Protein folding
The main driving force for folding water soluble globularprotein molecules is to pack hydrophobic side chains into the interior of the molecule , thus creating a HYDROPHOBIC CORE &HYDROPHILLIC SURFACE.
Problem- How to create such a hydrophobic core from a protein chain ???
-
?Protein folding
Compact (in general)Defined structure
Insulin
Molten Globule
1. Secondary structure that is present in a native protein forms within a few microsecond
2. This is because of hydrophobic collapse3. It is larger by (5-15%) in size of native conformations4. Side chains are not ordered/packed5. Structure fluctuation is much larger 6. Not thermodynamically stable
-
Folding protein moves over energy surface from unfolded to folded state:
unfolded
folded
Energy landscape governs folding
100
k BT
Deg
ree
of
nativ
enes
s
H= bond stretching + bending of angles + Bond rotations + van der waals interaction + electrostatic interaction
-
Folding is a complex process
Misfolding
Macromolecules must fold into correct shape to function properly: Structure Function
Disease(non-native structures)
Many diseases with large impacts involve protein misfolding:
Alzheimers Disease A peptide
Parkinsons Disease -synuclein
Creutzfeldt-Jakob disease Prion
Huntingtons Disease Huntingtin protein
Amyotrophic Lateral Sclerosis Superoxide dismutase
Type II Diabetes Amylin
-
Misfolding and aggregation are complex
nativelyfolded
amyloid fibrils
Many different species and steps involved in misfolding and aggregation:
protein synthesis unfolded
ribosome
partiallyunfolded
partially-unfoldedand aggregated
disorderedaggregates
degradedfragments
non-native structuredaggregates
misfolded
Chiti & Dobson, Annu. Rev. Biochem., (2006)
-
Force-extension curvesMove traps apart at constant rate to stretch handles and apply force to a single-molecule:
50 mM MOPS, 200 mM KCl, pH 7.0
unfolding
stretching handles
molecule unfoldsrefolding
Apparently two-state folding
molecule unfolds
-
Reconstructing landscapes from FECsBased on Jarzysnki equality for relating equilibrium free energy to non-equilibrium work:
Hummer & Szabo, PNAS (2001)
Many applications, never validated experimentally
Forc
e
Distance
Work
But energy is dissipated!
WGeqm
Recover equilibrium energy fromfluctuations in work done:
Prob
abili
ty
Work
Geqm
W
Jarzynski, Phys. Rev. Lett. (1997)
Jarzynski equality:
=
TkW
TkG
B
eqmnon
B
eqm expexp
Method to calculate free energy profile G(x)
-
Reconstructing energy landscape profilesUsing equilibrium probability distributions:
Exte
nsio
n (n
m)
Time (s)
Prob
abili
ty d
ensit
y
( ) ( )expB
G xP x
k T
( ) ( )logBG x k T P x =
Thousands oftransitions!
Extension (nm)
Free energy (kJ/mol)
Invert
Deconvolution to remove effects of compliant handles:
Prob
abili
ty
Extension (nm)
P(x)
deconvolution
Residual
Free
ene
rgy
(kJ/
mol
)
Extension (nm)
G(x)
raw data
Woodside et al., Science (2006)
parameter-free modelfor folding
Woodside et al., PNAS (2006)
Slow and time consuming experiments
-
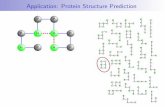
Computational approach for protein folding
1. Energy minimization(a) Steepest(b) Conjugated gradient
2. Monte Carlo Simulation(a) Random Sampling(b) Stimulated annealing
3. Molecular dynamics(a) Compute conformational change(b) Calculate trajectories at thermal condition and fond
the ensemble averaged physical quantity
-
Denaturants high temperatures
- cause protein unfolding, aggregation
low temperatures- some proteins are sensitive to cold denaturation
heavy metals (e.g., lead, cadmium, etc.)- highly toxic; efficiently induce the stress response
proteotoxic agents (e.g., alcohols, cross-linking agents, etc.)
oxygen radicals, ionizing radiation- cause permanent protein damage
chaotropes (urea, guanidine hydrochloride, etc.)- highly potent at denaturing proteins;often used in protein folding studies
Protein folding/unfolding
-
Force spectroscopy
AFM Optical Tweezers
-
Protein-Protein Interaction NetworksYeast ~6000 proteins, ~3 interactions per protein, i.e. ~>20,000 interactions. Humans ~100,000 interactions
Nat. Biotechnol. 18, 12571261 (2000)
Which two proteins will interact?
AND, which will not?
The ANSWER lies in the nature of theinteracting surfaces
A-B, A-C forms poorly matched surfaces, few weak bonds are formed, broken apart by thermal motion
A-D offers well matched surfaces, enough noncovalent bonds are formed to create a stable interface
-
Forces driving protein-protein interaction
Long-range attractive interactions
electrostatic steering
Short-range non-covalent forces:
Hydrophobic interactions
van der Waals attraction
Hydrogen bonds
Ion pairs
Other factors:
Shape and charge complementarity
Secondary structure
Amino acid composition
Slide Number 1Slide Number 2Slide Number 3Slide Number 4Slide Number 5Slide Number 6Slide Number 7Slide Number 8Slide Number 9Slide Number 10Slide Number 11Slide Number 12Slide Number 13Slide Number 14Slide Number 15Slide Number 16Slide Number 17Slide Number 18Slide Number 19Slide Number 20Slide Number 21Slide Number 22Slide Number 23Slide Number 24Slide Number 25Slide Number 26Slide Number 27Helical Wheel: Each residue can be plotted every 360/3.6=100 around a circle or spiralSlide Number 29Slide Number 30Slide Number 31Slide Number 32Slide Number 33Slide Number 34Slide Number 35Slide Number 36Ramachandran plotSlide Number 38Slide Number 39Slide Number 40Protein structure: overviewSlide Number 42Slide Number 43Slide Number 44Slide Number 45Slide Number 46Slide Number 47Slide Number 48Slide Number 49Slide Number 50Slide Number 51Slide Number 52Slide Number 53Slide Number 54Slide Number 55Slide Number 56Slide Number 57Slide Number 58Slide Number 59Slide Number 60Slide Number 61Slide Number 62DenaturantsSlide Number 64Slide Number 65Slide Number 66