Document S1. Fifteen Figures, Supplemental Experimental ...
-
Upload
duongxuyen -
Category
Documents
-
view
233 -
download
3
Transcript of Document S1. Fifteen Figures, Supplemental Experimental ...
Cell, Volume 133
Supplemental Data
TNF-α Induces Two Distinct
Caspase-8 Activation Pathways Lai Wang, Fenghe Du, and Xiaodong Wang
Supplemental Experimental Procedures
Reagents
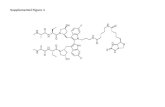
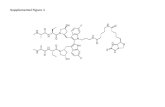
Smac mimetic and the biotinylated compound were synthesized as previously described
(Petersen et al., 2007). z-VAD and MG-132 was obtained from Calbiochem. Cycloheximide was
obtained from Sigma. The following antibodies were used for western-blot: caspase-3 (Cell
Signaling, 9662), caspase-8 (Cell Signaling, 9746), caspase-8 (BD Bioscience, 556466), c-FLIP
(Axxora, ALX-804-428), cIAP1 (R&D, AF8181), cIAP2 (BD Biosciences, 552782), CYLD (Axxora,
ALX-210-910), FADD (Stressgen, AAM-212E), Flag (Sigma, A8592), HA (Roche, 12013819001),
IκB-α (Santa Cruz Biotech, SC-371), Pan cIAP (R&D, MAB3400), PARP (Cell Signaling, 9542),
RIPK1 (BD Biosciences, 610458), RIPK1 (BD Biosciences, 551041), TNFRI (Santa Cruz Biotech,
SC-8436), TRADD (BD Biosciences, 610572), XIAP (BD Biosciences, 610717).
Plasmids and siRNA Oligoes
c-FLIP cDNA was amplified from a Hela cDNA library with primers containing a N-
terminal Flag epitope. It was then cloned into pCI-neo plasmid (Promega). HA-RIP construct was
generated in a similar fashion, but with an N-terminal HA epitope. FADD-DN (amino acid 81-208
of FADD) cDNA was also amplified using Hela cDNA library and subsequently cloned into a
modified pCI-neo plasmid, resulting in a fusion protein containing three Flag epitopes at the N-
terminus. p3xFLAG-CMV-7 mammalian expression plasmids for cIAP1 and cIAP2 as well as their
E3 ligase mutants H588A and H574A, respectively, were kindly provided by Dr. Chunying Du
(Stowers Institute). cIAP1 with a C-terminal Flag tag and its truncation constructs were amplified
with specific primers accordingly and subcloned into pCI-neo plasmid. For RIPK1-shRNA
construct, eight copies of RIPK1 shRNA (sequence as 5’-ccactagtctgacggataa-3’) were cloned into
pSuperior.puro vector using the protocol as previously described (Zhong et al., 2005). To generate
RIPK1 scramble expression constructs, RIPK1 cDNA with a C-terminal Flag epitope was first
subcloned into a modified pcDNA4/TO vector (Invitrogen) containing neomycin resistant gene. The
shRNA targeting site on RIPK1 was then mutated at six different sites without affecting amino acid
sequence. For kinase-dead version of RIPK1, lysine 45 was further mutated into alanine.
All siRNAs were purchased from Dharmacon. c-FLIP, caspase-8, RIPK1, and TRADD
were ON-TARGETplus siRNA pools of 4 oligoes. FADD was siGENOME siRNA pools of 4
oilgoes. For cIAP1, cIAP2, XIAP, and CYLD, individual oligo was used (cIAP1 target sequence 5’-
TTCGTACATTTCTCTCTTA-3’; cIAP2 target sequence 5’-AATGCAGAGTCATCAATTA-3’;
XIAP target sequence 5’-CCAGAATGGTCAGTACAAA-3’; CYLD target sequence 5’-
GAAGGCUUGGAGAUAAUGA-3’).
Recombinant Proteins
GST-TNF-α bacterial expression plasmid was kindly provided by Dr. Zhijian Chen (UT
Southwestern Medical Center at Dallas) and GST-TNF-α protein was generated as previously
described (Ea et al., 2006). GST-cIAP1 and GST-cIAP2 was expressed in E. coli strain BL21 (DE3)
and purified using Glutathione Sepharose 4B beads (Amersham Biosciences).
Cell Culture
Panc-1 pancreatic carcinoma, T98G glioblastoma, U-2OS osteosarcoma, MIA PACA-2
pancreatic carcinoma, and HEK-293T cells were obtained from ATCC and were cultured in DMEM
supplemented with 10% fetal bovine serum (FBS, Sigma) and 100-units/ml penicillin/streptomycin
(Hyclone). H2009 non-small cell lung carcinoma cells were obtained from the laboratory of Dr.
