graduate school of medicine, the University of Tokyo ...
Transcript of graduate school of medicine, the University of Tokyo ...

How molecular motor acts
2006.11.27. graduate school of medicine, the University of Tokyo Nobutaka Hirokawa
Life Science Seen from Molecular Motor

Low-angle, rotary shadowing electronmicrograph
Kinesin
KIF1A



Okada et al.Science 283:1152-, 1999






Optical Trapping Nanometry
Spatial resolution < 0.2 nmTemporal resolution < 1 ms
0.2μm beads⇒Small viscous drag
⇒ High temporal resolution
(Response time ≦ 0.25 ms)
Okada,Higuchi, and Hirokawa Nature 424:574-2003

Immobilization of KIF1A to Bead

Bidirectional and Step-wise Movement of a Single KIF1A Monomer

One ATP Hydrolysis Triggers One Stepping Movement.
Time (s)8 9 10
4080
120
-20 0 20
(nm)
(ms)-400
4080
120160
Cycle Time (ms)
0
200
400
600
800
0 200 400 600 800

Step size of KIF1A is 8×n nm, and varies stochastically.
02468
10
-32 32160-16step (nm)
12

Plus-end biased binding of KIF1A to the microtubule: ensemble average
Time (s)
on
0
2
4
-0.2 0 0.2
// MT⊥ MTControl
+-end
//^

Flush Ratchet Model
α β α β α βα β
- +
KIF1AATP
Phosphate
drelease ~0 nm
Dwell Detach
Load F Drag -gv
1D-diffusion under load
dbind ~3 nm
ADPBind
① ②
③④


Microtubule Structure Kikkawa et al.
JCB, 1994: Nature, 1995
Microtubule Microtubule-Kinesin complex

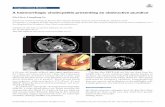
KIF1A-Microtubule Complex: cryo EM(15Å resolution)
Kikkawa et al. Cell 100: 241-, 2000

KIF1A-Microtubule Complex: cryo EM(15Å resolution)





ADP ATP(AMPPNP)
View from outside

ADP ATPView from outside


Tarzan Model
“Drag force” = diffusion might reflect the fluctuation of theflexible tether between KIF1A and tubulin (K-loop & E-hook).
TarzanTarzan

k i
Kikkawa et al. Nature 411: 439- ,2001


Structural Biological Analysis of kinesin motors
KIF1A alternately uses two loops to bind microtubules
Nitta,R. et al. Science 305:678- , 2004

Construction of KIF1A Motor Domain

Chemo-mechanical cycle of kinesin motors

Overall architecture of KIF1A
ATP(AMPPNP)
ADP-Pi (1)(ADP-AlF3)
ADP-Pi (2)(ADP-VO4)
ADP

Conformational changes of two switch regions

KIF1A*=ATP- Microtubule

KIF1A=ADP-Pi -Microtubule

KIF1A*=ADP-Pi -Microtubule

KIF1A=ADP -Microtubule

Nucleotide construct
wild type L12 L11 L8
AMPPNP 4.2±1.3 6.0±1.4 20.2±4.0 25.0±6.0
ADP 6.8±2.5 23.5±8.4 12.3±4.0 26.5±5.0
ATP 10.8±1.8 40.5±11.8 nd nd
ADP-AlFx 5.9±1.5 7.1±1.7 nd nd
ADP-Vi 21.4±4.3 167±66 nd nd
Equilibrium Binding Constants of KIF1A mutant

Design principle
of Walker-
type NTPase
s

Design principle of Walker-type NTPases

Model for the processive movement of the monomeric kinesin KIF1A



Univ. Tokyo
Takao NAKATAYoshimitsu KANAIYasuko NODASumio TERADAYosuke TAKEISen TAKEDAYasushi OKADAYosuke TANAKAMasahiko KAWAGISHIMitsutoshi SETOU
Laurent GUILLAUDRichard WONGJunlin TENGHarukata MIKIDae-Hyung SEOGChunjie ZHAOTerunaga NAKAGAWAMasahide KIKKAWAHiroaki YAJIMARyo NITTATadayuki OGAWA
UCSFRobert J. FletterickElena P. Sabrin
Univ. Hospital TokyoJunko TAKITAYasuhide HAYASHI
Niigata Univ. Brain Res. Inst.
Masaaki SAITOShoji TSUJI