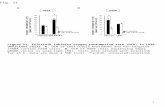
Expression Levels in Treated and Untreated Arabidopsis Thaliana
description
Transcript of Expression Levels in Treated and Untreated Arabidopsis Thaliana

Expression Levels in Treated and Untreated Arabidopsis ThalianaBree Dahle, Desiree Gates, & Tyler Healy
MATH 2040 Fall Semester 2010
Purpose
Methods
Results Conclusion
The purpose of this study is to observe the differences in gene expression of Rubisco Large, EF-1α, and EPSP of treated and untreated A. thaliana. We believe that the expression levels of Rubisco Large and EPSP will be higher in treated samples versus the untreated samples. We also believe that the expression levels of EF-1α will stay the same between treated and untreated samples because it is our internal control.
We homogenized A. thaliana leaves and extracted and purified the RNA using a hybrid RNeasy/Trizol method. The RNA was then synthesized into complementary DNA (cDNA) which was then run in the ABI 7500 Fast Real Time System. The expression levels on the genes were observed and critical threshold (Ct) values were calculated by the machine based on the absorbance versus cycle number. We then averaged the Ct’s for the untreated samples and made this our hypothesized average. The same was done to treated samples and these were our test averages. T-tests were done on all the sample averages which we then used to determine if we would reject or fail to reject our null hypotheses.
Hypotheses
EPSP:Ho: treated µ ct value ≤ untreated µ ct valueHA: treated µ ct value > untreated µ ct value (claim)Rubisco Large:Ho: treated µ ct value ≤ untreated µ ct valueHA: treated µ ct value > untreated µ ct value (claim)EF-1α:Ho: treated µ ct value = untreated µ ct value (claim)HA: treated µ ct value ≠ untreated µ ct value
Difficulties
-2.353 2.353
EF-1α
2.353
Rubisco Large and EPSP
SampleSample mean
Hypothesized mean
Sample Standard Deviation
(s)
Sample Size
t Critical Value
t Test Value
EF-1α 26.016 30.439 0.185 4 -2.353, 2.353 -47.816
Rubisco Large
18.817 20.123 0.292 4 2.353 -8.945
EPSP 43.79 37.629 0.895 4 2.353 13.768
Some of the difficulties that we encountered were:•Unreliable RNA•Human Error•Not enough data available•Wanted to do a 2-Sample Dependent Test•Not enough sample replicates
For the hypothesis that the treated and untreated samples of EF-1α will stay the same, we reject the Ho because the test statistic, -47.816, falls in the rejection regions of less than -2.353 or greater than 2.353. The data for EF-1α does not support the claim that the Ct value will stay the same between the treated and untreated samples. For the hypothesis that the treated samples of EPSP will be greater than the untreated samples, we reject the Ho because the test statistic, 13.768, falls in the rejection region of greater than 2.353. The data for EPSP does not support the claim that the Ct value for the treated samples would be greater than the Ct values of the untreated samples. For the hypothesis that the treated samples of Rubisco Large will be greater than the untreated samples, we fail to reject the Ho
because the test statistic, -8.945, does not fall in the rejection region of greater than 2.353. The data for Rubisco Large supports the claim that the Ct value for the treated samples would be greater than the Ct values of the untreated samples.
http://www.google.com/imgres?imgurl=http://users.rcn.com/jkimball.ma.ultranet/BiologyPages/A/arabidopsis.jpg&imgrefurl=http://users.rcn.com/jkimball.ma.ultranet/BiologyPages/A/Arabidopsis.html&usg=__HFWer2rBT_-J-xnF8dt9pdnDXDM=&h=368&w=279&sz=21&hl=en&start=0&zoom=1&tbnid=DTG2Aa6na4V_EM:&tbnh=170&tbnw=129&prev=/images%3Fq%3Darabidopsis%2Bthaliana%26um%3D1%26hl%3Den%26client%3Dfirefox-a%26rls%3Dorg.mozilla:en-US:official%26biw%3D1600%26bih%3D1010%26tbs%3Disch:1&um=1&itbs=1&iact=rc&dur=338&ei=dQwATaWLOdGpnQfo-IToDQ&oei=dQwATaWLOdGpnQfo-IToDQ&esq=1&page=1&ndsp=47&ved=1t:429,r:14,s:0&tx=37&ty=131