John Minna (UT Southwestern Medical Center at Dallas). Jurkat leukemia cells and RIPK1
deficient Jurkat cells were kindly provided by Dr. Adrian Ting (Mount Sinai School of Medicine,
New York, NY). H2009 cells and Jurkat cells were grown in HyQ RPMI-1640 medium (Hyclone)
supplemented with 10% FBS and 100-units/ml penicillin/streptomycin.
Methylene Blue Staining
For Supplementary Figure 2A, cell survival was accessed by methylene blue staining.
Briefly, cells were treated as indicated in figure legend. After 48 hours, cells were washed twice
with PBS (Invitrogen) and stained with 2% methylene blue (w/v) in 50% ethanol for 15 minutes with
shaking at room temperature (RT). Cells were then washed twice with PBS and air-dried before
being photographed.
Generation of Stable Cell Lines
To generate c-FLIP or FADD-DN overexpression cells, linearized plasmids were transfected
into Panc-1 cells as previously described. Forty-eight hours post transfection, cells were split and
placed in complete medium containing 1 mg/ml G418 (Calbiochem). After 2 to 3 weeks, clones
were lifted and tested for expression of the transgene. To generate H2009 tetracycline repressor
expression cells, an expression plasmid containing tetracycline repressor (TetR) was transfected into
H2009 cells and cells were selected with 10 μg/ml blasticidine (Invitrogen). Inducible RIPK1-
shRNA construct was then stably introduced into H2009-TetR cells, selected with 1 μg/ml
puromycin (Calbiochem). Finally, inducible wild type or kinase-dead version of RIPK1 rescue cells
were generated by transfecting RIPK1-shRNA stable cells with RIPK1 scramble expression
constructs and selected with 1 mg/ml G418.
Fluorogenic Caspase-3 Activity Assay
Cells were treated and subsequently pelleted. Cell pellet was resuspended in Buffer A (20
mM HEPES, 10 mM KCl, 1.5 mM MgCl2, 1 mM EDTA, 1 mM EGTA, 1 mM DTT, 0.1 mM PMSF,
and Complete Protease Inhibitor; Roche). The resuspended cell pellet was incubated on ice for 20
minutes before cells were broken by freezing in liquid nitrogen followed by thawing in 37°C
waterbath repetitively for 3 times. The resulting broken cell mixture was centrifuged at 20,000 g for
30 minutes. Protein concentration of the supernatant was determined by Commassie Plus Protein
Assay Kit (Pierce). Twenty micro-grams of cell lysates were incubated with 10 µM fluorogenic
caspase-3 substrate (Calbiochem) in a 20 μl system. Fluorescence reading was carried out as
previously described (Nijhawan et al., 2003).
Preparation of Cell Lysates with Lysis Buffer
Cell pellet was resuspended in lysis buffer (20 mM Tris-HCl, pH 7.4, 150 mM NaCl, 10%
glycerol, 1% Triton X-100, 1 mM Na3VO4, 25 mM β-glycerol-phosphate, 0.1 mM PMSF, and
Complete Protease Inhibitor) and vortexed for 20 seconds. The resuspended cell pellet was
incubated on ice for 20 minutes and centrifuged at 20,000 g for 30 minutes. Supernatant was
collected for further analysis.
Supplemental References
Ea, C.K., Deng, L., Xia, Z.P., Pineda, G., and Chen, Z.J. (2006). Activation of IKK by TNFalpha requires site-specific ubiquitination of RIP1 and polyubiquitin binding by NEMO. Mol Cell 22, 245-257. Micheau, O., and Tschopp, J. (2003). Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell 114, 181-190. Nijhawan, D., Fang, M., Traer, E., Zhong, Q., Gao, W., Du, F., and Wang, X. (2003). Elimination of Mcl-1 is required for the initiation of apoptosis following ultraviolet irradiation. Genes Dev 17, 1475-1486. Zhong, Q., Gao, W., Du, F., and Wang, X. (2005). Mule/ARF-BP1, a BH3-only E3 ubiquitin ligase, catalyzes the polyubiquitination of Mcl-1 and regulates apoptosis. Cell 121, 1085-1095.
Supplemental Figures
Figure S1. Smac mimetic induces TNF-α-dependent cell death in tumor cells from a variety of
tissue origins.
(A) T98G glioblastoma cells, (C) MIA PACA-2 pancreatic carcinoma cells, and (D) U-2OS
osteosarcoma cells were treated for 48 hours. (B) H2009 lung carcinoma cells were treated for 72
hours as indicated. Cell survival was determined by measuring ATP levels. Data were represented
as mean + standard deviation of duplicates. All experiments were repeated at least three times with
similar results.
Figure S2. TNF-α plus Smac mimetic induces caspase activation, and subsequent apoptosis.
(A) Panc-1 pancreatic carcinoma cells were treated with 100 nM Smac mimetic, 100 ng/ml TNF-α,
and/or 20 µM z-VAD for 72 hours. Cell viability was determined by methylene blue staining as
described in Experimental Procedures. (B) Same cell lysates used in Figure 1B were used for
caspase-8, caspase-3, PARP, and β-actin western-blot analysis. All experiments were repeated at
least three times with similar results.
Figure S3. Smac mimetic has no effect on TNF-α-induced NF-κB activation.
HEK-293T cells were transfected with a firefly luciferase reporter construct containing
three NF-κB binding sites together with a renilla luciferase plasmid as an internal control.
Forty-eight hour post transfection, cells were treated and were harvested after another 24
hours for luciferase activity using Dual-Glo Luciferase Assay System. Fold activation
was calculated by normalizing firefly luciferase activity/renilla luciferase activity of each
treatment to that of control treatment. Data were represented as mean + standard
deviation of duplicates. Experiments were repeated at least three times with similar
results.
Figure S4. c-FLIP cleavage requires caspase-8.
(A) Panc-1 cells were transfected with caspase-8 siRNA or control luciferase (Luc) siRNA. Forty-
eight hours post transfection, cells were treated with TNF-α plus Smac mimetic for three hours. Cell
lysates were collected and subjected to western-blot analysis of c-FLIP and β-actin levels. *
indicates a cross reactive band. (B) RNAi efficiency of caspase-8 siRNA. Forty-eight hours post
transfection, cell lysates from cells transfected with respective siRNA were collected and subjected
to western-blot analysis of caspase-8 and β-actin levels. All experiments were repeated at least two
times with similar results.
Figure S5. Elimination of c-FLIP mimics cycloheximide effect on TNF-α-induced apoptosis.
(A) Panc-1 cells were treated as indicated. Cell lysates were subject to western-blot analysis of
PARP and β-actin levels. (B) Panc-1 cells were pre-treated with cycloheximide or PBS vehicle for 1
hour prior to the treatment of TNF-α or PBS vehicle. After 48 hours, cell viability was determined
by measuring ATP levels. Data were represented as mean + standard deviation of duplicates. (C)
Panc-1 cells were transfected with c-FLIP siRNA or luciferase siRNA. Forty-eight hour post
transfection, cells were treated with TNF-α for the indicated time. PARP and β-actin levels were
measured by western-blot analysis. (D) c-FLIP knockdown efficiency. Forty-eight hours post
transfection, cell lysates from cells transfected with respective siRNA were collected and subjected
to western-blot analysis of c-FLIP and β-actin levels. * indicates a nonspecific cross reactive band.
(E) Forty-eight hour post transfection, cells were treated with or without TNF-α for additional 24
hours. Cell viability was determined by measuring ATP levels. Data were represented as mean +
standard deviation of duplicates. The effect of c-FLIP RNAi was verified using individual oligoes
from siRNA pool (data not shown). All experiments were repeated at least three times with similar
results.
Figure S6. RIPK1 is essential for TNF-α plus Smac mimetic induced
apoptosis in all cells examined.
(A) T98G cells, (B) H2009 cells, (C) MIA PACA-2 cells, and (D) U-2OS cells were transfected with
indicated siRNA. Forty-eight hours post transfection, cells were treated with or without TNF-α plus
Smac mimetic. For MIA PACA-2 cells, 10 ng/ml TNF-α was used to treat the cells. After
additional 48 hours, cell viability was determined by measuring ATP levels. Data were represented
as mean + standard deviation of duplicates. All experiments were repeated at least three times with
similar results.
Figure S7. The effect of RIPK1 knockdown on NF-κB activation.
H2009 RIPK1-shRNA cells were pre-treated with or without 0.03 mg/ml tetracycline for
72 hours prior to any additional treatment. (A) Cells were then treated with or without
Smac mimetic or TNF-α for another 72 hours. Cell viability was determined by
measuring ATP levels. Data were represented as mean + standard deviation of duplicates.
(B) Cells were treated with TNF-α for the indicated time and cell lysate were collected
for western-blot analysis of IκB, c-FLIP, and β-actin levels. Experiments were repeated
at least three times with similar results.
Figure S8. Knockdown of TRADD enhances RIPK1-dependent caspase-8 complex
formation.
Panc-1 cells were transfected with the indicated siRNAs. Forty-eight hours post
transfection, cells were treated with TNF-α plus Smac mimetic plus z-VAD for the
indicated time. Cells were harvested and analyzed for caspase-8 immunocomplex as
described in Experimental Procedures. Experiments were repeated twice with similar
results.
Figure S9. E3 ligase activity of cIAP ring finger domain is required for Smac mimetic induced
degradation.
HEK-293T cells were transfected with either wild-type or ring finger mutant forms of Flag-cIAP1 (A)
or Flag-cIAP2 (B) for 48 hours and then treated with Smac mimetic as indicated. Cell lysates were
collected at the indicated time points. Flag and β-actin antibodies were used for western-blot analysis.
Experiments were repeated at least three times with similar results.
Figure S10. cIAP1 is the major endogenous cIAP.
Western-blot analysis of cIAP1/2 levels using cell lysate and recombinant GST-cIAP fusion proteins.
(A) cIAP1 in 50 µg cell lysate is comparable to 2.5 ng of recombinant GST-cIAP1. (B) cIAP2 in 50
ug cell lysate is about half of 0.75 ng of recombinant GST-cIAP2. By calculation, endogenous cIAP1
is about 0.05% of total cellular protein while cIAP2 is only about 0.01%.
Figure S11. Triple knockdown of XIAP, cIAP1 and cIAP2 partially reproduces Smac
mimetic effect.
Panc-1 cells were transfected with luciferase (Luc), XIAP, cIAP1, and/or cIAP2 siRNAs. Forty-
eight hours post transfection, cells were treated for another 48 hours (A) or 3 hours (B). (A) Cell
viability was determined by measuring ATP levels. Data were represented as mean + standard
deviation of duplicates. (B) Cell lysates were collected and subjected to western-blot analysis of
PARP and β-actin. (C) XIAP, cIAP1 and cIAP2 RNAi efficiency. Forty-eight hours post
transfection, cell lysates from cells transfected with respective siRNA(s) were collected and
subjected to western-blot analysis of XIAP, cIAP1, cIAP2, and β-actin levels. All experiments
were repeated at least three times with similar results.
Figure S12. Mapping of RIPK1 or Smac mimetic binding sites on cIAP1.
HEK-293T cells were transfected with the indicated plasmids. Cells were harvested after 48 hours.
Cell lysates were immunoprecipitated with M2 Flag agarose beads (A), or avidin agarose beads in
the presence of 100 nM biotinylated Smac mimetic (B) as described in Experimental Procedures.
Flag, HA, and β-actin antibodies were used for western-blot analysis. * indicates a non-specific
cross reactive band. All experiments were repeated at least three times with similar results.
Figure S13. Model of TNF-α induced two caspase-8 activation pathways.
Upon TNF-α stimulation, Smac mimetic stimulates the release of RIPK1 from TNF-α receptor and
the formation of a RIPK1/FADD/caspase-8 death inducing signaling complex by activating cIAP
auto-degradation. This process requires the action of a RIPK1 deubiquitinating enzyme, CYLD.
On the other hand, cycloheximide or c-FLIP RNAi induces a RIPK1-independent caspase-8
activation pathway by removing c-FLIP inhibition (this pathway was modeled based on (Micheau
and Tschopp, 2003)). Details of this model are described in text.
Figure S14. Smac mimetic induces more FADD/caspase-8 complex formation than
cycloheximide.
Panc-1 cells were treated as indicated. Cycloheximide was added 1 hour prior to other
treatments. Cells were harvested 4 hours after treatment for caspase-8
immunoprecipitation. The levels of PARP and β-actin in the cell lysates and the levels of
caspase-8 and FADD in the immunocomplex were analyzed by western-blot.
Experiments were repeated three times with similar results.
Figure S15. TNF-α induced cell death in Jurkat cells.
Wild-type and RIPK deficient (RIPK1-/-) Jurkat cells were treated as indicated for 48 hours. (A) Cell
viability was determined by measuring ATP levels. Data were represented as mean + standard
deviation of duplicates. (B) Cell lysates were collected for westernblot analysis of RIPK1, c-FLIP
and β-actin levels. Experiments were repeated twice with similar results.